A multi-omics approach for biomarker discovery in neuroblastoma: a network-based framework
Published in Cancer and Protocols & Methods
Neuroblastoma (NB) is one of the leading causes of cancer-associated death in children. MYCN amplification is a prominent genetic marker for NB, and its targeting to halt NB progression is difficult to achieve. Therefore, an in-depth understanding of the molecular interactome of NB is needed to improve treatment outcomes. Analysis of NB multi-omics unravels valuable insight into the interplay between MYCN transcriptional and miRNA post-transcriptional modulation. Moreover, it aids in the identification of various miRNAs that participate in NB development and progression.
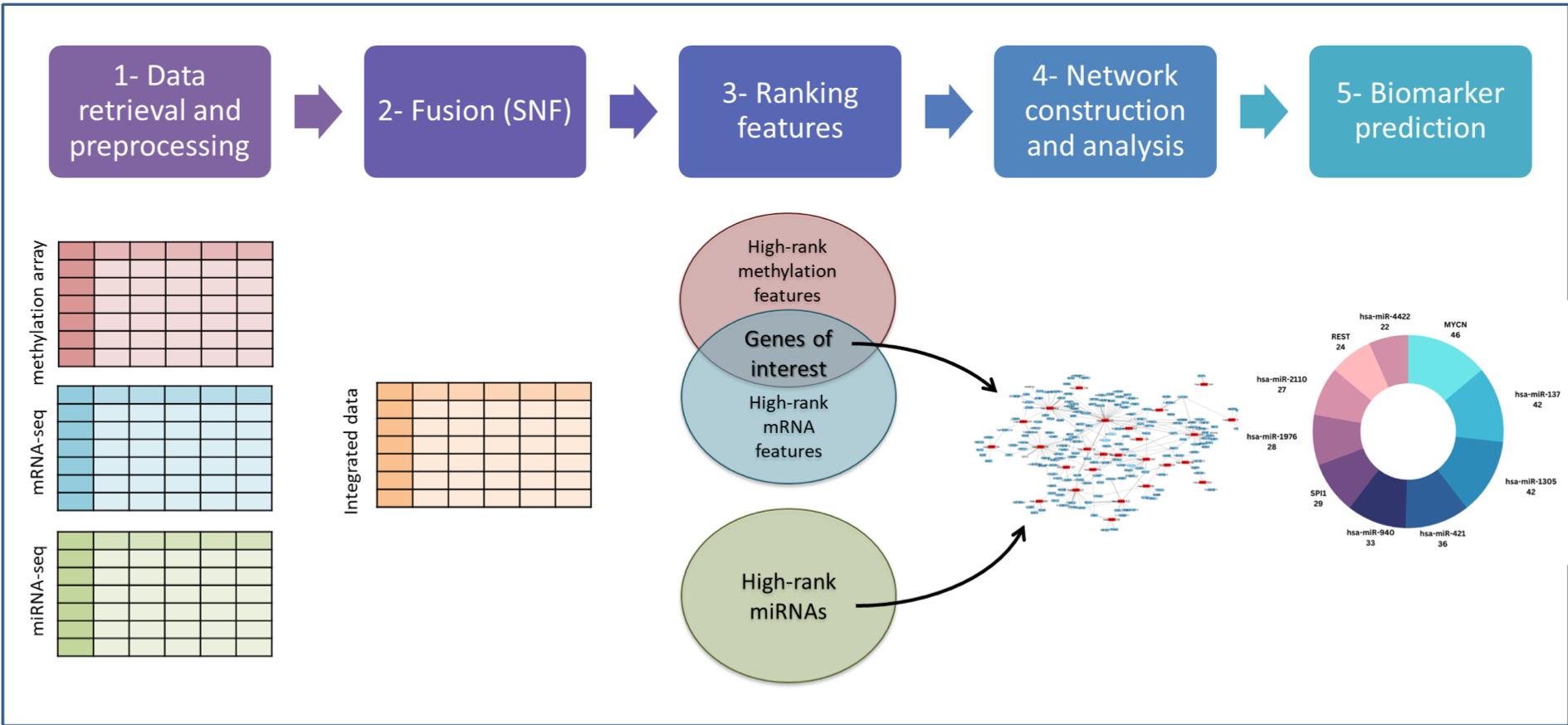
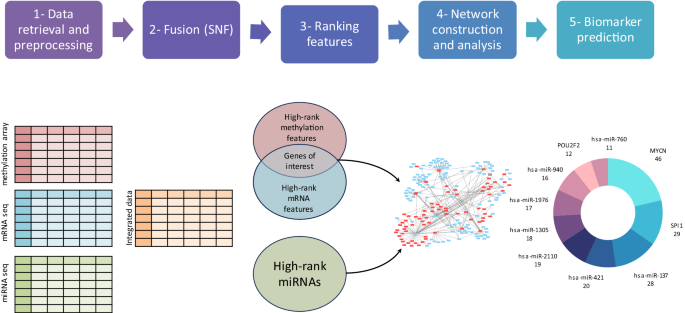
This study proposes an integrated computational framework with three levels of high-throughput NB data (mRNA-seq, miRNA-seq, and methylation array). Similarity Network Fusion (SNF) and ranked SNF methods were utilized to identify essential genes and miRNAs. The specified genes included both miRNA-target genes and transcription factors (TFs). The interactions between TFs and miRNAs and between miRNAs and their target genes were retrieved where a regulatory network was developed. Finally, an interaction network-based analysis was performed to identify candidate biomarkers. The candidate biomarkers were further analyzed for their potential use in prognosis and diagnosis.
The candidate biomarkers included three TFs and seven miRNAs. Among them, the roles of MYCN, hsa-miR-137, hsa-miR-421, and hsa-miR-2110 in NB tumor development and progression have been studied and proven. On the other hand, the rest of the predicted biomarkers in our study, such as SPI1, POU2F2, hsa-miR-1305, hsa-miR-1976, hsa-miR-940 and hsa-miR-760, could serve as potential biomarkers to halt NB tumorigenicity. In addition, their role in other tumor development and progression has been studied. Our regulatory network shows that they interact with some well-studied NB biomarkers, including MYCN, which support their under-studied implication in NB development.
In conclusion, analyzing cellular interactome to identify potential biomarkers is a promising approach that can contribute to optimizing efficient therapeutic regimens to target NB vulnerabilities. This study proposes a deeper understanding of MYCN interactome, providing candidate routes for targeted therapies in NB.
Follow the Topic
-
npj Systems Biology and Applications
An online Open Access journal dedicated to publishing the premier research that takes a systems-oriented approach and encourages studies that integrate, or aid the integration of, data, analyses and insight from molecules to organisms and broader systems.
Related Collections
With Collections, you can get published faster and increase your visibility.
Systems mechanobiology
Publishing Model: Open Access
Deadline: Jun 28, 2026
Systems immunology: multi-omics approaches, dynamical modeling and novel agentic AI approaches
Publishing Model: Open Access
Deadline: Sep 12, 2026

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in