VinDr-CXR: The largest public chest X-ray dataset with radiologist-generated annotations for machine learning-based computer-aided diagnosis (CAD)
Published in Research Data
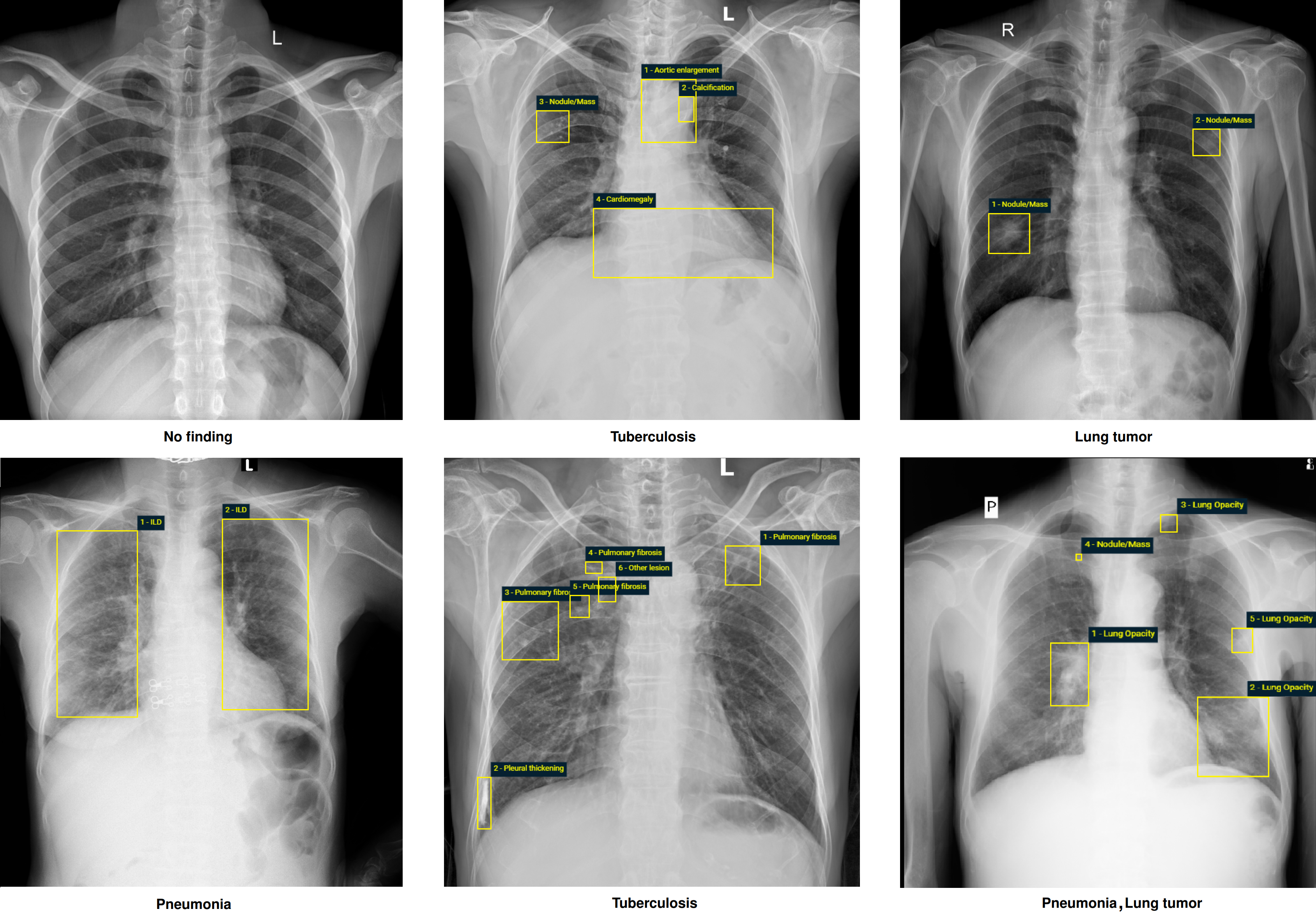
In 2022, we published the paper "VinDr-CXR: An open dataset of chest X-rays with radiologist annotations" in Scientific Data. We also released the whole dataset on PhysioNet. After just one year, the paper reached almost 100 citations and now became a standard, benchmarking dataset for developing and evaluating machine learning, deep learning, and computer vision for chest X-ray interpretation. Below we would like to share our story.
