Unifying Multi-site MRI Harmonization Via Content-Style Representation Disentanglement
Published in Computational Sciences
The need for multi-site MRI harmonization
Structural MRI provides detailed high-resolution images of the brain's anatomy for visualizing different types of brain tissue, including gray matter, white matter, and cerebrospinal fluid. This in turn is crucial for identifying the size, shape, and integrity of various brain structures, especially for various brain abnormalities and diseases. Large-scale multi-site MRI studies are increasingly common, allowing much greater sample sizes not possible in the past decades (e.g., n > 10,000) for greater power in detecting subtle changes in association with brain development, aging, and diseases. However, harmonization of imaging data across sites remains a largely unsolved challenge, resulting in non-biological variation across scanners that negatively affects study outcomes.
Retrospective harmonization refers to the process of methodologically adjusting and standardizing MRI data collected from different sites or scanners to ensure consistency and comparability across all sites, aiming to minimize non-biological variability introduced by factors such as different scanner models, manufacturers, software versions, imaging protocols, and environmental conditions across study sites. Traditionally, statistics-driven methods are used to align intensity distributions of multi-site imaging data, but these methods are typically applied on the image globally without catering to variation in fine local details.
Advances in machine learning (ML) are suggesting the possibility of new paradigms in multi-site harmonization using image synthesis. One of the key advantages of ML-based methods, particularly those based on deep learning, is the automatization and standardization of tedious and time-consuming tasks.
Learning harmonization without paired data
Deep learning harmonization methods have been shown to significantly outperform classical methods in dealing with the complexity of anatomical details due to its prominent feature extraction capability. However, a crippling limitation of most deep learning methods to date is that they rely on supervised training and hence require scanning the same subjects (a.k.a. traveling phantoms) multiple times at the different sites. This is apparently very challenging, especially when images in sufficient quantity are needed in practice for reliable model training.
By contrast, unsupervised image-to-image translation techniques, such as CycleGAN, utilize unpaired images, which eliminates the necessity for paired data. However, these methods encounter scalability challenges in extensive multi-site scenarios, as they are designed to learn mappings between pairs of sites. This pair-wise approach is less effective because it fails to leverage the comprehensive information accessible across all sites. Additionally, these unsupervised strategies risk altering anatomical structures in the process of adjusting image contrast.
Our solution: Content-style disentanglement
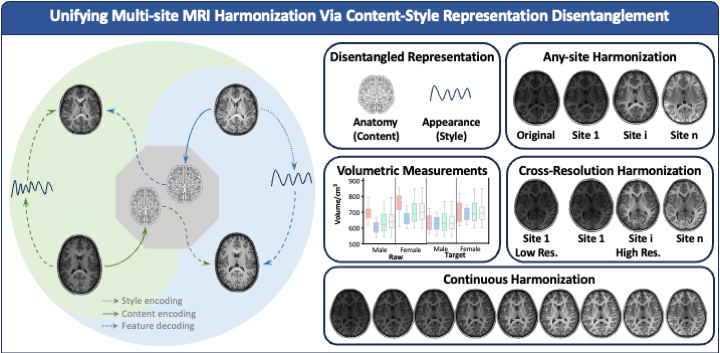
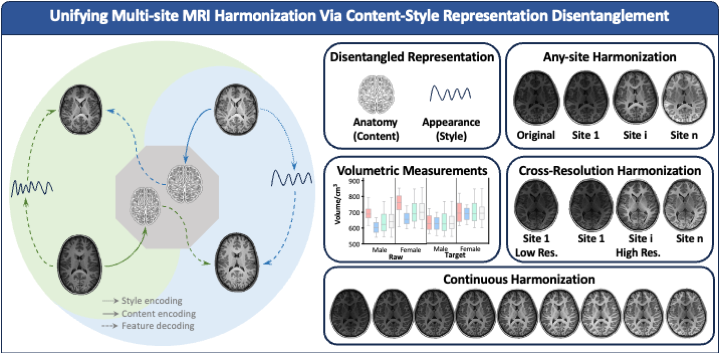
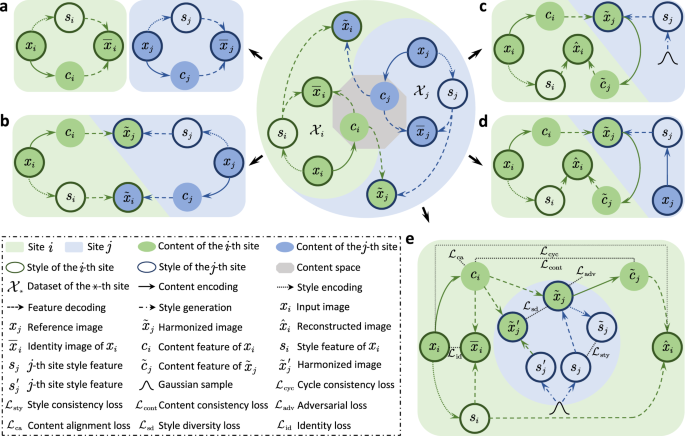
Our recent Communications Engineering paper "Learning Multi-site Harmonization of Magnetic Resonance Images Without Traveling Human Phantoms" presents a unified deep learning framework for simultaneous multi-site harmonization. Our method, multi-site unsupervised representation disentangler (MURD) treats simultaneous multi-site harmonization as a unified unsupervised image-to-image translation across multiple domains, decomposing an image into anatomical content that is invariant across sites and appearance style (e.g., intensity and contrast) that is dependent on each site. Harmonized images are generated by combining the content of an image with styles specific to the different sites, as illustrated in Figure 1. Our method requires only a small amount of unpaired data for highly efficient training and is therefore a game changer and allows data in a wide range of existing studies, conducted via different imaging protocols, to be harmonized without additional data collection.
Potential impact
MURD's ability to harmonize across sites without altering anatomical details could significantly improve the quality and consistency of MRI scans obtained from different locations. Such harmonization is crucial for accurate diagnosis and treatment planning, especially in cases where patients' scans are taken at different facilities. In large-scale multi-site clinical studies, such as those related to brain development or neurodegenerative diseases, ensuring consistent imaging data is essential. MURD allows for the retrospective harmonization of existing study data, enhancing the reliability and accuracy of research outcomes. MURD can also be instrumental in training AI models for medical image analysis as consistent and harmonized data are vital for developing algorithms that are accurate and generalizable across various imaging settings. Additionally, MURD enables the sharing of imaging data for second opinions or specialist consultations without the concern of variability due to differences in imaging equipment, promoting greater collaboration among healthcare providers. Besides, eliminating the need for human phantoms to travel between sites for the purpose of harmonizing MRI data reduces costs and logistical complexities. This efficiency can lead to faster diagnosis and treatment, benefiting both healthcare providers and patients.
Follow the Topic
-
Communications Engineering
A selective open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of engineering.
Related Collections
With Collections, you can get published faster and increase your visibility.
Generative AI for mechanical engineering design and optimization
Publishing Model: Open Access
Deadline: Jun 30, 2026
Engineering Solutions in Wind Energy Systems: Design, Efficiency, and Sustainability
Publishing Model: Open Access
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in