Towards Operational Intelligence: Machine learning for real-time aggregated prediction of hospital admission for emergency patients
Published in Healthcare & Nursing
To date, most applications of Artificial Intelligence (AI) to healthcare have been applied to clinical questions about diseases and patient care [1]. Now that many hospitals have electronic health records (EHRs), there is potential to use AI for operational purposes [2]. Patient flow in hospitals is difficult to plan for, because hospitals are highly connected systems in which capacity constraints in one area (for example, lack of ward beds) block the flow of patients from other locations. Management of emergency admissions is particularly complex, requiring bed managers to balance a known quantity (planned inpatient admission) with an unknown one (an uncertain number of emergency patients requiring beds).
In hospitals with EHRs, staff record patient data at the point of care, creating an opportunity to use real-time data for operational planning. In this study, we present a prediction pipeline that uses live EHR data for patients in a hospital emergency department (ED) to generate short-term, probabilistic forecasts of emergency admissions.
One advantage of using real-time EHR data for operational planning is that it can make use of accumulating data about each patient. To calculate each patient’s probability of admission, we trained 12 Machine Learning (ML) models on data available at successively longer elapsed times after the patient visit began. Figure 1 shows which features in the data were most important in each of the 12 models. At the outset, little is recorded about the patient other than age, arrival method, prior admission history and triage scores, so these are the most important. As the elapsed time increases, these diminish in importance as other signals of likely admission (like vital signs, lab results and consult requests) become available.
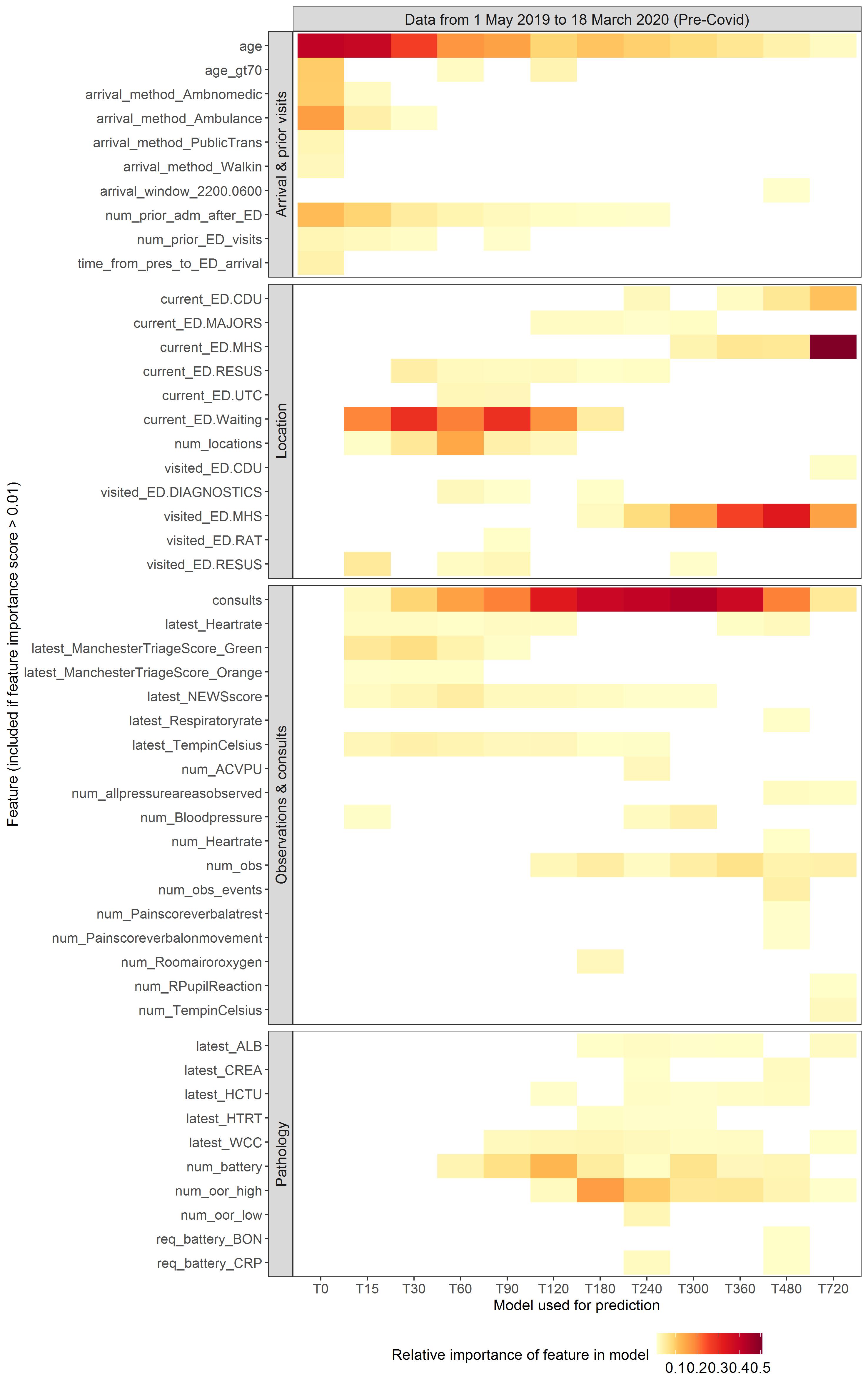
Figure 1: Feature importance for each of 12 ML models trained on data available at successively longer elapsed times since arrival
The vertical axis lists the features in each model, presented for ease of interpretation in four groups (visit data, location history, observations and consults, and pathology). See the paper for a glossary explaining these terms. The horizontal axis shows each of the 12 models. T0 was a model trained only on data available at 0 min after each patient’s arrival was first recorded in the EHR. T15 was trained on data available up to 15 min after arrival, and so on. The colour intensity reflects the relative importance of each feature within each model.
