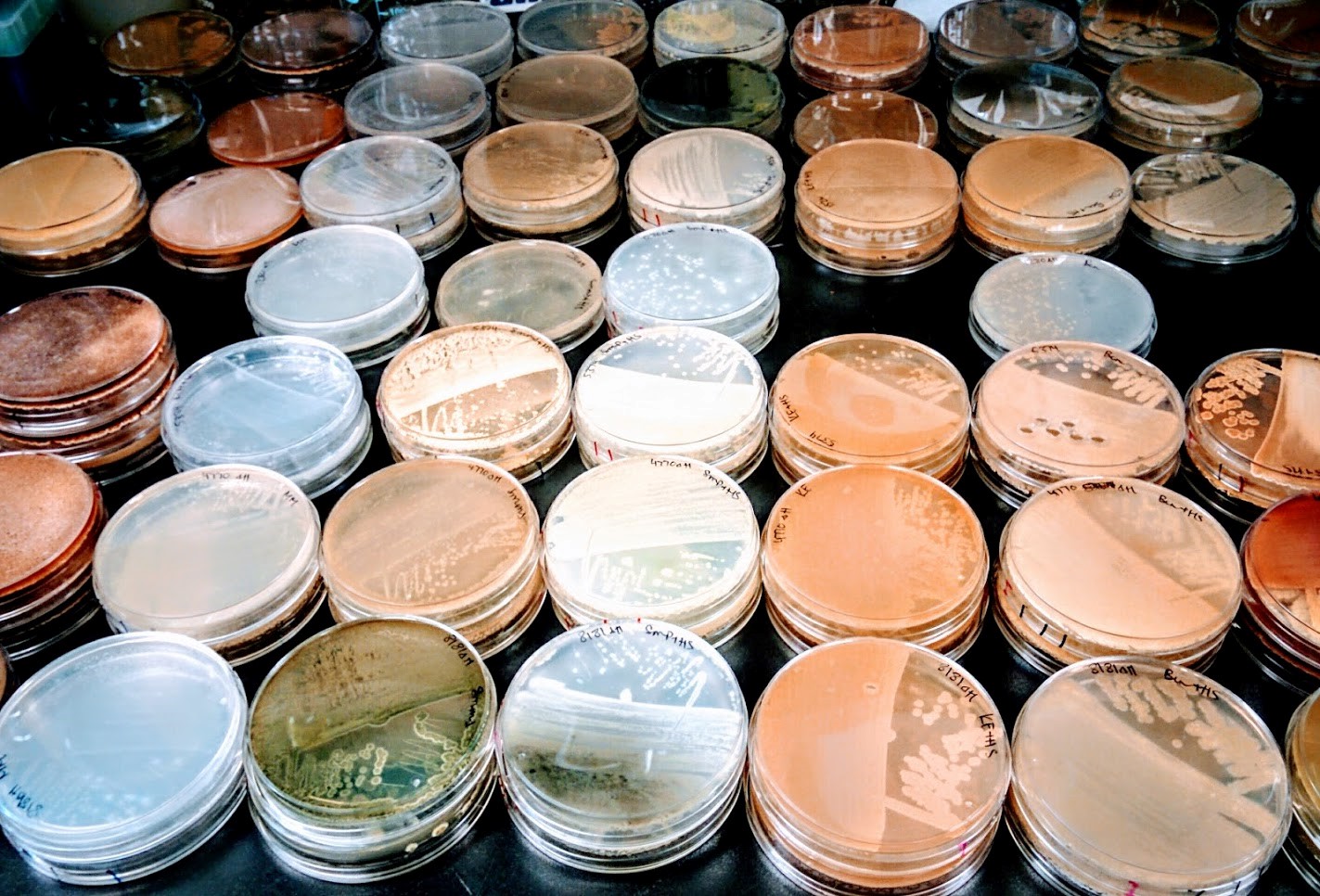
From the soil to the gut, bacteria in the environment do not live in monocultures grown in rich media, but in complex communities where they must outcompete their neighbours for resources and space. On the offensive side, antibiotics are one way a bacterium may inhibit growth of its neighbour and gain a fitness advantage. The bacterial order Actinomycetale is particularly gifted in the production of antibiotics, encoding them in regions of their genome called biosynthetic gene clusters (BGCs). These specialized small molecules display exquisite chemical complexity and biological specificity for their given target. Exploring these bacterial inventions offers humankind the opportunity not only to understand a day-in-the-life of a microbe, but also to take inspiration for the development of novel therapeutics. Thinking about what selective pressure might exist on a bacterium and how we can leverage these predictions in discovery strategies can be a fruitful and elegant solution for identifying novel bioactive compounds.