Why did we build the dataset?
Most existing chest radiograph datasets depend on automated rule-based labelers that either use keyword matching or an NLP model to extract disease labels from free-text radiology reports. These tools can produce labels on a large scale but, at the same time, introduce a high rate of inconsistency, uncertainty, and errors. These noisy labels may lead to the deviation of deep learning-based algorithms from reported performances when evaluated in a real-world setting. Furthermore, the report-based approaches only associate a CXR image with one or several labels in a predefined list of findings and diagnoses without identifying their locations.
There are a few CXR datasets that include annotated locations of abnormalities but they are either too small for training deep learning models or not detailed enough. The interpretation of a CXR is not all about image-level classification; it is even more important, from the perspective of a radiologist, to localize the abnormalities on the image. This partly explains why the applications of computer-aided detection (CAD) systems for CXR in clinical practice are still very limited.
We faced major challenges
Building high-quality datasets of annotated images is costly and time-consuming due to several constraints: (1) medical data are hard to retrieve from hospitals or medical centers; (2) manual annotation by physicians is both time-consuming and expensive; (3) the annotation of medical images requires a consensus of several expert readers to overcome human error; and (4) it lacks an efficient labeling framework to manage and annotate large-scale medical datasets.
Our approach
The building of the VinDr-CXR dataset is divided into three main steps: (1) data collection, (2) data filtering, and (3) data labeling. Between 2018 and 2020, we retrospectively collected more than 100,000 CXRs in DICOM format from local PACS servers of two hospitals in Vietnam.
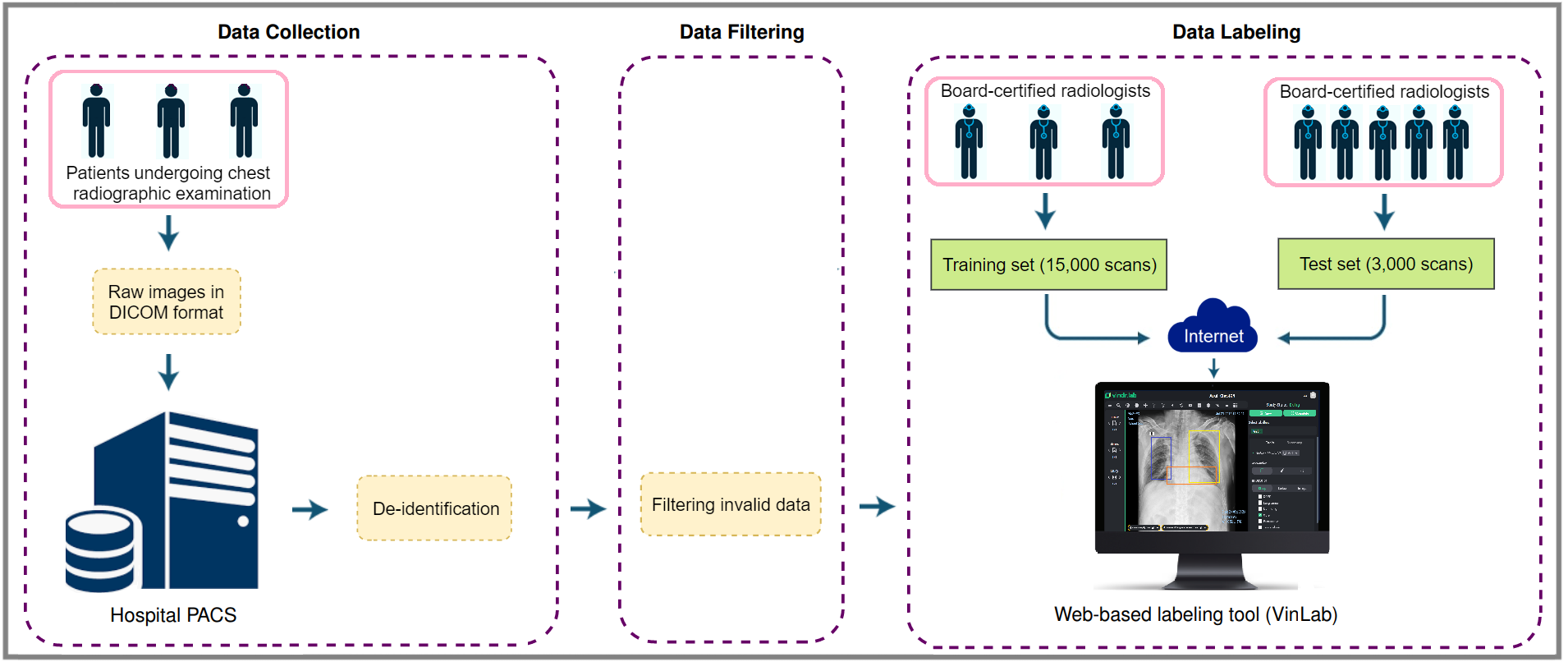
hospital’s PACS and got de-identified to protect patient’s privacy; (2) invalid files, such as images of other modalities, other body parts, low quality, or incorrect orientation, were automatically filtered out by a CNN-based classifier; (3) A web-based labeling tool, VinDr Lab, was developed to store, manage, and remotely annotate DICOM data: each image in the training set of 15,000 images was independently labeled by a group of 3 radiologists and each image in the test set of 3,000 images was labeled by the consensus of 5 radiologists.
