C-type lectin receptor 2d forms homodimers and heterodimers with TLR2 to negatively regulate IRF5-mediated antifungal immunity
Published in Healthcare & Nursing, General & Internal Medicine, and Immunology
Prior research predominantly concentrated on elucidating the function of CLEC2D as a ligand, with particular emphasis on unraveling the impact of CD161-CLEC2D interaction in shaping tumor immunity and antiviral responses. Our series of investigations endeavors, focusing on CLEC2D, have uncovered an elusive facet of this receptor, heretofore overlooked - its prowess as a recognition receptor for exogenous carbohydrates in fungal infection. This remarkable revelation accentuates the pivotal role of CLEC2D in orchestrating the delicate balance required for effective regulation of fungal infections.
CLEC2D, a member of the C-type lectin receptor family, has merited limited attention in the realm of fungal infections, unlike its extensively studied counterparts such as Dectin-1 and Dectin-2, which have been identified to specifically recognize β-glucans and mannans of fungi, initiating immune responses. However, the lately captivating series of revelations has engendered a profound fascination regarding the enigmatic role of CLEC2D in the realm of fungal immunity.
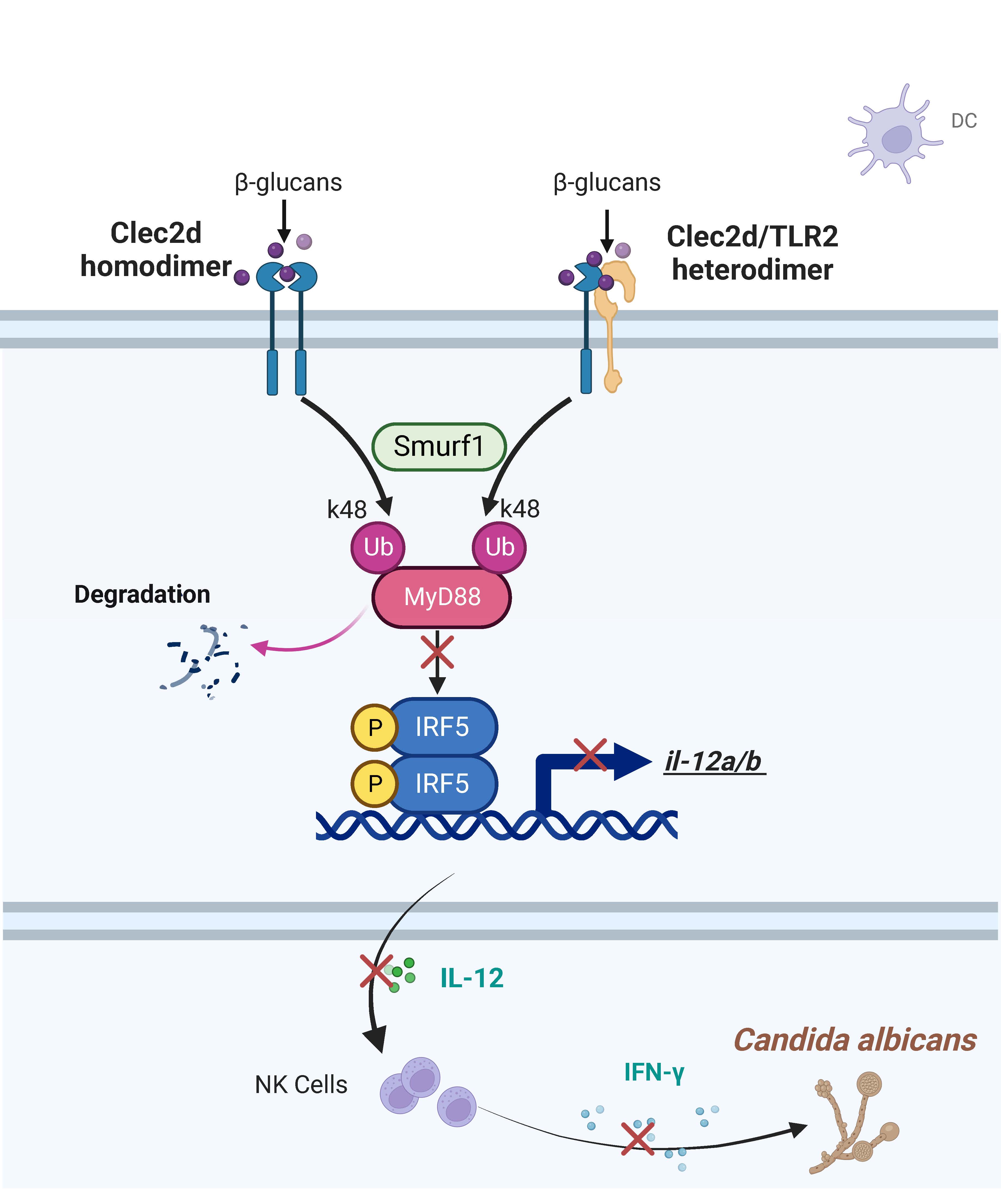
Previous studies have discerned CLEC2D 's affinity for binding high-molecular-weight sulfated glycoproteins, while our recent series of discoveries has elegantly unraveled the mysterious veil of CLEC2D as a receptor in recognizing fungal surface polysaccharides during fungal infections. We have found that in Cryptococcus neoformansinfection, CLEC2D recognizes glucuronoxylomannan to potentiate the immunosuppressive activity of neutrophilic myeloid-derived suppressor cells (MDSCs) by initiating p38-mediated production of the enzyme arginase-1, which inhibits T cell-mediated antifungal responses [1]. Here, in Candida albicans infection, CLEC2D recognizes and bind to β-glucan to induce ubiquitination and degradation of MyD88, inhibiting the activation of transcription factor IRF5 and subsequent IL-12 production. Notably, we have also elucidated that CLEC2D operates independently of CD161, serving as an inhibitory receptor in MDSCs and dendritic cells, thereby contributing to the negative regulation of antifungal immunity. These findings expand the previously perceived role of CLEC2D paired with CD161 as an immune checkpoint in NK cells.