Setting out to use some of these ideas to discover new antibiotics, we were particularly interested in finding new compounds that targeted the highly conserved intracellular protease, ClpP. It seemed likely that such a compound might exist given the ubiquity of ClpP and the druggability of proteases in general. Furthermore, if we were to identify a suitable compound, it could be effective against the pathogen Mycobacterium tuberculosis, which unlike most other bacteria, requires ClpP indispensably for growth. Spoiler alert – the antibiotic we identified is probably too unstable to be useful in the clinic in its present form – but nonetheless this was our motivation.
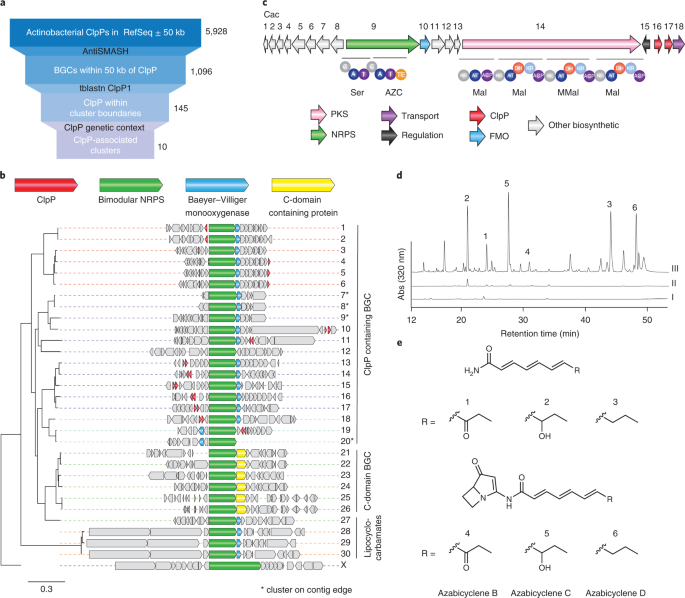
To identify compounds targeting ClpP, we were inspired by a previously developed technique called target-directed genome mining. This strategy makes use of that, to avoid self-intoxication, BGCs encoding antibiotics contain resistance genes that sometimes take the form of an antibiotic-insensitive copy of the targeted housekeeping protein. Therefore, by searching for these duplicated housekeeping genes, one can predict BGCs encoding antibiotics with a given function. This strategy lent itself well to ClpP-directed compounds, and our search turned up a large family of related BGCs all associated with ClpP and spread across many phyla of bacteria. We chose to work on one such BGC found in the organism Streptomyces cattleya as a representative.
In real life, this is where the project stalled and sat on the backburner for about four years. The BGC in S. cattleya was transcriptionally silent, and even once we were able to activate the cluster through heterologous expression and some promoter swapping gymnastics, the products we identified, a family of compounds called azabicyclenes, were not active against ClpP. In fact, azabicyclenes and their larger BGC family have been extensively characterized in the literature, yet nobody has previously reported any association or activity having to do with ClpP. This puzzled us to no end but driven by the conviction that ClpP was involved somehow, we finally came back to the project in time for the final 6 months of my PhD.
Since the easily detectable azabicyclene metabolites did nothing to ClpP, we searched for ClpP inhibition in the heterologous producer organism. These efforts were based on ClpP regulation in Streptomyces spp., many of which have five ClpP isoforms that form heterocomplexes and are regulated by ClpP-dependent degradation of transcription factors. By measuring which clpP homologs were turned on/off, we detected evidence of ClpP inhibition in the BGC containing cells. We predicted that the relevant metabolite was a covalent inactivator and thus unstable, and indeed could detect modification of ClpP in vitro through whole protein mass spectrometry.
At this point, while we could detect ClpP inhibition using spent media, we still found the inhibitor to be unstable and we always lost activity when we tried traditional activity-guided purification. Retrospectively, this makes sense given that nobody had previously noticed ClpP inhibition associated with these well known BGCs. To overcome its instability, we used the predicted mass of the inactivator to guide isolation, which allowed us to purify the molecule in a directed and gentler manner. In my last three weeks in the lab, we solved the structure of the resulting ClpP inhibitor, which we named clipibicyclene. This potent inactivator has a bicyclocarbamate warhead that undergoes nucleophilic attack of the active site Ser of ClpP. The easily detectable azabicyclenes were in fact red herrings, just shunt metabolites.
