Amyloid-like p53 is a prognostic marker in serous ovarian cancer
Published in Cancer
A protein’s folding state is crucial for its native function. Under certain conditions, proteins can misfold and aggregate into amyloid fibrils leading to insoluble protein deposits. Neurodegenerative diseases, such as transmissible spongiform encephalopathy (e.g., mad cow disease or Creutzfeldt-Jakob disease), Alzheimer’s disease, or Parkinson’s disease, are characterized by the conformation change of a native protein into a cross-β-sheet enriched amyloid protein (1). Recently, the concept of amyloid-like protein aggregation has been extended to cancer, particularly to the tumor suppressor protein p53 (2). The TP53 gene is mutated in more than half of all cancers, with an exceptionally high mutation rate in ovarian cancer (OC). High-grade serous ovarian cancer, which accounts for more than 70% of all epithelial OC, carries a TP53 mutation in more than 95% of cases (3). OC cancer is characterized by poor 5-year survival of only 50% (4). Despite more than 40 years of research, there are currently no p53-targeting therapies approved (5).
In the last few years, numerous studies have investigated amyloid-like p53 and suggested that p53 aggregation plays a role in cancer development and progression, as well as chemotherapy resistance (6, 7). However, the clinical implications of amyloid-like p53 in serous OC remain largely unknown due to the lack of adequate methods for the reliable detection of p53 aggregates. Importantly, targeting amyloid-like p53 has become a growing field, and several drugs have been developed that show promising results (8, 9). Considering the poor prognosis of OC patients, new biomarkers, which help stratify patients and have the potential to guide treatment options or, ideally, be a drug target itself, are an utmost need.
Study objectives
We recently established the p53-Seprion-ELISA, which for the first time allows the quantitative detection of p53 aggregates (10). Compared to the current state-of-the-art method for p53 aggregation detection, the co-immunofluorescence assay, the ELISA has a shorter turnaround time and allows high throughput analysis (11). In this study, we aimed to elucidate the role of amyloid-like p53 in serous OC using the p53-Seprion-ELISA.
P53 aggregation is not associated with a specific mutation.
In this work, we analyzed fresh-frozen tissue specimens of 81 newly diagnosed OC patients using our p53-Seprion-ELISA to quantify the amount of p53 aggregates within the tumor. We detected amyloid-like p53 in more than half of all samples and in more than 80% of tumors bearing a single nucleotide substitution (missense mutation). The previous literature describes that specific missense mutations are so-called “aggregation-prone” mutations. Thus, we investigated this in our cohort and found that p53 aggregation is not associated with “aggregation-prone” mutations. Additionally, specific TP53 mutations led to the formation of amyloid-like p53 in some patients, while in other patients who carried the same mutation no aggregates could be detected. This finding led us to the conclusion that a TP53 mutation does not necessarily induce the formation of amyloid-like p53 and that co-factors should be considered in future studies.
To verify the p53-Seprion-ELISA, we established an additional assay for detecting and quantifying p53 aggregates, the so-called proximity ligation assay (PLA). This assay has the advantage of being applicable on FFPE tissue sections, which are the most common source of biological specimens in clinical routine, thereby enabling broader applicability of p53 aggregation diagnostics.
Amyloid-like p53 is an independent prognostic marker in serous ovarian cancer patients.
Further, we elucidated the prognostic impact of p53 aggregates in serous OC. We could detect a small proportion of patients with extensive amounts of p53 aggregates. This subgroup of patients was associated with prolonged progression-free survival in the initial univariate analysis. Next, we performed a multivariable analysis, adjusted for known prognostic factors, such as age, tumor stage, and residual tumor present after primary surgery. Thereby, we found that p53 aggregates are an independent prognostic marker in serous OC.
P53 aggregation is accompanied by increased apoptosis and immune response.
Since our findings were unexpected and in contradiction to the current literature, we aimed to explore the underlying reasons for the better survival of patients with high amounts of p53 aggregation. We found that these patients had increased cell death (apoptosis) levels in their tumors. Additionally, the analysis of corresponding plasma samples revealed that p53 aggregation is associated with elevated p53 autoantibody titers, indicating an enhanced immune response.
