Biosynthetic brilliance and blunders in virginiamycin M assembly
Published in Bioengineering & Biotechnology
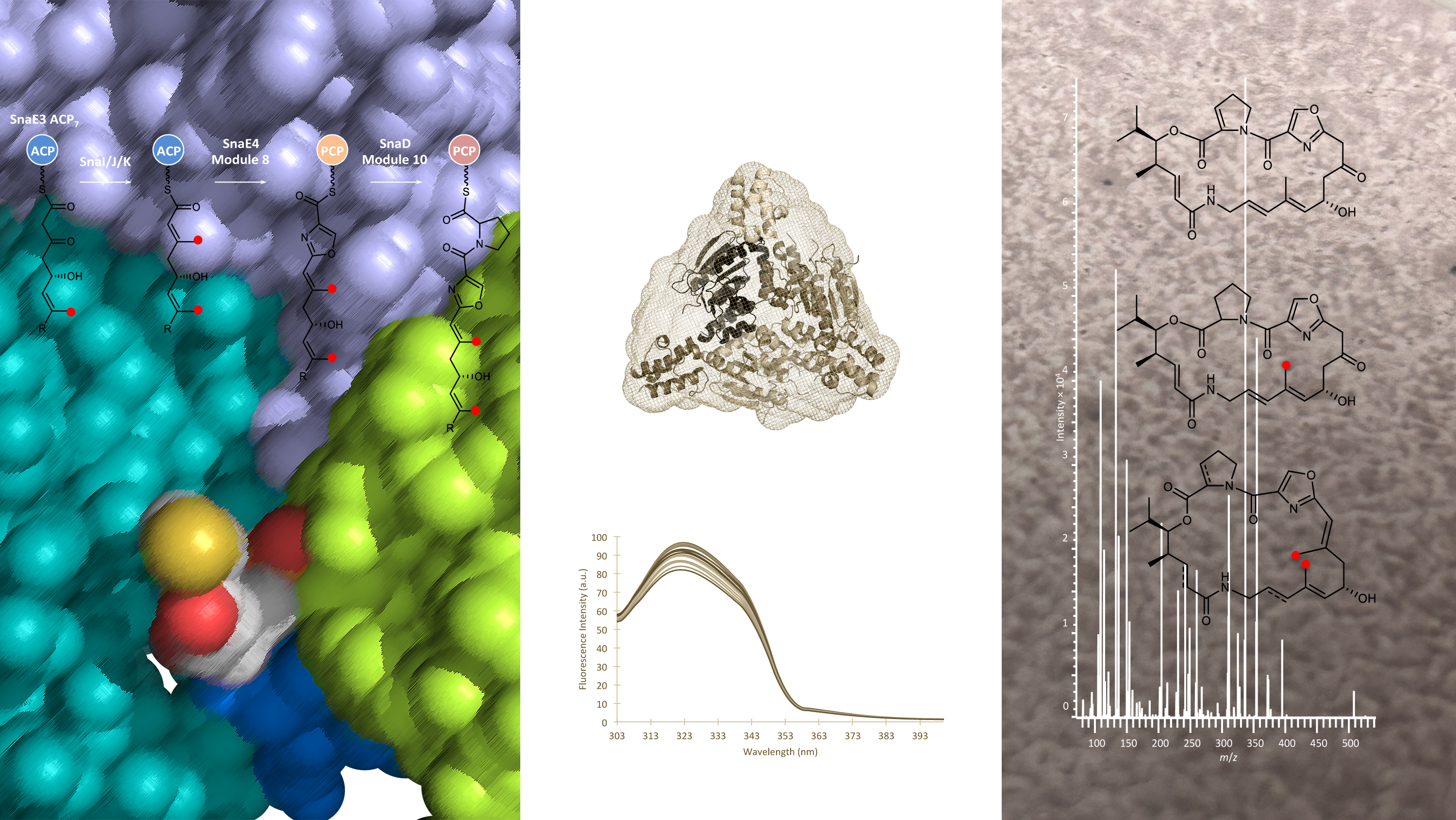
The polyketides are highly-complex small molecules made by multiple types of organisms including bacteria1. We are interested in this type of specialized metabolite because a large number of polyketides are deployed in the clinic, notably as front-line antibiotics and anti-cancer drugs2,3. A prominent example is the Streptomyces-derived virginiamycin M4, a modified (semi-synthetic) form of which is used in a combination therapy Synercid® to combat multi-resistant bacterial infections.
Another remarkable feature of polyketides is how they are constructed. The mega-enzymes which are responsible – the polyketide synthases (PKSs) – are among the biggest and most complex catalysts in all of Nature5. More astonishing still, the PKSs act like molecular-scale assembly lines, as each task in building the metabolites is performed by a dedicated region of the proteins called a ‘domain’1. Specifically, for every stage in polyketide construction, there is a domain which chooses the next building block, a domain which adds it to the growing chain, and a variable number of domains which chemically tinker with the functional groups to establish the final product. Throughout all of these steps, the polyketide remains attached to small, non-catalytic domains called acyl carrier proteins (ACPs). Both the catalytic and ACP domains are collected into functional units called ‘modules’, hence the name ‘modular PKSs’ for these types of systems.
