Citizen-science to chart the oral microbiome
Published in Microbiology
World of mouth
We are not alone. every square inch of our internal and external body surface is populated by a rich and diverse microbial community. Our mouth is not an exception. In fact it is estimated that each milliliter of saliva may contain about 100 million microbial cells, and over 700 bacterial genera have been described in this niche. Identifying the composition of the oral microbiome in adolescents and how it may change with varying dietary and hygiene habits was the main research purpose of the “Stick out your tongue” project (see website). A particularity of this project is that we took a citizen-science approach, aiming to involve citizens in all steps of the research, from hypothesis generation, through sample collection, to data analysis and interpretation. This approach has enabled us to access a large number of samples and data from a so-far poorly studied population (healthy adolescents) but has also represented many challenges. The scientific results have been published recently (see our publication). I here provide a personal account of how this project went ahead,
It all started by flexing the muscles of our institute core facilities. This was not a project to be ran by a research group but rather an institutional effort that would showcase our Institute’s capacity to undertake challenging projects. Different core facilities such as the sequencing and the bioinformatics units were in charge of processing and analyzing all the samples up to a point that my research group – with expertise in microbial genomics- could perform more in depth analysis and interpretation. The problem number one was that the samples were to be collected in a two-months frame and a large number of samples would have to be analyzed in a record time. The problem number two was that this type of analyses had never been done by these core facilities before, and the project was starting in a few months. After a very stressing period of testing various protocols of sample collection, extraction, amplification, analyses, etc, we had a working protocol, tested over and over on samples taking from many kind volunteers at the CRG.
Scientists on the road
In
parallel we had to prepare the actual logistics of sample collection.
Accessing samples from high-schools all over Spain was a nice idea.
But how to get there within a reasonable time period? Yes, the idea
of a van carrying whatever was necessary to collect the samples and
traveling throughout the Spanish geography came to our mind. Although
we initially thought about this as a crazy idea, it ended up being
the most cost-effective and reasonable one. If the van would be
operated by someone able to drive, communicate science, and perform
the initial sample processing, this would allow us to collect about
2000 samples from around 40 different locations in Spain in around
eight weeks. But who would take this job? Who would be willing to
live on the road for two months, sleep on a van next to a portable
centrifuge and a freezer, and be ready to engage with teenagers from
all around the country, almost every day? Fortunately we knew such
person: Luis Bejarano, one of the co-authors of the study and perhaps
the one providing the most key contribution. The sample collection
logistics involved coordinating with dozens of high-schools with the
valuable help of dozens of science or biology teachers who
volunteered to participate in the study. In addition, the
communication department put much effort in preparing communication
material and making the arrival of our van being noticed by the local
media in every one of our stops. I myself had the change to
participate in several of the sample collection events and discuss
directly with participants. The two months of the sample collection
phase were of the most intensive of the project, with problems
arising almost every other day. But with Luis on the road and Annick
Labeeuw at the control center in Barcelona we successfully overcame
every logistic challenge.