We worked closely with hospital bed managers to understand how to make these individual-level predictions most useful to them. From their point of view, knowing the probability that a particular patient will be admitted is less valuable than knowing in aggregate how many patients to plan for. In this respect, a prediction tool that can provide a probability distribution for the number of admissions in a given prediction window is more useful than one that solely estimates the probability of admission at the patient level. Also, such projections must allow for the number of patients not on the ED at the prediction time who will arrive later, and be admitted within the window.
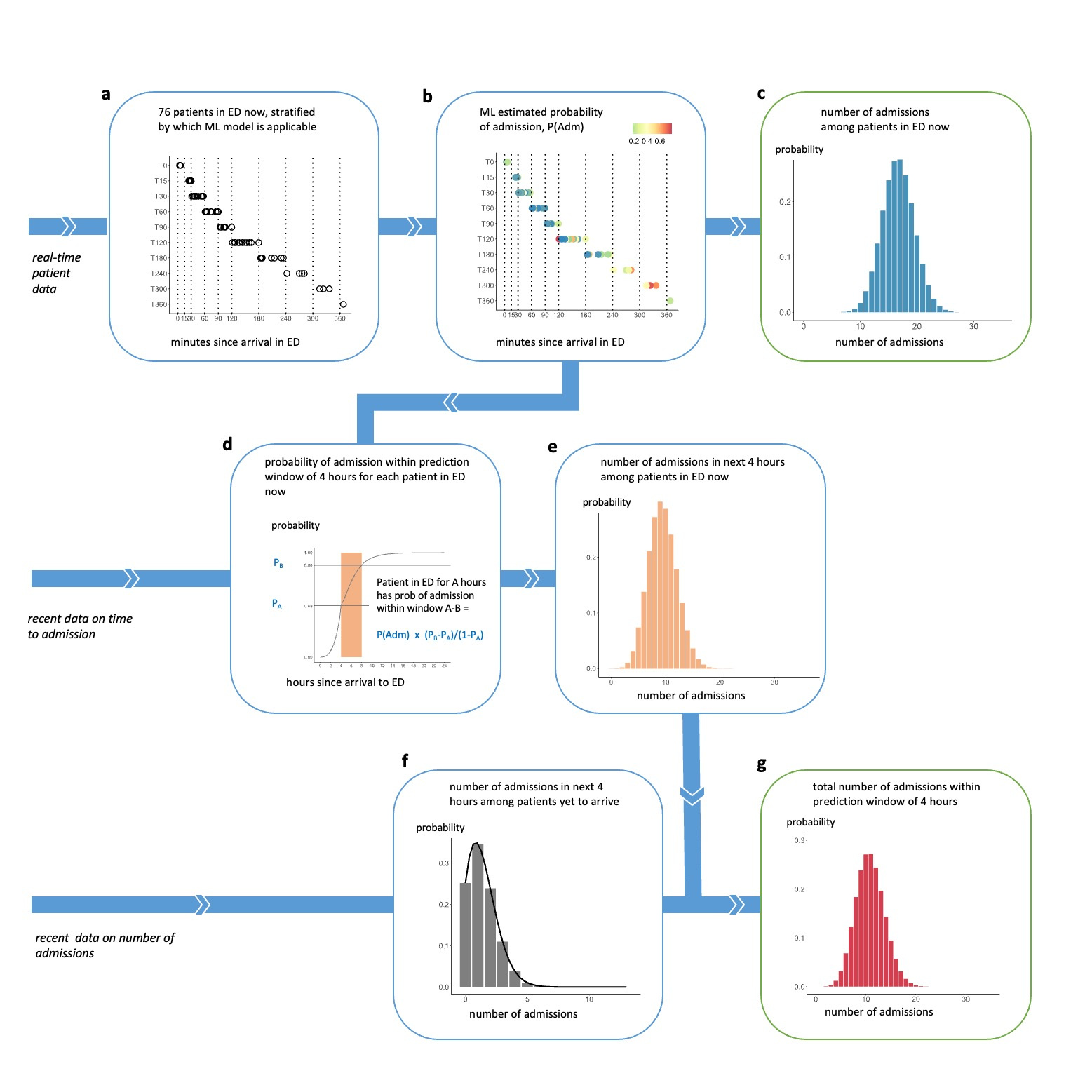
We therefore developed a pipeline that begins by applying ML to live data for each patient currently in the ED, as described above, and follows a series of steps to convert the ML predictions into aggregate predictions for the total number of admissions to plan for. The seven steps are shown in Figure 2, using a real example of patients in the ED at 16:00 on 11 May 2021.
