What can enzymes teach us about working together?
Published in Chemistry
Much of biochemical research relies on our knowledge of which enzymes catalyze which reactions. Consider the use of enzymes as biocatalysts, which is often limited by the variety transformations that can be catalyzed by known enzymes. Some great strides have been made expanding this “biocatalytic toolbox” through the use of directed evolution1, reinventing the scope of reactions catalyzed by known enzyme classes through mechanistic insight2,3, and de novo enzyme design4,5, but consider just how many enzymes are already encoded in the world’s genetic diversity that have never been studied! Countless new genomes are deposited in public databases every day, yet only a small fraction has been biochemically characterized6–9.
The field of Genome Mining aims to explore this “metabolic dark matter” for novel enzymes, as well as entire metabolic pathways, usually with the eventual goal of discovering new bioactive natural products10. Because of this, historically, genome mining efforts have been primarily guided by sequence homology to enzymes known to be involved in the biosynthesis of well-established medicinal natural products. This approach has been very successful in discovering enzymes involved in the biosynthesis of new members of known classes of molecules, such as polyketides, non-ribosomal peptides, RiPPs and terpenes.
But, what if you wanted to discover enzymes with truly new activities?
Previously, I would rely on my intuition: I would annotate pfam11 domains in a genome, and simply scroll through the whole genome until I noticed a pattern that I did not recognize as any primary or secondary metabolism familiar to me, after which I would dig through the literature to make sure the genes had not been studied before. This is how I initially discovered the enzyme responsible for the oxidative cyclization of prodigiosin to form cycloprodigiosin in the marine bacterium Pseudoalteromonas rubra12. This time around however, I wanted to formalize my workflow from one relying on intuition, into one that I could share with the world.
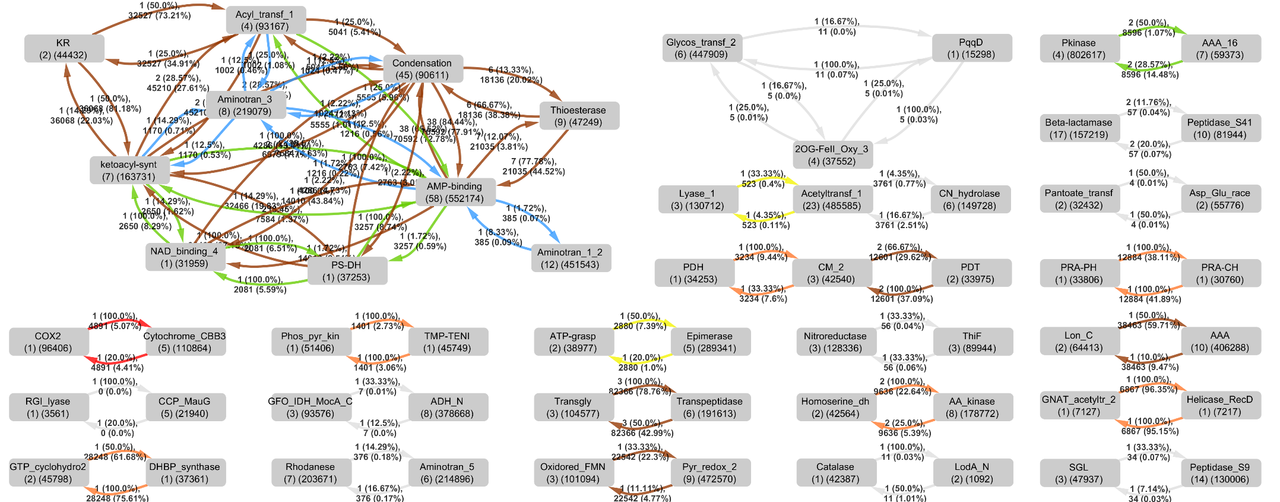
I focused on multi-domain enzymes due to their track record as impressive catalysts: think of the amazing chemistry catalyzed by polyketide synthases, nonribosomal peptide synthetases, or the enzymes recently found to catalyze reticuline epimerization13,14 and N-nitrosation15,16. I found that it helped me visualize the diversity of multi-domain enzymes if I drew them as a network. In parallel, I made a database of all annotated multi-domain enzymes. Now, instead of manually looking up every domain combination found in the literature, I was able to query the database and quickly color-code the networks based on annotation status.