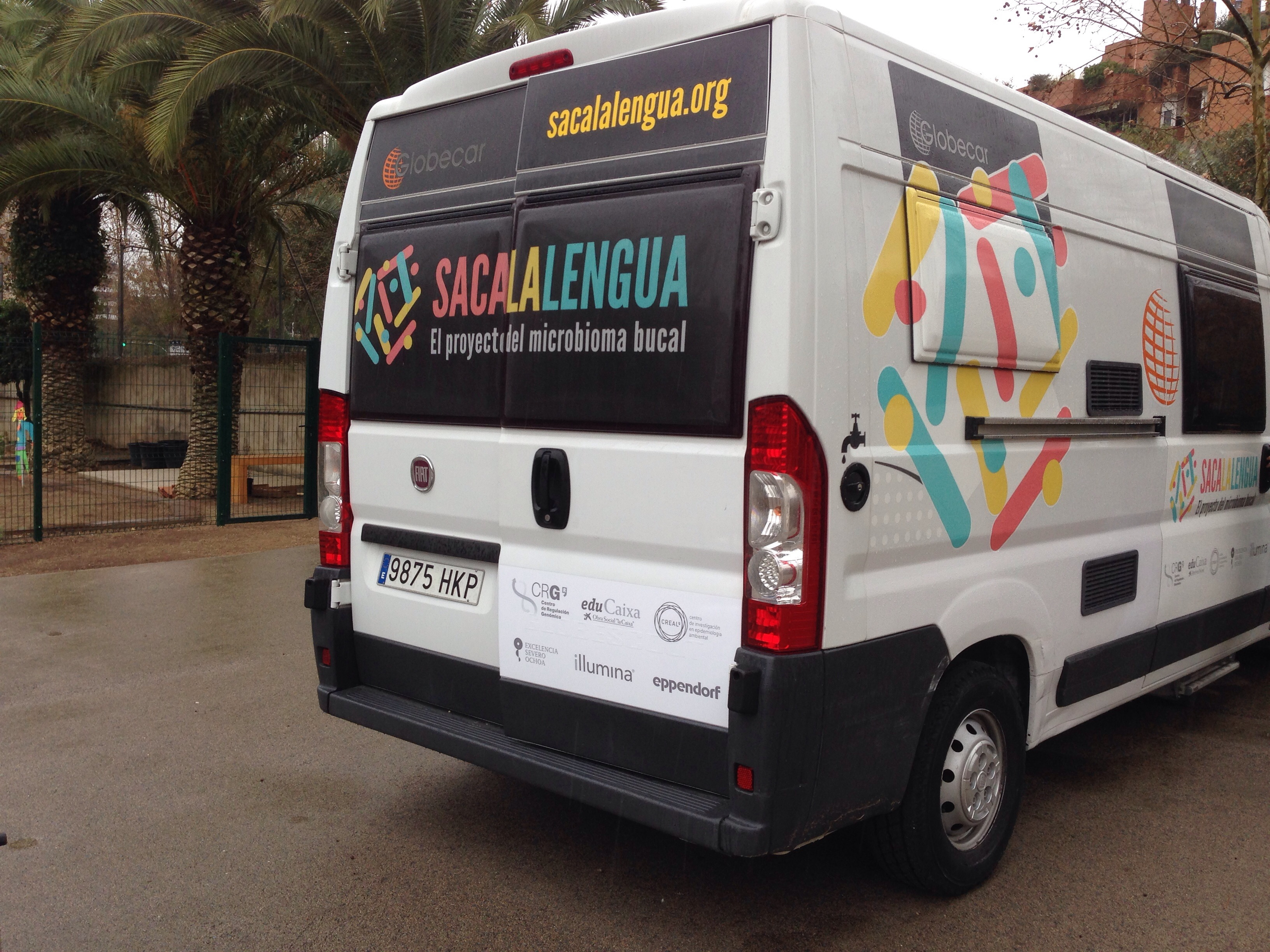

Our fully-equipped bus
Engaging adolescents
Once the samples arrived to Barcelona the problems were not over, but these were mostly within the terrain in which we scientists are more trained (technical problems with sequencing infrastructure, delivery of some reagents, logistics of scalability, etc). Additionally we had the challenge of involving study participants over the whole project. Citizens had already contributed by proposing questions to investigate by adding them to a questionnaire filled by all study participants. Many of these questions reflected questions similar to those initially proposed by the research team (e.g. effect of diet) but others we would have never considered on our own (i.e. does having a piercing in the mouth affect oral microbiota?). In addition, study participants were definitely involved in sample and data collection. However, we needed to have a way to have them involved in the analysis and interpretation part of the study. This represented a real challenge as the analyses are rather sophisticated and needed particular skills. We approached this by having a workshop with teachers of the participating schools were we discussed at length on the project objectives and how to get students involved in analyzing the projects data. We came out with several ideas that consisted in a series of challenges with the real data were the students could participate. Participants could classify sequencing reads to a given taxonomic level by using sequence comparison tools and compare their classification with our automated pipeline, teams could test their own hypothesis about any relationships between any of the variables measured within the project and, finally, teams could propose new original ways to represent the vast amount of data.

