A simple animal reveals fascinating insights in intracellular symbionts
Published in Microbiology
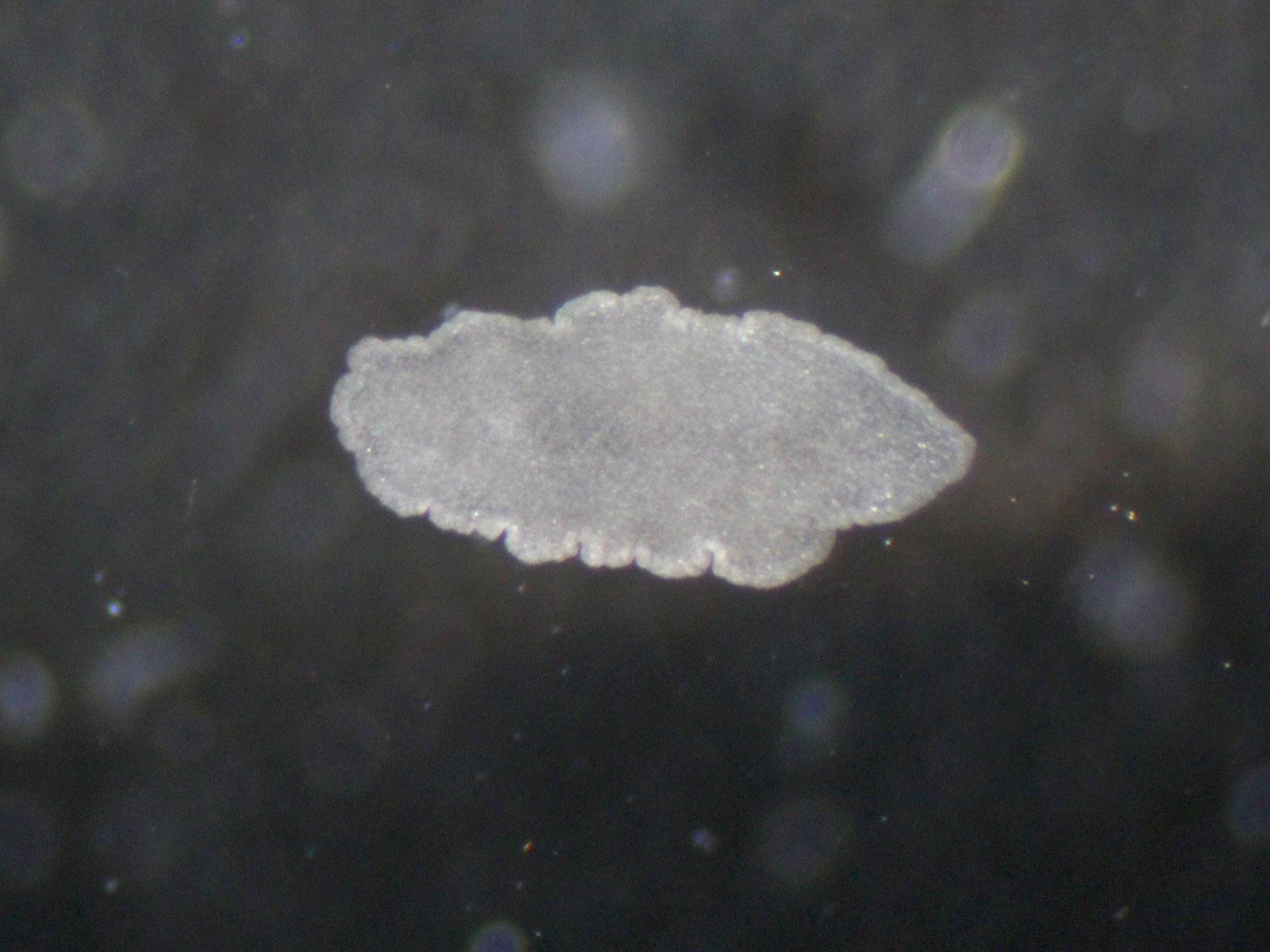

The story of our paper started many decades ago with the German zoologist Karl Grell. In his efforts to show that the small, transparent, mouth and gutless animals called Trichoplax belong to a new phylum – the Placozoa - and are not larvae of cnidarians, he studied them using electron microscopy. He observed, among many other morphological features, bacteria-like structures in “den Zisternen des rauhen endoplasmatischen Retikulums” – the cisternae of the rough endoplasmic reticulum (rER).