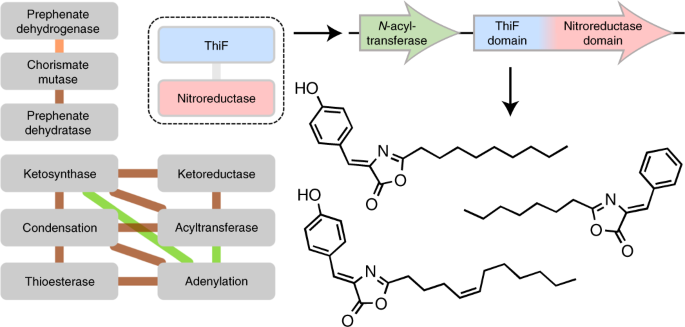
I subjected the genome of my old friend P. rubra to this algorithm, which I called “CO-ED”, and found that it coded for a di-domain enzyme whose combination of domains, a “ThiF” and a “Nitroreductase” domain, had never been characterized together before. These domains are known to catalyze rather interesting reactions by themselves. Imagine what they could do when working together!
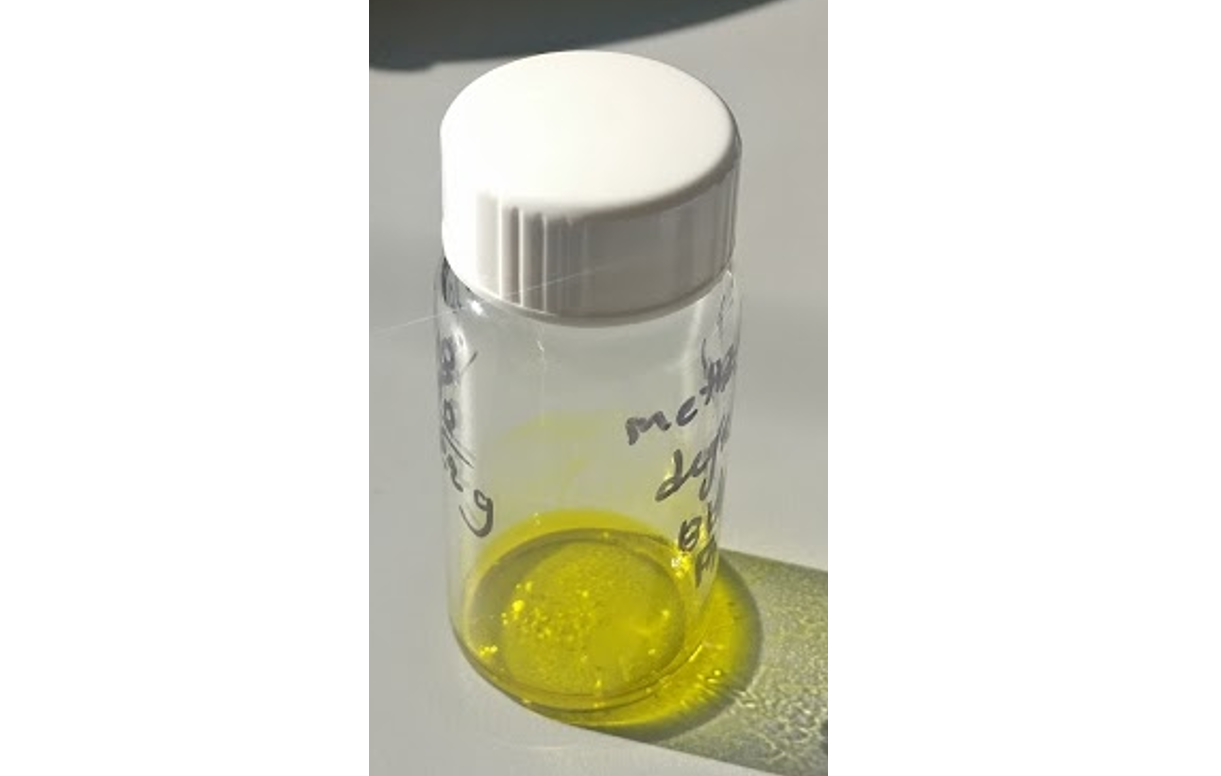
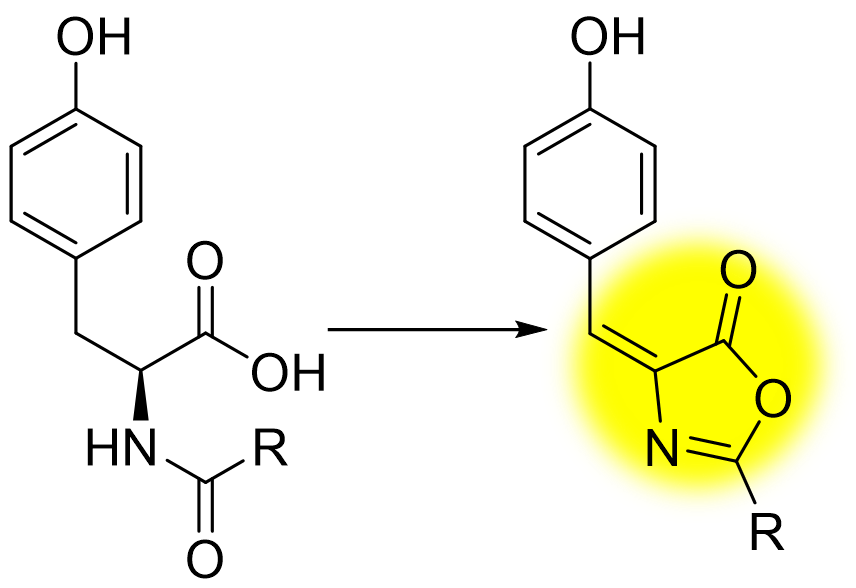
In P. rubra and other organisms, the gene coding for this enzyme (which we call oxzB) was consistently located adjacent to another gene, oxzA. Going forward, the project would require cloning the oxzAB gene pair, heterologously expressing these genes in E. coli, determining if any new metabolic products were produced and if so, isolating and identifying them. Believing this would be a valuable set of skills to learn for an undergraduate student, I recruited a talented undergraduate student, Julia Asay, who immediately got off on the right foot. Her first PCRs in the lab were successful, and she got colonies on her first transformation. While we were miniprepping the DNA to sequence-verify the constructs, we noticed a bright yellow color appear while adding “Solution #2” (the basic SDS solution to lyse the cells). Upon scaling the culture, we noticed that the