Teachers thinking on how to engage students
Stomatotypes, tap water, and oral microbiome
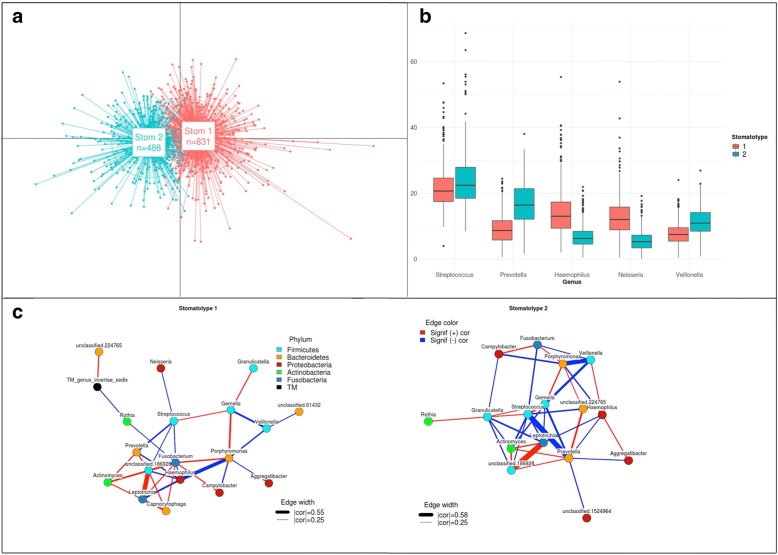
In parallel, the team of scientists involved in the project crunched the data in search for interesting new correlations. The dataset was impressive with almost 2000 processed samples and a rich set of meta-data from the questionnaire. For several reasons, the project used a meta-barcondig approach (amplifying and sequencing a region of the 16S gene) rather than a genome shotgun approach, and thus our data basically consisted on relative proportions of microbial taxa within each analyzed samples. We found many correlations, some of which had already been described. One of the main preliminary hypothesis for this project was to test whether samples from rural areas would be different from those of urban areas, and the sampling sites were selected accordingly. However, we did not find any significant differences between these two environments. We found however some regional differences in several of the bacterial taxa. When plotting the abundance of some of these taxa on a map, these suggested to us that regional differences may be playing a role. Among other potential factors, the composition of drinking water was an obvious candidate. Hydrography of Spain is quite peculiar with strong differences in the mineral composition of tap water among regions. In addition, the oral microbiome is obviously exposed to tap water, which we use for drinking, cooking and our hygiene. We crossed our data with those from other studies of the tap water composition of the same regions and found that several water parameters and bacterial taxa were significantly correlated. Although some components of water such as fluoride have been related to the prevalence oral diseases such as caries, to our knowledge, no study so far had reported correlations between the drinking water and the human oral microbiome composition. I consider this to be one of our most significant findings. Many other correlations with dietary or hygiene habits are reported, as well as a general structuring of the microbiome variability into two major types (which we called stomatotypes). Importantly the dominant compositional differences between these two types are present in other studies that performed similar clustering approaches of their data in very different populations, suggesting these two stomatotypes may represent two of the possible broad types of microbial ecosystems that can be present in our mouth.
Our results comprise an interesting set of hypothesis that need to be further tested with more direct and controlled approaches. Citizen-science has enabled us to access a type and a scale of data that would have been difficult through an standard approach. However, it also has come with its own challenges. For instance some peers may cast doubts on sample collection or experiments done out of the controlled space of a laboratory, far from expensive devises and performed by highly motivated but inexperienced citizens. In addition, it takes us scientists out of our comfort zone, forcing us to face unexpected challenges and making the whole process of discovery a bit more cumbersome. Yet, I think it is one of the bests ways to engage with the public. Citizens that to some degree live a research project from within will be more prepared to understand the intricacies of science and will be more prepared to critically assess information from scientific discoveries and understand their implications.
Follow the Topic
-
Microbiome
This journal hopes to integrate researchers with common scientific objectives across a broad cross-section of sub-disciplines within microbial ecology. It covers studies of microbiomes colonizing humans, animals, plants or the environment, both built and natural or manipulated, as in agriculture.
Related Collections
With Collections, you can get published faster and increase your visibility.