It is not rare for the CLR family to exert immune functions through dimerization during fungal infections. Our previous findings have revealed that Dectin-3 and Dectin-2 form a heterodimeric pattern-recognition receptor (PRR) for host defense against fungal infection [2]. Interestingly, former research has found that the inhibitory receptor NKR-P1A on natural killer (NK) cells forms homodimers and bridges two CLEC2D molecules to form polymer complexes, which revealed a unique higher avidity binding organization by NKR-P1A that multimerizes CLEC2D, bolstering effective inhibitory signaling by an otherwise weak ligand-receptor affinity interaction [3]. In this study, we discovered that CLEC2D interacts across families with TLR2, forming a heterodimer during C. albicans infection. Remarkably, our findings revealed an intriguing pattern: in comparison to the TLR2 homodimers, CLEC2D exhibited an augmented capacity for binding β-glucan of C. albicans when forming heterodimers with TLR2. This increased binding strength translated into a more potent inhibitory effect, strikingly mirroring our earlier observations pertaining to the inhibitory significance of CLEC2D in C. neoformans infection, which also coincidentally exemplified a fascinating parallelism with those previous discoveries regarding the functional role of CLEC2D polymerization.
We found that the mechanism of CLEC2D dimerization-induced negative regulation of IRF5-mediated antifungal immunity, is driven by CLEC2D's role in promoting ubiquitination and degradation of MyD88. Of note, MyD88 is commonly regarded as a classical adaptor for TLR signaling. This underscores the intertwined and mutually reinforcing nature of CLR and TLR signaling pathways, collectively weaving a robust network within the PRR system to combat microbial infections. These findings provide fresh inspiration for investigating inter-family receptor interactions.
[1] Li YN, Wang ZW, Li F, Zhou LH, Jiang YS, Yu Y, Ma HH, Zhu LP, Qu JM, Jia XM. Inhibition of myeloid-derived suppressor cell arginase-1 production enhances T-cell-based immunotherapy against Cryptococcus neoformans infection. Nat Commun. 2022 Jul 14;13(1):4074. doi: 10.1038/s41467-022-31723-4. PMID: 35835754; PMCID: PMC9283461.
[2] Zhu LL, Zhao XQ, Jiang C, You Y, Chen XP, Jiang YY, Jia XM, Lin X. C-type lectin receptors Dectin-3 and Dectin-2 form a heterodimeric pattern-recognition receptor for host defense against fungal infection. Immunity. 2013 Aug 22;39(2):324-34. doi: 10.1016/j.immuni.2013.05.017. Epub 2013 Aug 1. PMID: 23911656.
[3] Bláha J, Skálová T, Kalousková B, Skořepa O, Cmunt D, Grobárová V, Pazicky S, Poláchová E, Abreu C, Stránský J, Kovaľ T, Dušková J, Zhao Y, Harlos K, Hašek J, Dohnálek J, Vaněk O. Structure of the human NK cell NKR-P1:LLT1 receptor:ligand complex reveals clustering in the immune synapse. Nat Commun. 2022 Aug 26;13(1):5022. doi: 10.1038/s41467-022-32577-6. PMID: 36028489; PMCID: PMC9418145.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in