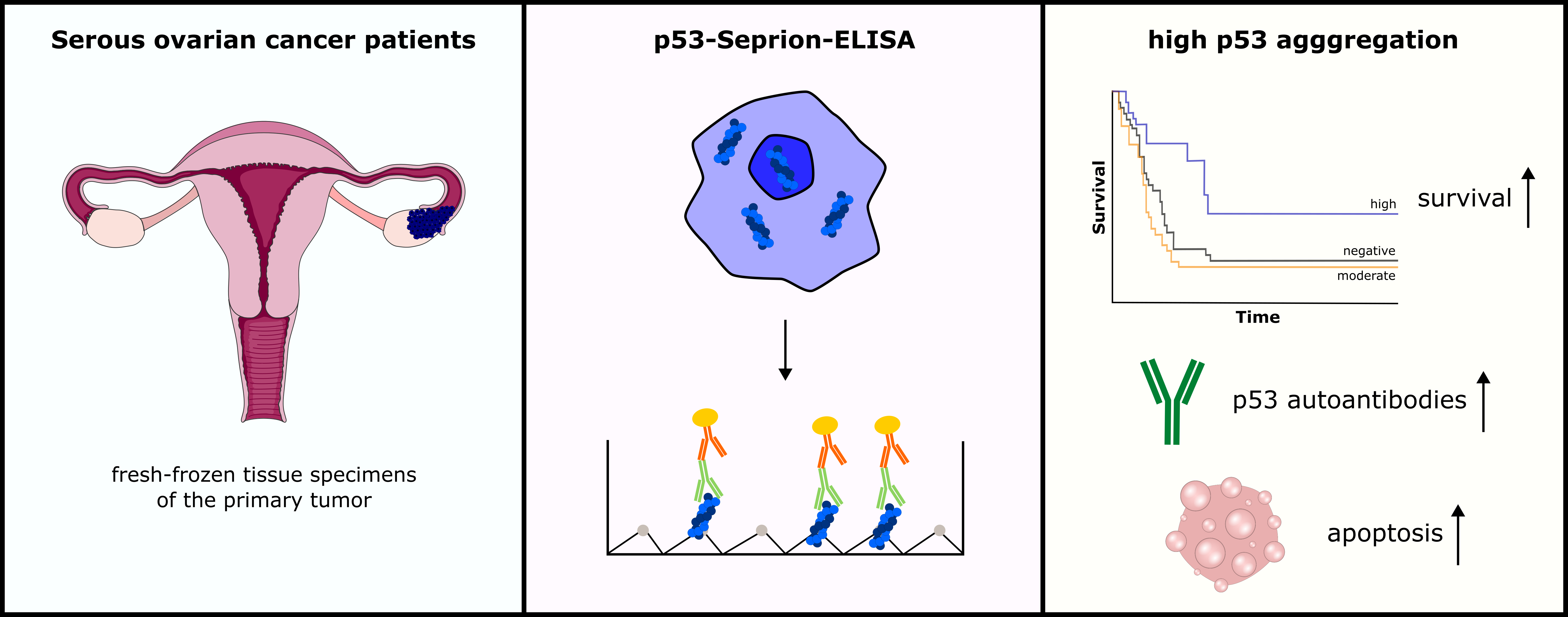
Conclusion and what is next:
To the best of our knowledge, this is the first study that applies a quantitative method to detect p53 aggregates and reveals that extensive p53 aggregation is associated with improved patient outcomes. These findings could pave the way for studies investigating p53 aggregation-targeting therapeutic strategies. Since p53 is mutated in half of all cancers, implementing the p53 aggregation status in future clinical trials could help stratify patients for personalized therapy options.
References
- Knowles TPJ, Vendruscolo M, Dobson CM. The amyloid state and its association with protein misfolding diseases. Nature reviews Molecular cell biology. 2014;15(6):384-96.
- Ano Bom AP, Rangel LP, Costa DC, de Oliveira GA, Sanches D, Braga CA, et al. Mutant p53 aggregates into prion-like amyloid oligomers and fibrils: implications for cancer. The Journal of biological chemistry. 2012;287(33):28152-62.
- Cancer Genome Atlas Research N. Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474(7353):609-15.
- Surveillance E, and End Results (SEER) Program Cancer Stat Facts: Ovarian Cancer 2023 [May 20, 2023]. Available from: https://seer.cancer.gov/statfacts/html/ovary.html.
- Wallis B, Bowman KR, Lu P, Lim CS. The Challenges and Prospects of p53-Based Therapies in Ovarian Cancer. Biomolecules. 2023;13(1).
- Yang-Hartwich Y, Soteras MG, Lin ZP, Holmberg J, Sumi N, Craveiro V, et al. p53 protein aggregation promotes platinum resistance in ovarian cancer. Oncogene. 2015;34(27):3605-16.
- Navalkar A, Paul A, Sakunthala A, Pandey S, Dey AK, Saha S, et al. Oncogenic gain of function due to p53 amyloids occurs through aberrant alteration of cell cycle and proliferation. Journal of cell science. 2022;135(15).
- Soragni A, Janzen DM, Johnson LM, Lindgren AG, Thai-Quynh Nguyen A, Tiourin E, et al. A Designed Inhibitor of p53 Aggregation Rescues p53 Tumor Suppression in Ovarian Carcinomas. Cancer cell. 2016;29(1):90-103.
- Palanikumar L, Karpauskaite L, Al-Sayegh M, Chehade I, Alam M, Hassan S, et al. Protein mimetic amyloid inhibitor potently abrogates cancer-associated mutant p53 aggregation and restores tumor suppressor function. Nature communications. 2021;12(1):3962.
- Maritschnegg E, Heinzl N, Wilson S, Deycmar S, Niebuhr M, Klameth L, et al. Polymer-Ligand-Based ELISA for Robust, High-Throughput, Quantitative Detection of p53 Aggregates. Analytical chemistry. 2018;90(22):13273-9.
- Heinzl N, Koziel K, Maritschnegg E, Berger A, Pechriggl E, Fiegl H, et al. A comparison of four technologies for detecting p53 aggregates in ovarian cancer. Frontiers in oncology. 2022;12.
Follow the Topic
-
Oncogene
This journal aims to make substantial advances in our knowledge of processes that contribute to cancer by publishing outstanding research.






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in