bacterial pellets were somewhat yellow, which could be intensified by adding base (see figure), but the color would eventually fade due to degradation. It appeared that our heterologous construct was producing some sort of pigment that we hypothesized was phenolic. Adding extra tyrosine to the culture medium improved our yields of the metabolite, so we suspected tyrosine was its precursor.



Thanks to its bright color under basic conditions, we were able to follow our molecule throughout its purification by means of an unusual TLC “stain”: sodium hydroxide in ethanol, which would result in a startlingly bright yellow spot
NMR indeed showed a phenolic moiety, as well as what appeared to be a fatty acyl chain, however the structure connecting the two proved difficult to elucidate as it contained only a single hydrogen atom. After scaling up our heterologous expression, we were able to isolate 9 mg of the molecule, and through the use of a 1.7 mm cryoprobe were able obtain a solution that was sufficiently concentrated (over 0.5 M!) to perform 1,1-ADEQUATE and 1H-15N HMBC NMR experiments, which let us determine that the molecule was an oxazolone. While oxazolones are rare in nature, the existence of an oxazolone previously isolated from a red alga, almazolone17, gave us confidence in our assignment. Since our oxazolone was derived from tyrosine, we named it “tyrazolone”. Cloning several oxzAB gene pairs from other organisms showed that they too coded for tyrazolones, as well as phenylalanine-derived “phenazolones”. We also showed that the di-domain enzyme, OxzB, was responsible for forming an oxazolone heterocycle from an N-acyl amino acid, the first enzyme known to catalyze such a reaction, and a potentially promising biocatalyst.
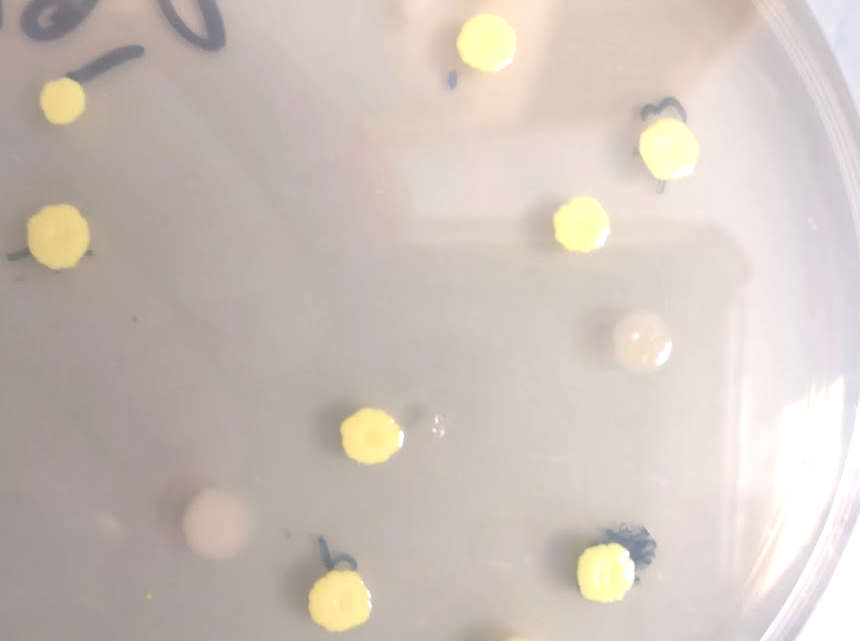
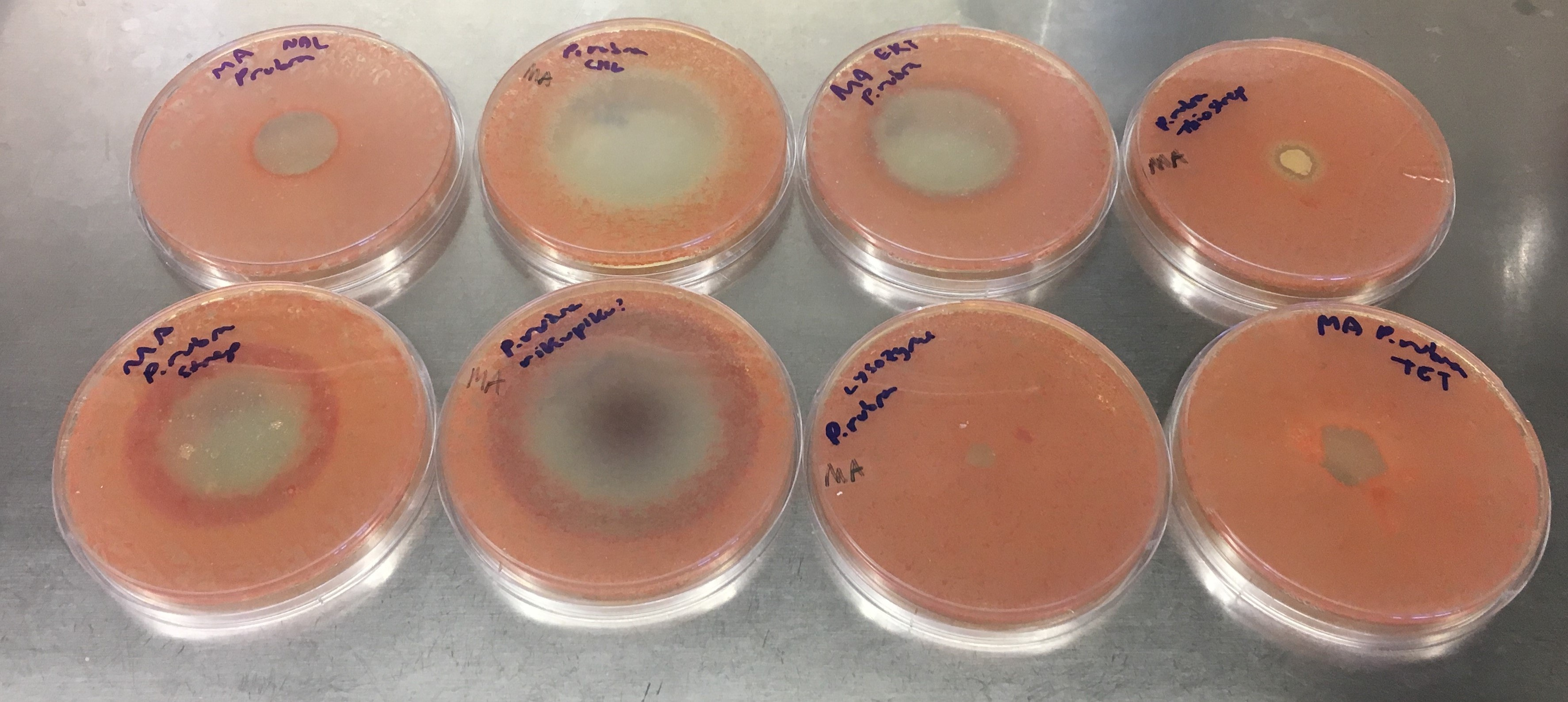
One question we had was whether P. rubra actually produces oxazolones, or if their formation was an artifact of heterologous expression. We were unable to detect any oxazolones in P. rubra growing under laboratory conditions, but it is known that sometimes organisms require a stressor to start producing a natural product18. I asked Julia to dig into the freezer, grab all antibiotics that she could find, and expose P. rubra to them (see figure). Satisfyingly, some antibiotics induced oxazolone production in P. rubra, verifying that they are indeed true natural products. We can only guess at the biological role of these molecules; they do not exhibit antibacterial activity, but we know their biosynthetic genes are present in several dozen sequenced (predominantly aquatic) bacteria. The fact that they are induced by antibiotic stress and that they are somewhat unstable suggests maybe they act as signals?