Virginiamycin biosynthesis also involves a set of enzymes (called VirC, VirD and VirE) which are distinct from the PKS, and therefore act ‘in trans’ on the chains attached to the ACPs4. The job of this enzymatic ‘cassette’ is to install a β-methyl group into the polyketide chain at a specific point in the assembly process6. Virginiamycin is not alone is possessing a β-methyl, as many other polyketides incorporate β-methyls and other types of β-branching modifications7.
Given the high therapeutic value of polyketides, many labs world wide have been aiming for decades to generate improved analogues of these structures by genetic engineering of the PKS enzymes8. In this context, it would be particularly interesting to be able to introduce β-branching at will into polyketides, as these types of functionalities modulate both the biophysical and biological activities of the molecules7. However, the success of such efforts depends on gaining an intimate understanding of PKS structure and enzymology. In this work, we sought to decipher how β-branching occurs during virginiamycin M βassembly, and in particular, how the β-methylation cassette ‘choses’ at which point to intervene during the biosynthesis − information which is necessary to enable site-specific modification of other polyketides.
To answer this question, we succeeded in visualizing the interaction between VirD and its preferred ACP partner in atomic detail using X-ray crystallography. Analysis of the ACP−VirD structure revealed that the choice of substrate to modify depends heavily on the surface features of the ACP domain – i.e. the precise pattern of charged, polar and hydrophobic amino acids surrounding the attached polyketide chain – and not on differences in overall ACP architecture, as had been proposed9. ACPs are small domains, and therefore the number of surface residues available to drive these specific interactions is limited. Indeed, to our surprise, we discovered using binding tests in vitro, that the cassette enzymes recognize a second ACP domain in the same PKS. We further showed that this interaction results, in about 1 out of every 100 molecules, in a second β-methylation reaction, giving rise to a previously unidentified virginiamycin analogue. While this may not seem like much, the amounts of this derivative relative to the parental virginiamycin M are comparable to those typically achieved via PKS synthetic biology8.
It had been assumed previously that β-modification reactions occurring during polyketide assembly are highly programmed, and therefore faithful6,7. Our results show conclusively that PKSs sometimes make mistakes, and therefore that it would be worthwhile to re-analyze bacterial growth extracts for metabolites arising from other biosynthetic errors. It may even be possible to boost the yields of such compounds by deliberately disabling the control mechanisms which normally suppress their synthesis. Given the large number of PKS pathways which incorporate trans-acting enzymes10, this biosynthetic loophole may be widely exploitable.
For more detailed information on our work, please see the original article: “Decrypting the programming of β-methylation in virginiamycin M biosynthesis” in Nature Communications (https://doi.org/10.1038/s41467-023-36974-3).
References
- Weissman, K. J. 1.02 - Bacterial Type I Polyketide Synthases. in Comprehensive Natural Products III (eds. Liu, H.-W. (Ben) & Begley, T. P.) 4–46 (Elsevier, 2020). doi:10.1016/B978-0-12-409547-2.14644-X.
- Demain, A. L. Importance of microbial natural products and the need to revitalize their discovery. J. Ind. Microbiol. Biotechnol. 41, 185–201 (2014). doi: 10.1007/s10295-013-1325-z
- Demain, A. L. & Vaishnav, P. Natural products for cancer chemotherapy. Microb. Biotechnol. 4, 687–699 (2011). doi: 10.1111/j.1751-7915.2010.00221.x.
- Pulsawat, N., Kitani, S. & Nihira, T. Characterization of biosynthetic gene cluster for the production of virginiamycin M, a streptogramin type A antibiotic, in Streptomyces virginiae. Gene 393, 31–42 (2007). doi: 10.1016/j.gene.2006.12.035
- Grininger, M. Enzymology of assembly line synthesis by modular polyketide synthases. Nat. Chem. Biol. In press (2023) doi:10.1038/s41589-023-01277-7.
- Calderone, C. T. Isoprenoid-like alkylations in polyketide biosynthesis. Nat. Prod. Rep. 25, 845–853 (2008). doi: 10.1039/b807243d
- Walker, P. D., Weir, A. N. M., Willis, C. L. & Crump, M. P. Polyketide β-branching: diversity, mechanism and selectivity. Nat. Prod. Rep. 38, 723–756 (2021). doi: 10.1039/d0np00045k
- Weissman, K. J. Genetic engineering of modular PKSs: from combinatorial biosynthesis to synthetic biology. Nat. Prod. Rep. 33, 203–230 (2016). doi:10.1039/c5np00109a
- Haines, A. S. et al. A conserved motif flags acyl carrier proteins for β-branching in polyketide synthesis. Nat. Chem. Biol. 9, 685–692 (2013). doi: 10.1038/nchembio.1342
- Helfrich, E. J. N. & Piel, J. Biosynthesis of polyketides by trans-AT polyketide synthases. Nat. Prod. Rep. 33, 231–316 (2016). doi: 10.1039/c5np00125k
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Healthy Aging
Publishing Model: Open Access
Deadline: Dec 31, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in