Fast forward many years, when the University of Hawaii zoologists Margaret McFall-Ngai and Mike Hadfield reached out to Harald Gruber-Vodicka and Nicole Dubilier, asking if they would be interested in taking a closer look at the symbiont. Mike has been avidly collecting placozoans from his seawater tanks, and together with Margaret observed bacteria in every individual they looked at. One of the first challenges was dealing with the cryptic diversity of host species in placozoans. We finally established a Trichoplax lineage from Hawaii that originated from a single animal. This made sure that we could connect the outcome of all the different methods and analyses we applied.

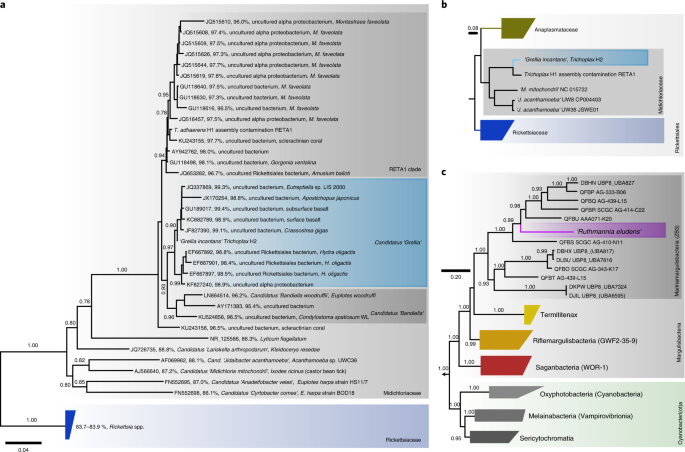
With the lineage established, we built on our expertise at working with minute animals, and performed metagenomic analyses of single individuals. The metagenomes revealed a stable community, but surprisingly it was not only one bacterium that we found, but two that were always present. This highlighted one of the most important take home lessons of this project: Only by going down to the single individual level and avoiding any form of sample pooling, we could reliably distinguish symbiotic associations from contaminations or infections of a small fraction of the population.
One of the symbionts – we named Ca. Grellia incantans - is a close relative to rickettsial parasites which are infamous for causing devastating diseases such as scrub fever or Rocky Mountain spotted fever. The other one – we called Ca. Ruthmannia incantas - is a member of the remote phylum Margulisbacteria, which was discovered only a few years ago. In fact, the phylum was only known from sequencing data and now with Trichoplax growing in the lab, we could not only image Margulisbacteria for the first time, but also have experimental access to them.
To disentangle this ménage à trois Nikolaus Leisch was critical with his fluorescence and electron microscopy expertise. The first big hurdle was the strong autofluorescence of the Trichoplax animals which made the go-to method of visual identification using Fluorescence in situ hybridization (FISH) highly challenging. The second “big” hurdle was that the cells of Trichoplax are minute compared to most other eukaryotic cells – 3-4 times smaller and in the size range of a typical bacterium. Imaging structures within those cells quickly became challenging as we approached the resolution limit of light microscopy. Correlating the phylogenetic identification of the symbionts with electron microscopy showed that each bacterium colonizes specific host cells. The third hurdle was figuring out if Grelllia really resides in the rER. Since the original observations made by Grell, research has shown that many closely related bacteria are capable of hiding within host cells by decorating their vacuoles with ribosomes and “pretending” to be part of the rough ER. This raised the question if Grellia similarly camouflages itself. Using serial section-based electron tomography, we were able to trace the outer membrane of the nucleus, which forms the rough ER membrane. Looking at the first reconstruction, which showed that Grellia truly lives in the ER, was one of the greatest “Heureka” moments in this project.
What was puzzling for us was that Grellia lives inside one of the most important organelles of the eukaryotic cell and the placozoans still grow rapidly. The association appears to have no detrimental effects and we can think of at least two explanations. Only a few Grellia cells are present in each cell. This means that they do not overrun the host cell, unlike their close relatives, which camouflage themselves as rER, reproduce rapidly within their hiding spot, deplete the cells energy storages and eventually cause the death of the host cell. Additionally, although Grellia has the genes to steal energy (ATP) from their host, our transcriptomics analyses revealed that they don't express these genes. All this indicates a moderated lifestyle.
One of the most surprising parts of this study came when we screened the SRA database for Grellia. Our sequence analysis of samples from all over the globe showed that Grellia can be found everywhere in all aquatic habitats. This discovery raised a multitude of new questions: Where do these Grellia live – in other hosts or are they free-living? Is this intra-ER lifestyle more common than we thought?
You can find the paper at http://dx.doi.org/10.1038/s41564-019-0475-9
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in