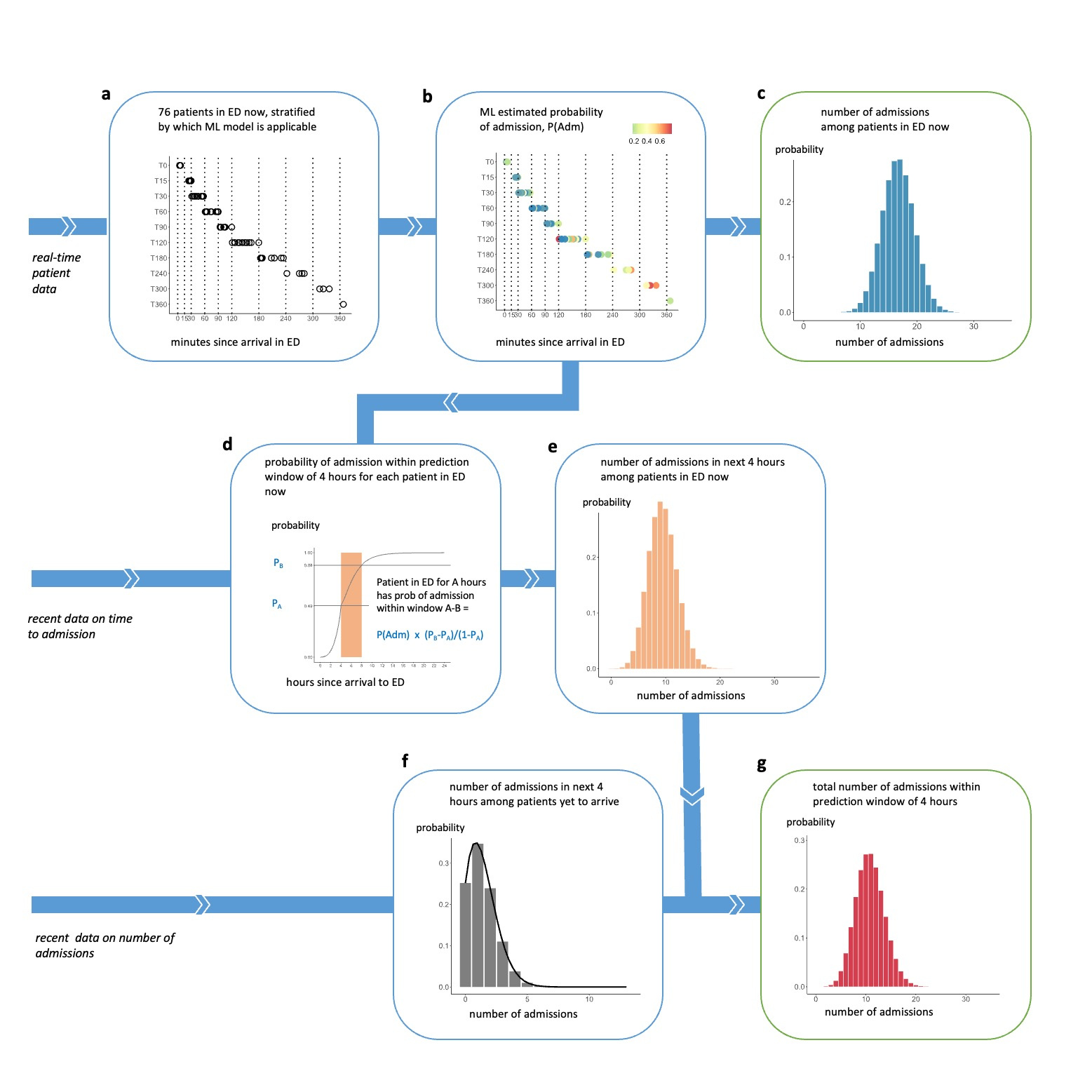
Figure 2 Illustration of the seven-step prediction pipeline using a real example, predicting the number of admissions within 4 hours after 16:00 on 11 May 2021
a illustrates the roomful of patients in the ED at 16:00, grouped according to how long they had been in the ED since arrival. b shows each patient’s probability of admission, generated using a set of ML models. These are combined in c into a probability distribution for the number of admissions among this roomful of patients. d shows how, for each patient, the probability of admission within the next 4 hours would be calculated from recent data on time to admission, taking into account the length of time the patient has already been in the ED prior to 16:00. e shows a probability distribution for the number of admissions among the roomful of patients within a prediction window of 4 hours. f shows a probability