Common Conditions of use Elements – Deceptively simple use conditions.
Published in Social Sciences, Computational Sciences, and Biomedical Research
Written by Spencer Gibson, Maria Sánchez, Pim Kamerling and Esther van Enckevort.
The initial concept for the Common Conditions of use Elements (CCEs) stemmed from an idea of two of its’ authors (Anthony Brookes and Esther van Enckevort). This was the simplification of use conditions down into their core concepts, to which rules could be appended specifying whether a use was permitted or not.
The vision was that a collection of such terms and rules could be used to summarise use conditions, without being constrained to predetermined permissions, as is the case with alternative approaches, which can compound the use type and permission (i.e. DUO 0000014: for research use only). This is especially the case when considering uses outside of the non-profit or academic arena. For instance, the DUO term DUO_0000018: not for profit, non-commercial use compounds “not for profit” and “non-commercial use”, with no complementary permissive statement(s). While such ontologies can define types of use extremely precisely, this process requires careful encoding by experts who are familiar with a given ontology.
With a focus on producing an elemental and easily understood set of terms, a core group of four people (Maria Sánchez, Mariapia Iermito, Pim Kamerling, and Spencer Gibson) from biobanks, registries and data governance was formed. We started the process by identifying “core concepts” for bioresources use. Each term was to conform to four key principles, in that they should be: elemental, neutral, relevant, and generic. The review process looked at various kinds of consents, material and data transfer agreements, as well as our own working processes to see what the elemental conditions of use were. The challenge was knowing where to draw the line, as the goal was a framework that would unify different worlds that may need to summarise use conditions for bioresources from across the field of rare disease research. The input from the wider group of researchers in the European Joint Program on Rare Disease (EJP-RD) was key in redirecting the core group when a more complex concept was encountered. Also, liaison with the group working on the Digital Use Conditions (Jeason. F. et al. 2024. https://www.nature.com/articles/s41597-024-03280-6) helped the reciprocal and iterative development of both.
Once the initial process of how to extract the core terms had been established the process of producing the initial set of core terms seemed to just flow. As a result, agreement on the first set of CCEs, thought to be potentially applicable to biobanks and registries or data platforms, took a year and a half. However, this was just a list a small group of researchers had devised amongst themselves and others had already tried similar approaches. The real question was, would it work in the real world?
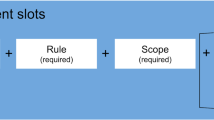
This required the input from those “in the field”. A group of “testers” (from within European biobanking and ERN communities), tested and provided feedback on the initial terms, which helped refine them into a more relevant and useful set. Finally, we had the product: 20 terms representing the most relevant, atomic, non-directional conditions of use for a bioresource. These terms could in conjunction with rules, such as those in the DUC structure, along with a short and simple piece of explanatory free text (where needed), represent the resource's policies for data and sample sharing as well as discovery.
Our objective was that the resultant framework would be both human and machine readable to facilitate the creation of easily interpreted “profiles” for a given bioresource. The profiles should be a simple summary of the main aspects of a bioresource’s use conditions (akin to a nutritional label), allowing a researcher to have a sense of whether their use was likely to be approved before undertaking the more formal access request. Not only would it potentially weed out uses that would be “show stoppers” but it could also serve to make the bioresources in the project more Findable, Accessible, Interoperable and Re-usable (F.A.I.R.) (Wilkinson M.D., et al 2016 https://doi.org/10.0.4.14/sdata.2016.18), by making their use conditions easily accessible. Indeed, the “showstoppers” was one of the use cases for DUC/CCE profiles that an independent group of testers highlighted in their initial training session on how to devise profiles using the DUC/CCE profile generation tool (https://ducejprd.le.ac.uk), prior to its’ potential adoption in the European Project on Neurodegenerative Disease (https://epnd.org). The testers felt that this could reduce the burden on access committees by preventing access request being made for uses that were clearly not permitted.
The group felt that Scientific Data was the most appropriate journal to open this standard up to the world to help with the governance of data and sample sharing, given its’ interest in the reuse of data. While the DUC framework and CCEs are independent, as the saying goes “the whole is greater than the sum of its’ parts” and so the articles for both were written in parallel with the aim of presenting them side by side due to their complementary nature.
While the current format relies on a free text element to clarify the basic rule, we hope that this could be computer interpretable in future, either by employing an appropriately trained large language model or more simply by applying some basic restrictions on the free text such as: not introducing exemptions, duality or dependencies, so that the main term and rule meaning are not contradicted. Initial trials have given positive results using ChatGPT and locally run LLM models.
While the current system has been developed within the context of rare diseases, there is no reason why the general design principals for CCE terms and their implementation within the DUC schema could not be applied to other areas. The authors hope that the main take away that readers will get from both papers, is the process for defining and using CCEs within the DUC structure. We hope they will be inspired to create their own CCE terms for their specific use case. However, the adoption of any system relies on its’ accessibility for the end user. With this in mind, the authors would caution against defining terms that use language outside of that which is commonly used within the target user base.
Follow the Topic
-
Scientific Data
A peer-reviewed, open-access journal for descriptions of datasets, and research that advances the sharing and reuse of scientific data.
Related Collections
With Collections, you can get published faster and increase your visibility.
Genomics in freshwater and marine science
Publishing Model: Open Access
Deadline: Jul 23, 2026
Genomes of endangered species
Publishing Model: Open Access
Deadline: Jul 01, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in