I find it fascinating how, P. rubra, an organism I had been studying for years, is capable of biosynthesizing molecules that had never been isolated before! There are thousands of unannotated multidomain enzymes encoded in publicly available genomes, and genome sequencing is not slowing down anytime soon. This project taught Julia hands-on skills in microbiology, molecular biology, biochemistry, organic chemistry and analytical chemistry. Each of those thousands of enzymes could be an opportunity for another student to similarly learn a myriad of new skills! A pre-computed CO-ED network for all proteins in Uniprot19 can be found as supplemental data to the manuscript, or you can run the CO-ED workflow on your own data at http://enzyme-analysis.org.
Sesame Street was right about working together: enzyme domains working together can achieve remarkable chemical transformations, working across scientific disciplines makes for intellectually rewarding research, and working together in the lab is a great learning opportunity!
For more information, please read our article in Nature Chemical Biology at https://www.nature.com/articles/s41589-021-00808-4
- Zeymer, C. & Hilvert, D. Directed evolution of protein catalysts. Annu Rev Biochem 87, 131–157 (2018).
- Chen, K. & Arnold, F. H. Engineering new catalytic activities in enzymes. Nat. Catal. (2020). doi:10.1038/s41929-019-0385-5
- Sandoval, B. A. & Hyster, T. K. Emerging strategies for expanding the toolbox of enzymes in biocatalysis. Curr Opin Chem Biol 55, 45–51 (2020).
- Kiss, G., Çelebi-Ölçüm, N., Moretti, R., Baker, D. & Houk, K. N. Computational enzyme design. Angew Chem Int Ed Engl 52, 5700–5725 (2013).
- Hilvert, D. Design of protein catalysts. Annu Rev Biochem 82, 447–470 (2013).
- Gerlt, J. A. et al. The enzyme function initiative. Biochemistry 50, 9950–9962 (2011).
- Hanson, A. D., Pribat, A., Waller, J. C. & de Crécy-Lagard, V. Unknown proteins and orphan enzymes: the missing half of the engineering parts list – and how to find it. Biochem J 425, 1–11 (2009).
- Schnoes, A. M., Brown, S. D., Dodevski, I. & Babbitt, P. C. Annotation error in public databases: misannotation of molecular function in enzyme superfamilies. PLoS Comput Biol 5, e1000605 (2009).
- Ellens, K. W. et al. Confronting the catalytic dark matter encoded by sequenced genomes. Nucleic Acids Res 45, 11495–11514 (2017).
- Medema, M. H., de Rond, T. & Moore, B. S. Mining genomes to illuminate the specialized chemistry of life. Nat Rev Genet (2021). doi:10.1038/s41576-021-00363-7
- El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic Acids Res 47, D427–D432 (2019).
- De Rond, T. et al. Oxidative cyclization of prodigiosin by an alkylglycerol monooxygenase-like enzyme. Nat Chem Biol 13, 1155–1157 (2017).
- Farrow, S. C., Hagel, J. M., Beaudoin, G. A. W., Burns, D. C. & Facchini, P. J. Stereochemical inversion of (S)-reticuline by a cytochrome P450 fusion in opium poppy. Nat Chem Biol 11, 728–732 (2015).
- Winzer, T. et al. Plant science. Morphinan biosynthesis in opium poppy requires a P450-oxidoreductase fusion protein. Science 349, 309–312 (2015).
- Ng, T. L., Rohac, R., Mitchell, A. J., Boal, A. K. & Balskus, E. P. An N-nitrosating metalloenzyme constructs the pharmacophore of streptozotocin. Nature 566, 94–99 (2019).
- He, H., Henderson, A. C., Du, Y.-L. & Ryan, K. S. A two-enzyme pathway links L-arginine to nitric oxide in N-nitroso biosynthesis. J Am Chem Soc 141, 4026–4033 (2019).
- Guella, G., N’Diaye, I., Fofana, M. & Mancini, I. Isolation, synthesis and photochemical properties of almazolone, a new indole alkaloid from a red alga of Senegal. Tetrahedron 62, 1165–1170 (2006).
- Seyedsayamdost, M. R. High-throughput platform for the discovery of elicitors of silent bacterial gene clusters. Proc Natl Acad Sci U S A 111, 7266–7271 (2014).
- The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res 46, 2699 (2018).
Follow the Topic
-
Nature Chemical Biology

An international monthly journal that provides a high-visibility forum for the chemical biology community, combining the scientific ideas and approaches of chemistry, biology and allied disciplines to understand and manipulate biological systems with molecular precision.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in