Animal Gut Nutrition and Greenhouse Gas Mitigation
Animal Microbiome, Journal of Animal Science and Biotechnology and Microbiome call for submissions to the collection on Animal Gut Nutrition and Greenhouse Gas Mitigation.
Efforts to reduce greenhouse gas emissions from livestock systems increasingly hinge on innovations in animal gut nutrition. The dynamic relationship between the gut microbiome and nutrient utilization plays a pivotal role in shaping methane output, feed efficiency, and overall sustainability. Advances in microbial ecology—particularly in understanding the role of gut microbiome in nutrient metabolism—are opening new pathways for mitigating emissions while enhancing productivity. These developments support the implementation of climate-smart agricultural strategies to address climate change and its impacts.
Looking ahead, continued research in this field has the potential to yield innovative solutions such as targeted probiotic supplementation, which could further optimize gut function and enhance nutrient absorption. These advancements may lead to reduced greenhouse gas emissions while improving animal health and productivity. By deepening our understanding of the animal gut microbiome, we can contribute significantly to sustainable agricultural practices that benefit both the environment and food security.
We invite researchers to contribute to this special Collection on Animal Gut Nutrition and Greenhouse Gas Mitigation. Topics of interest include but are not limited to:
- Animal Gut Microbiome and Feed Efficiency
- Greenhouse Gas Mitigation Strategies
- Rumen Fermentation Dynamics
- Nutrient Utilization in Livestock
- Probiotic Supplementation Effects
- Sustainable Livestock Production Practices
- Climate-Smart Agriculture Innovations
This Collection supports and amplifies research related to SDG 13, Climate action.
All submissions in this collection undergo the relevant journal’s standard peer review process. Similarly, all manuscripts authored by a Guest Editor(s) will be handled by the Editor-in-Chief of the relevant journal. As an open access publication, participating journals levy an article processing fee (Animal Microbiome fees, Journal of Animal Science and Biotechnology fees, Microbiome fees). We recognize that many key stakeholders may not have access to such resources and are committed to supporting participation in this issue wherever resources are a barrier. For more information about what support may be available, please visit OA funding and support, or email OAfundingpolicy@springernature.com or the Editor-in-Chief of the journal where the article is being submitted.
Publishing Model: Open Access
Deadline: Sep 04, 2026
The Apple Microbiome
Microbiome and Environmental Microbiome are calling for submissions to our Collection on the Apple Microbiome.
With world apple production estimated at 84 million tons, the microbiome of the apple has significant implications for agriculture, food security, and human health. Understanding the complex interactions between apple plants and their associated microbial communities can lead to improved crop management strategies, enhanced fruit quality and longevity, and sustainable agricultural practices. Recent advances have highlighted the role of specific bacteria and fungi in promoting plant health and resilience against specific pathogens. Moreover, detailed profiling of these microbial communities, revealing their diversity and functional potential facilitate exciting future developments, such as the identification of beneficial microbial consortia for biocontrol and the formulation of tailored probiotic treatments for both plants and humans. By advancing our collective understanding in this area, we can work towards a more sustainable and resilient agricultural system.
Topics of interest include but are not limited to:
-Microbial diversity and function associated with apples
-Effects of soil health and rhizosphere interactions on apple production
-Impact of climate change on the apple microbiome
-Role of the apple microbiome in fruit quality
-Microbiome-driven strategies for disease resistance
This collection is open for submissions from all authors on the condition that the manuscript falls within both the scope of the collection and the journal it is submitted to.
All submissions in this collection undergo the relevant journal’s standard peer review process. Similarly, all manuscripts authored by a Guest Editor(s) will be handled by the Editor-in-Chief of the relevant journal. As an open access publication, participating journals levy an article processing fee (Microbiome, Environmental Microbiome). We recognize that many key stakeholders may not have access to such resources and are committed to supporting participation in this issue wherever resources are a barrier. For more information about what support may be available, please visit OA funding and support, or email OAfundingpolicy@springernature.com or the Editor-in-Chief of the journal where the article is being submitted.
Publishing Model: Open Access
Deadline: Aug 05, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in