distribution over the number of patients who have not yet arrived, who will be admitted in the next 4 hours, generated by a Poisson equation from recent data on number of admissions. g shows the final probability distribution for the number of admissions, including patients in the now and those yet to arrive, within the prediction window of 4 hoursOur predictions outperformed a six-week rolling average benchmark that is conventionally used in the sector to predict daily admission numbers, as shown in Figure 3, and improved on the benchmark (which only projects up to midnight) by enabling predictions for short time-horizons of 4 and 8 hours at various times throughout the day.
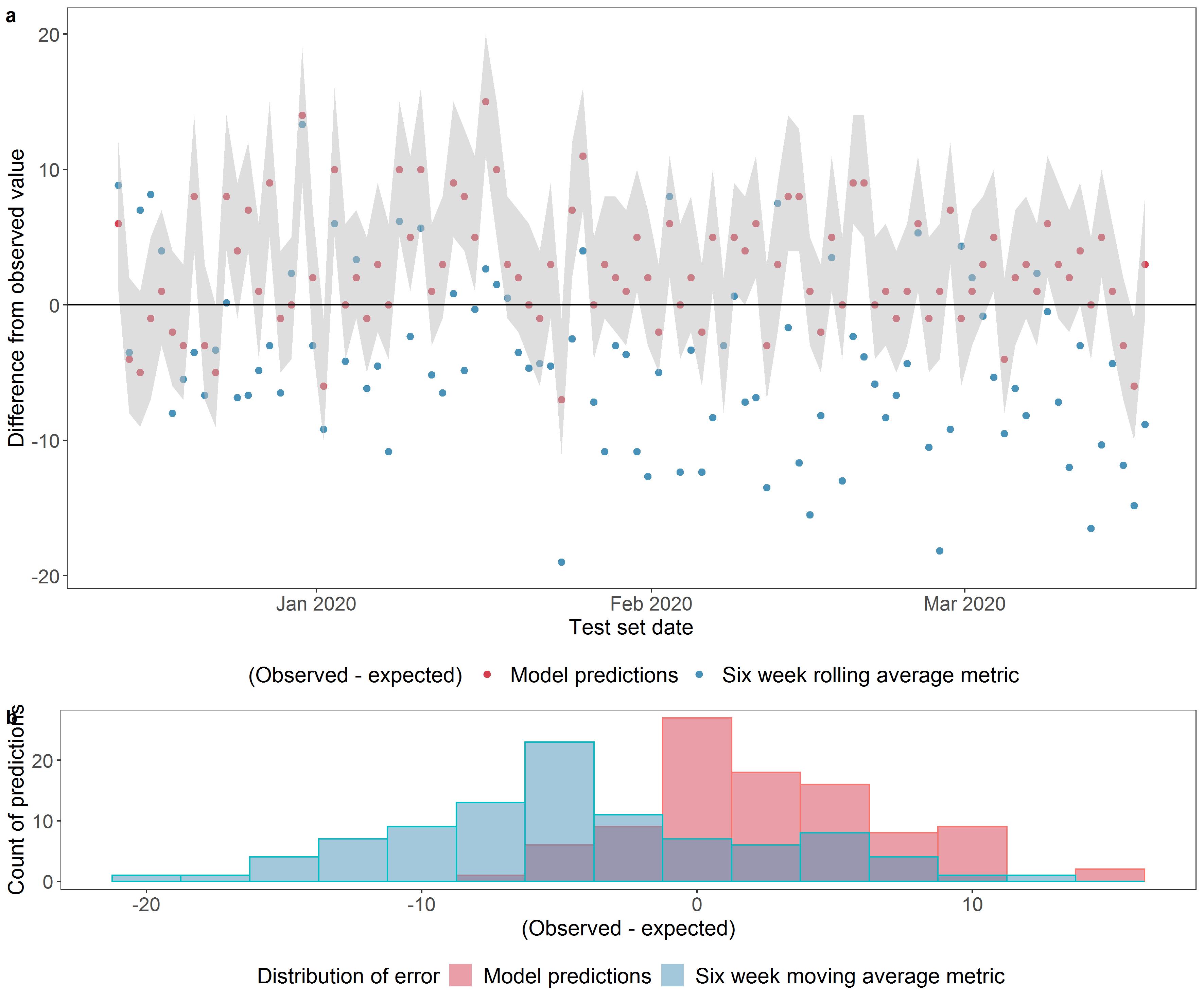
Figure 3: Comparing model predictions with a six-week rolling average benchmark for number of admissions within a prediction window of 8 hours after 16:00, including patients who are yet to arrive.
