Bio-GO-SHIP: Generating high resolution surface ocean metagenomes through international collaboration
Published in Research Data

A research ship never sleeps. Scientists and crew members open bleary eyes at all hours, pour warm mugs of coffee or tea, and get to work. Countless hours are spent diligently watching as water is pumped through a vast array of equipment. Despite the stress and long work days, inspiration comes from taking a moment to admire the majesty of the open ocean. Trivia nights, dress up days, birthdays, and holiday celebrations break up the monotony of the journey and build a sense of camaraderie as you share a home and a common purpose. Oceanography is at its core a collaborative science, which is perhaps its greatest strength.
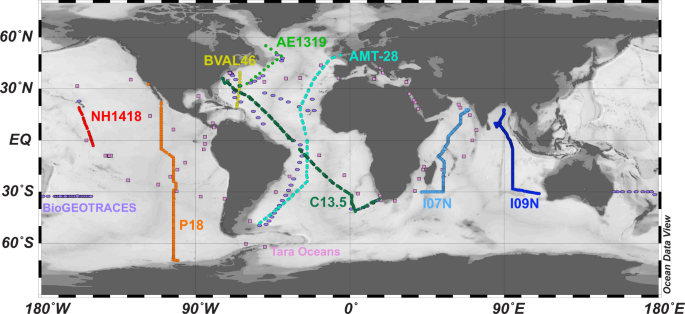
International collaboration is fundamental to the new Bio-GO-SHIP initiative. A total of 7 data collectors and an additional 5 lab members spent 228 days at sea and many months in the lab generating 971 ocean metagenomes from 932 locations across the globe, the highest spatial resolution characterization of the surface ocean microbiome to date (Larkin et al. 2021). Another ~125 scientists from countries across the globe contributed to 95 metadata variables associated with this dataset. None of which would be possible without ~200 crew members from 8 deployments and 6 research vessels as well as an indispensable array of coordinators at both academic and governmental institutions. Moreover, a veritable army of data curators have worked to ensure that this data is findable, accessible, interoperable, and reusable, or FAIR. Although navigating the complexities of international data licensing in pursuit of the FAIR doctrine can be a circuitous process, the result is data resources with the ability to transform our fundamental understanding of science.
The ocean represents 70% of our planet’s surface and, by volume, 99% of the earth’s living space. Thus, ocean observation and sampling remain a major scientific challenge. Significant progress in both spatial and temporal sampling of the ocean has been made through autonomous programs such as Argo, SOCCOM, and the new Global Ocean Biogeochemical Array (GO-BGC). However, ship-based campaigns offer a wider range of potential biogeochemical measurements, increased accuracy, and full water-column coverage. Thus, programs such as the Global Ocean Ship-based Hydrographic Investigations Program (GO-SHIP), which seeks to produce high spatial and vertical resolution measurements of key state variables on decadal transects, and the Plymouth Marine Lab’s annual Atlantic Meridional Transect (AMT) are critical for understanding both ocean biogeochemistry and climate change impacts on ocean environments. The high-resolution data produced by these ship-based programs helps reduce sample noise and error rates and allows for more accurate characterization of elemental flux, dynamic chemical balances, and overall ecosystem function. However, on a global scale, in situ measurements of marine biological parameters has lagged behind measurements of chemical and physical characteristics.
With this dataset, we sought to use the wealth of data stored in microbial genomes as a “biosensor” of ecosystem processes. Through their rapid generation times, microbial genomes integrate the effects of environmental change. Moreover, shifting microbial communities and genomic content may indicate changes in environmental conditions before similar shifts can be detected in ocean physics or chemistry. For example, our initial analysis of this dataset has demonstrated that the gene content of critical ocean microbes can be used as a quantitative and systematic indicator of Nitrogen, Phosphorus, and Iron limitation, even when these nutrients are below chemical detection limits in the surface ocean (Ustick et al. 2021).
This work builds upon the success of previous ocean metagenome projects such as the Global Ocean Survey, Tara Oceans, and bioGEOTRACES. With a median distance between samples of 26.5 km, our dataset is unique in its ability to link biological ‘omics measurements including microbial diversity and functional traits with ocean geochemistry. This work will allow for more detailed examinations of the relationship between microbial communities and parameters including nutrient flux, microbial production and respiration, carbon chemistry, and mesoscale dynamics such as eddy structure. At the same time, our dataset highlights significant gaps in our measurements of the global ocean metagenome. For example, high latitude environments, the Central and Western Pacific Ocean, and ecosystems below the euphotic zone have all been largely under-sampled when it comes to ‘omics measurements. Luckily, the only way to obtain these critical measurements is to pack up our equipment, our favorite caffeinated beverage, and take to the high seas.
Follow the Topic
-
Scientific Data
A peer-reviewed, open-access journal for descriptions of datasets, and research that advances the sharing and reuse of scientific data.
Related Collections
With Collections, you can get published faster and increase your visibility.
Genomics in freshwater and marine science
Publishing Model: Open Access
Deadline: Jul 23, 2026
Genomes of endangered species
Publishing Model: Open Access
Deadline: Jul 01, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in