Our paper describing the contribution of alterations of components of the ubiquitin-mediated proteolysis system (UPS) to tumorigenesis has just been published in Nature Cancer. We are proud to be authors of one of the first papers to this new journal, and we hope that it contributes to extend the reach of our work within the cancer research community.
For almost a decade now, our group has developed both methods and analyses to the discovery of genes and mutations involved in the development of tumors --i.e., driver genes and mutations. As we (and others) analyzed the first large pan-cancer cohorts of tumor whole exomes, some genes involved in core cellular functions, such as the maintenance of chromatin integrity, the splicing machinery, and the UPS were recurrently identified as cancer drivers1–4. Further studies clarified that alterations in these genes contributed to the misregulation of oncogenes or tumor suppressors and thus became positively selected in tumorigenesis5,6.
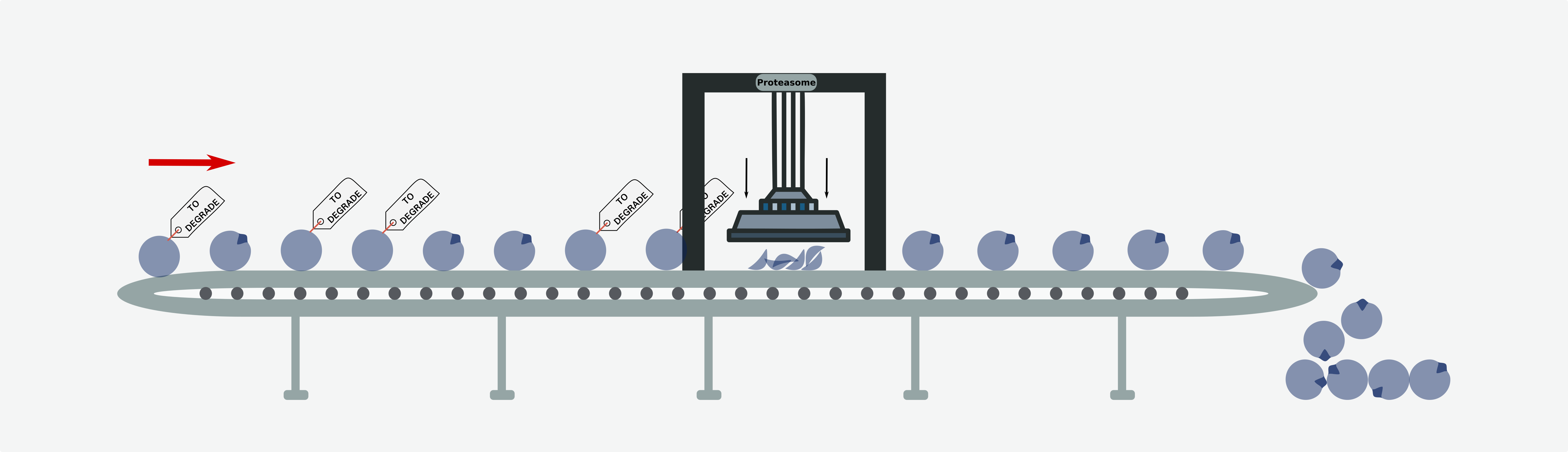
In homeostasis, the UPS contributes to the strict control of the abundance of proteins7,8. For instance the periodic variations in cell cycle regulators is achieved through this mechanism9. Specific regions of the protein structure (degrons) are recognized by an array of E3-ubiquitin ligases (E3), thus triggering their ubiquitination and subsequent degradation. Impairment of the recognition of a degron in a protein --via mutations of their sequence or that of its cognate E3-- may lead to its abnormal accumulation, and eventually tumorigenesis (in the case of oncoproteins). Our knowledge of degrons is so limited --around 200 instances are known for a couple dozen E3s-- that the total contribution of the alterations of UPS to tumorigenesis has remained obscure.
To uncover this contribution, we thus first systematically identified novel degron instances across the human proteome via machine learning, and we confirmed their validity using the mutations observed in primary tumors and cancer cell lines. The pattern of mutations in many of these novel degron instances show the signals of positive selection that denote their involvement in tumorigenesis. We demonstrated that overall, mutations disrupting E3s or degrons contribute represent more than one tenth of all driver mutations across more than thirty cancer types.
In our view, the relevance of this paper is not limited to the new insights into cancer biology. It has also produced an annotated list of potential novel degrons, which constitute a valuable resource to UPS researchers and protein engineers. It has introduced a novel framework for the analysis of the influence of any type of alterations (in cis or trans) on the stability of a given protein, as well as a novel score of the impact of mutations in degrons. It has also fostered the development of two new methods to identify amino acid sequence motifs under positive selection. All these resources are shared via the Extended Data Tables of the paper and a bitbucket repository (https://bitbucket.org/account/user/bbglab/projects/PD).
References
Follow the Topic
-
Nature Cancer
This journal aims to provide a unique forum through which the cancer community will learn about the latest, most significant cancer-related advances across the life, physical, applied and social sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Cancer Neuroscience: from mechanisms to therapy
Publishing Model: Hybrid
Deadline: Jan 30, 2027



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in