a shows the difference between the observed number of admissions and the expected value from the probability distribution for the pipeline model predictions (the red dots) and between the observed number of admissions and expected value from the benchmark (the blue dots). Where the expected value equals the observed value, the dots fall on the x axis (y = 0). The grey shaded band represents the range of probability between the 10th and the 90th centile of the cumulative probability distribution of the model. b shows the distribution of errors (difference between observed and expected). The pipeline model gave a mean absolute error (MAE) of 4.0 admissions (mean percentage error of 17%) versus 6.5 (32%) for a benchmark metric.
There are challenges associated with developing models for real-time implementation. The models cannot rely on data collated at the end of each day (as is often the case in hospital management reporting), or on data that is coded retrospectively (as often happens for coding of patient diseases). Our models achieved comparable performance to other studies, using only data available in real-time. If models are to be used operationally, their performance needs to be sustained even as care provision, patient characteristics and the systems used to capture data evolve. We found that operational variations in how long it took a patient to be admitted during the first year of the Covid-19 pandemic affected our time-window predictions. The staged nature of the seven-step pipeline enabled us to identify where data drift was occurring and retrain models to allow for this drift.
Real-time operational models need to cover the ‘last mile’ of AI deployment [3]; this means that the applications can run end-to-end without human intervention. This last mile is the most neglected, leading to calls for a delivery science for AI, in which AI is viewed as an enabling component within an operational workflow, rather than an end in itself [4]. We created an application to run four times daily, applying each step of the pipeline to generate predictions and emailing these to bed managers. An example of the email output is shown in the paper [5]. Our work provides a practical example of a ML-based modelling approach that is designed and fit for the purpose of informing real-time operational management of emergency admissions.
[1] Yu, K. H., Beam, A. L. & Kohane, I. S. Artificial intelligence in healthcare. Nat. Biomed. Eng. 2, 719–731 (2018).
[2] Pianykh, O. S. et al. Improving healthcare operations management with machine learning. Nat. Mach. Intell. 2, 266–273 (2020).
[3] Cabitza, F., Campagner, A. & Balsano, C. Bridging the “last mile” gap between AI implementation and operation: “data awareness” that matters. Ann. Transl. Med. 8, 501 (2020)
[4] Li, R. C., Asch, S. M. & Shah, N. H. Developing a delivery science for artificial intelligence in healthcare. Npj Digit. Med. 3, 1–3 (2020).
[5] The code for this project can be viewed at https://github.com/zmek/real-time-admissions
The contributors to this research are Zella King, Joseph Farrington, Martin Utley, Enoch Kung, Samer Elkhodair, Steve Harris, Richard Sekula, Jonathan Gillham, Kezhi Li, and Sonya Crowe.
The work was funded by grants from the Wellcome Institutional Strategic Support Fund (ISSF) UCL and Partner Hospitals and the NIHR UCLH Biomedical Research Centre. Some of the contributors were funded by the National Institute for Health Research and NHSX (see paper for award reference numbers). The views expressed in this publication are those of the authors and not necessarily those of the National Institute for Health Research, NHSX or the Department of Health and Social Care.
Follow the Topic
-
npj Digital Medicine
An online open-access journal dedicated to publishing research in all aspects of digital medicine, including the clinical application and implementation of digital and mobile technologies, virtual healthcare, and novel applications of artificial intelligence and informatics.
Related Collections
With Collections, you can get published faster and increase your visibility.
Evaluating the Real-World Clinical Performance of AI
Publishing Model: Open Access
Deadline: Jun 03, 2026
Impact of Agentic AI on Care Delivery
Publishing Model: Open Access
Deadline: Jul 12, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in