After the Paper | A new chapter in cereal domestication | Xie et al. first discovered the key gene related to sorghum glume coverage
Published in Bioengineering & Biotechnology
In a study published online in Nature Communications, Chinese scientists have identified a key gene involved in sorghum glume coverage during cereal domestication. This research is conducted by Qi Xie's team from the Institute of Genetics and Developmental Biology, Chinese Academy of Sciences.
Loss of glume coverage leads those seeds are detached from sticky glumes, is a symbolic event in crop threshing process and crop domestication. The hulled character carried by wild crop species can resist the invasion of animals and fungi and ensure their own reproduction. However, this hulled trait is greatly weakened during crop domestication due to its inconvenient for the manual or mechanized grain threshing, processing and breeding. So, the naked character has been selected by humans. Sorghum is one of the earliest cultivated gramineous crops. It has a rich phenotypic variation in glume coverage, among which the grain sorghum accessions are mostly naked. Scholars took glume coverage character as a typical index to distinguish sorghum subspecies eighty years ago, but the molecular basis of sorghum glume coverage is still remains unknown.
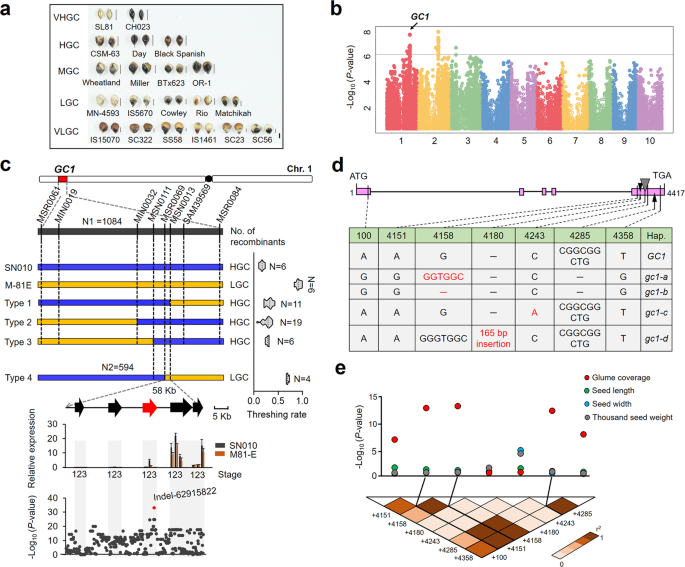
Through the application of genetics, bioinformatics and molecular biology, we demonstrate that natural variation of GC1 (glume coverage 1), a major locus controls glume coverage in sorghum. GC1 encodes an atypical G protein γ subunit. Once the candidate gene was cloned by combined map-based cloning and Genome-wide association study (GWAS), we detected sequence variations between the natural alleles. Four different types of natural variations occurred around the fifth exon of GC1, which lead to truncated mutations at the C-terminus of GC1, resulting in the reduced glume coverage. We then verified the gene function of GC1. Full-length GC1-overexpression plants show significantly decreased glume coverage while GC1-knockout mutants show greatly higher glume coverage, indicating that full-length GC1 negatively regulates glume coverage.
At the beginning, we were surprised that the phenotype of GC1-knockout, as is opposite to that of GC1-truncated version. By deep analysis we found the all of the truncated types harbor the intact Gγ-like domain, which may indicate that the GC1-truncated versions are also functioning in plants. Therefore, we overexpressed a truncated gc1 protein, and found that all of the T0 generations of gc1-overexpression plants showed significantly reduced glume coverage than both wild-type and full-length GC1-overexpression plants. These results show that the N-terminal G protein γ subunit is a key factor for negative regulation of glume coverage. We also overexpressed a C-terminus truncated homologous protein SiGC1124 in millet, and found glume coverage of the positive T0 transgenic plants were also significantly decreased. These results indicate G protein γ subunit functioning as a negative regulator is conserved in in cereals.
Since all natural mutations occur at the end of GC1, we continued to explore why the C-terminal truncated mutations reduce glume coverage in sorghum. Biochemical experimental results shows that these C-terminal truncated versions may reduce their degradation mediated by 26S proteasome, which accumulate significantly higher level of G protein γ subunit, and resulting in reducing glume coverage. Further results of cytology and transcriptome experimental results show that gc1-a, a natural truncated version, inhibits glume cell proliferation in the early stages of young panicles by downregulating the expression of Cyclin-CDK related genes, resulting in a low glume coverage.
In order to deeply reveal a molecular mechanism of GC1 involved in glume development, we performed an IP-MS assay to find a patatin-related phospholipase SbpPLAII-1 may play an important role as a key component of G protein signal. SbpPLAII-1 is a positive regulator that can promote glume cell proliferation process by upregulating the expression of Cyclin-CDK related genes. Furthermore, we found that compared with GC1, gc1-a, which is more stable in vivo, could promote the degradation of SbpPLAII-1, and then inhibit the function of SbpPLAII-1, resulting in the reduction of sorghum glume coverage (Fig. 3). In addition, selection analysis shows that GC1 gene was strongly artificially selected in naked sorghum cultivars. The genotype-based geographical distribution results show that the Sahel region, where the annual sorghum yield reaches 65% of total Africa, is a potential domestication center of sorghum.
In conclusion, our studies first identify a key gene GC1 controlling glume coverage, and deeply reveal a molecular mechanism of G protein signal involved in glume development in sorghum, which may provide gene resources for naked-grain breeding in sorghum, millet and other gramineous crops. In the future, site-specific truncated mutation in Gγ will also fast achieve the rapid de novo domestication of wild cereal germplasm resources.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in