About the dataset
The dataset contains more than 100,000 chest X-ray scans that were retrospectively collected from two major hospitals in Vietnam. Out of this raw data, we released 18,000 images that were manually annotated by a total of 17 experienced radiologists with 22 local labels of rectangles surrounding abnormalities and 6 global labels of suspected diseases. The released dataset is divided into a training set of 15,000 and a test set of 3,000. Each scan in the training set was independently labeled by 3 radiologists, while each scan in the test set was labeled by the consensus of 5 radiologists. All images are in DICOM format and the labels from training and test sets are made publicly available.
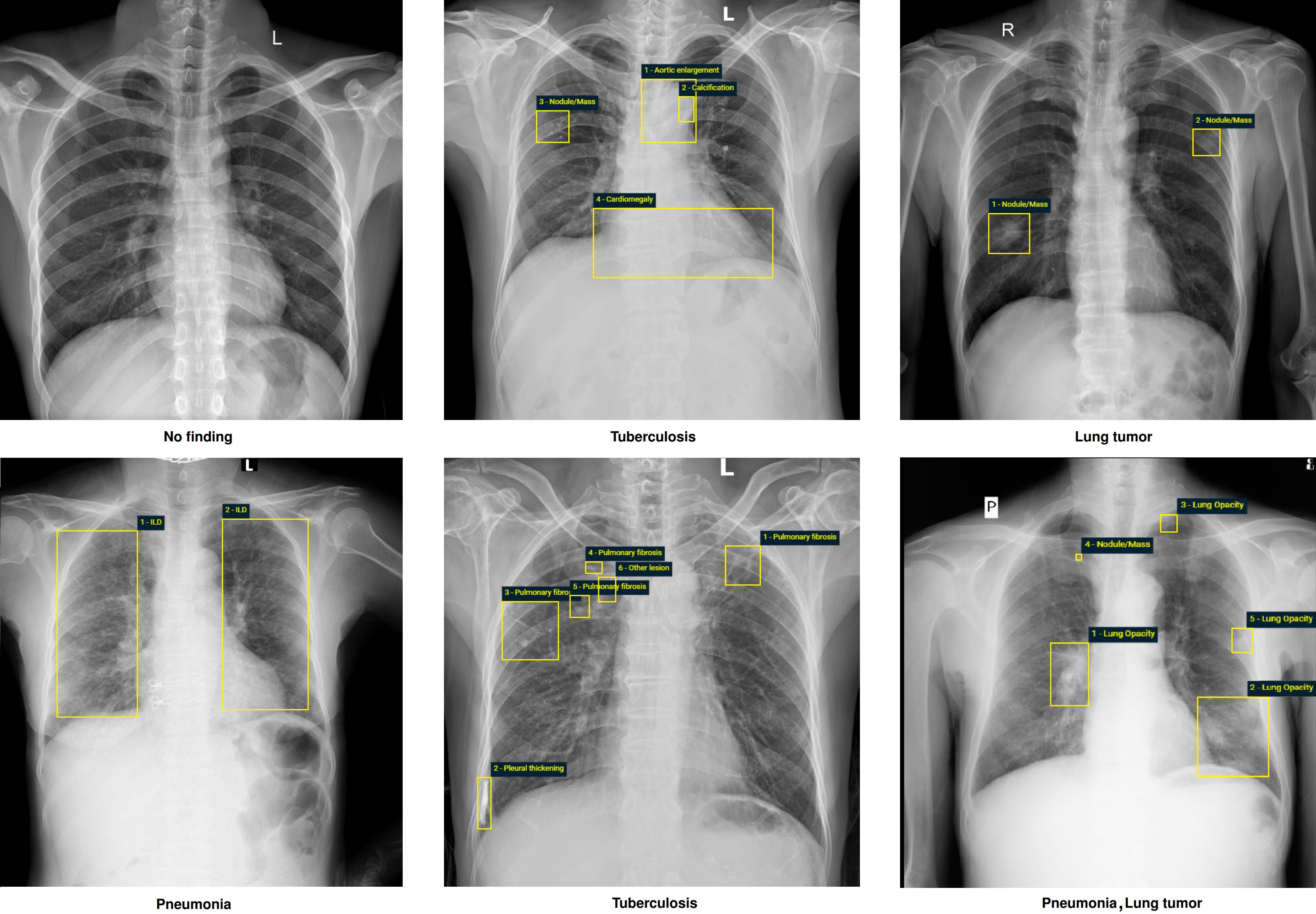
Again, we believe that large-scale, open, and high-quality data are the key to bringing medical AI algorithms to clinical settings and improving patient care. Besides the VinDr-CXR[1]. we commit our time and efforts to create more and more open datasets to release them to the research community. In 2023, we introduced the "VinDr-Mammo: A large-scale benchmark dataset for computer-aided diagnosis in full-field digital mammography" [2] and "PediCXR: An open, large-scale chest radiograph dataset for interpretation of common thoracic diseases in children" [2]. We also released novel data called VinDr-SpineXR: A large annotated medical image dataset for spinal lesions detection and classification from radiographs[3]. We believe that these imaging resources will play an important role in the development and validation of machine learning and deep learning algorithms for medical imaging research [4,5,6,7,8,9].
References
- Nguyen, Ha Q., et al. "VinDr-CXR: An open dataset of chest X-rays with radiologist’s annotations." Scientific Data 9.1 (2022): 429.
- Nguyen, Hieu Trung, et al. "VinDr-Mammo: A large-scale benchmark dataset for computer-aided diagnosis in full-field digital mammography." MedRxiv (2022): 2022-03.