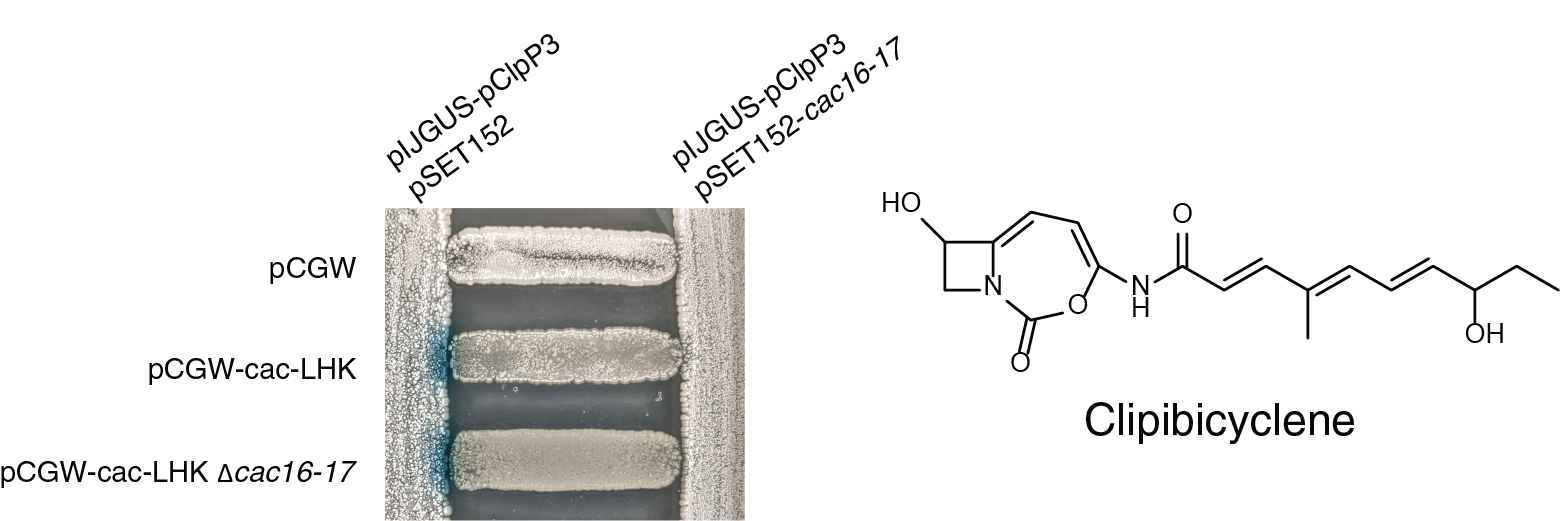

Altogether, within three months of first observing covalent inhibition of ClpP in crude media, we characterized the role of clipibicyclene resistant ClpPs in the BGC, the activity spectrum of clipibicyclene, and solved the crystal structure of clipibicyclene modified E. coli ClpP. Remarkably, unlike most other promiscuous protease inactivators, clipibicyclene is specific to ClpP inhibition up to 25 fold over the four other proteases we tested. Since ClpP is the sole protease encoded in the BGC, it makes sense that clipibicyclene displays selectivity for it. Even though ClpP is the only essential protease in Streptomyces, several other Ser proteases play important biological functions (e.g. Lon and Htr), and so it would be detrimental for the cell to produce a compound that inhibits them. It is interesting that the BGCs for other indiscriminate protease inhibitors produced by Streptomyces spp., including antipain and chymostatin, encode no known resistance mechanism (i.e. protease) other than export. This sets clipibicyclene/ClpP apart from these promiscuous inhibitors and supports that this scaffold has been shaped by evolution to specifically inhibit ClpP, likely as a competitive tool for bacterial warfare.
One of the most interesting things about clipibicyclene is that it belongs to a much larger family of BGCs, many but not all of which are associated with ClpP. These BGCs are exceptionally widespread and common across bacterial taxa, and so offers a unique model to study how and why some BGCs are more common than others. What are the principles driving its spread and diversification? How useful are these specialized metabolites for bacterial competition? How much cross-resistance is there among BGC family members? If not ClpP-associated, what bioactive molecules do the BGCs encode? I hope that clipibicyclene’s identification will pave the way to answer some of these questions.
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in