- Nguyen, Hieu T., et al. "VinDr-SpineXR: A deep learning framework for spinal lesions detection and classification from radiographs." Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, September 27–October 1, 2021, Proceedings, Part V 24. Springer International Publishing, 2021.
- Pham, Hieu H., et al. "Interpreting chest X-rays via CNNs that exploit hierarchical disease dependencies and uncertainty labels." Neurocomputing 437 (2021): 186-194.
- Tran, Thanh T., et al. "Learning to automatically diagnose multiple diseases in pediatric chest radiographs using deep convolutional neural networks." Proceedings of the IEEE/CVF International Conference on Computer Vision. 2021.
- Pham, Hieu H., et al. "An Accurate and Explainable Deep Learning System Improves Interobserver Agreement in the Interpretation of Chest Radiograph." IEEE Access 10 (2022): 104512-104531.
- Nguyen, Ngoc Huy, et al. "Deployment and validation of an AI system for detecting abnormal chest radiographs in clinical settings." Frontiers in Digital Health (2022): 130.
- Nguyen, Huyen TX, et al. "A novel multi-view deep learning approach for BI-RADS and density assessment of mammograms." 2022 44th Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC). IEEE, 2022.
- Le, Khiem H., et al. "Learning from multiple expert annotators for enhancing anomaly detection in medical image analysis." IEEE Access 11 (2023): 14105-14114.
Follow the Topic
-
Scientific Data
A peer-reviewed, open-access journal for descriptions of datasets, and research that advances the sharing and reuse of scientific data.
Related Collections
With Collections, you can get published faster and increase your visibility.
Genomics in freshwater and marine science
Publishing Model: Open Access
Deadline: Jul 23, 2026
Genomes of endangered species
Publishing Model: Open Access
Deadline: Jul 01, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in