Coupling T cell differentiation to the lung cancer mutanome via neoantigen surveillance
Published in Cancer

The contribution of neoantigen reactive T cells to anti-cancer immunity has become increasingly well-established over the last decade1. A key role for neoantigen-driven tumor recognition in humans is supported by the observations that i) patients with greater non-synonymous mutational burden (TMB) exhibit favourable clinical response to checkpoint blockade2,3, ii) neoepitope-specific T cells can be detected in clinical samples2–5, iii) adoptive T cell therapies that elicit radiographic responses contain neoantigen reactive T cell clones6 and iv) footprints of immune escape are enriched in tumors with an elevated neoepitope load7,8. However, the mechanisms underpinning neoantigen-specific T cell responses remain poorly characterized. Understanding how the neoantigen-driven T cell response is orchestrated at the cellular and molecular level may help to optimise cellular therapies, identify novel immunotherapy targets and define clinically relevant biomarkers.
In 2016 Prof Sergio Quezada, Prof Charles Swanton and I set out to decipher the T cell response to non-small cell lung cancer (NSCLC) in the context of the ‘Tracking Cancer Evolution through Therapy’ (TRACERx) study9. For the previous decade I had been fascinated by T cell exhaustion and terminal differentiation; two hypofunctional states induced by chronic or repetitive antigen engagement that occurred in an antigen-load-dependent manner, best characterised in viral infection10,11. More recently, similar trajectories of progressive T cell dysfunction were elegantly shown to be a feature of neoantigen reactive T cells in mouse models of cancer12. I reasoned that T cell differentiation could be leveraged as a readout for ongoing or prior (neo)antigen encounter, and more specifically, that if neoantigens fuelled a program of intra-tumoral T cell differentiation in NSCLC, this would manifest as a TMB-dependent skewing of progenitor to dysfunctional T cell subsets. Identifying T cell populations that were redistributed as a function of TMB would provide valuable insight into the cellular mechanisms of neoantigen driven responses and guide future experiments. The first step in testing this hypothesis was to integrate high dimensional cytometry with paired WES data in surgical resections from untreated NSCLC patients.
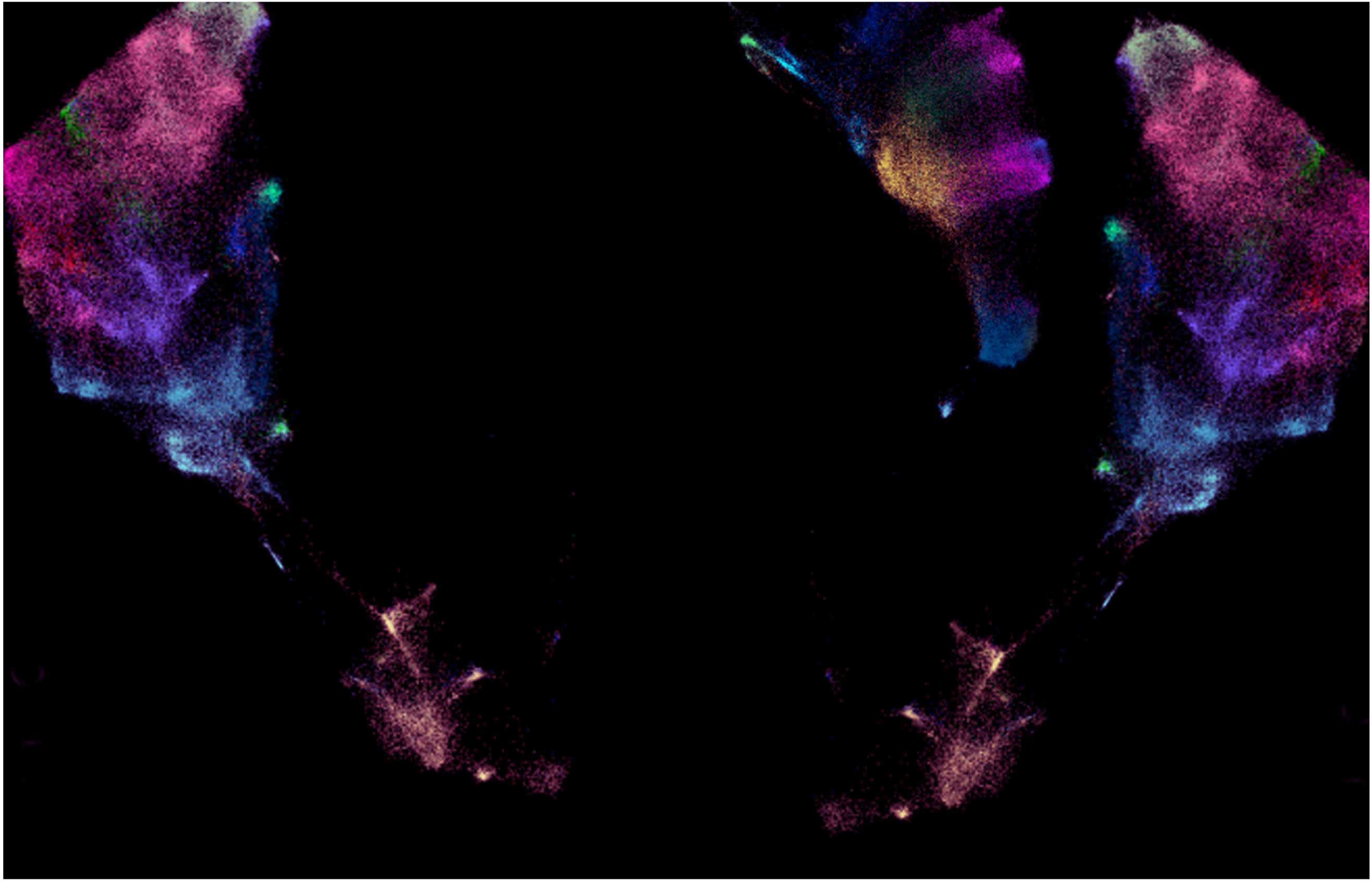
The subsequent analyses (powered by a co-ordinated effort with our co-authors), revealed populations of TCF7+ progenitor-like cells in equilibrium with subsets of PD-1hi dysfunctional (Tdys) and terminally differentiated, CD57+ dysfunctional (TDT) CD4 and CD8 T cells in NSCLC. Most importantly, the balance of these populations was shaped by TMB, favouring a high dysfunctional: progenitor T cell ratio in patients with increased numbers of mutations. Further experiments revealed T cell receptor sequence overlap, molecular reprogramming and similarity to MHC-multimer specific neoantigen reactive T cells amongst these cells; collectively indicative of a neoantigen-driven lineage trajectory. In the absence of immunotherapy, this imbalance in T cell homeostasis was associated with poor clinical outcome, suggesting that progenitor to dysfunctional subset conversion accompanies a fatal loss of local immune fitness. Thus, we had highlighted T cell intrinsic cellular and molecular pathways potentially involved in the progressive decay of neoantigen surveillance that occurs in the absence of immunotherapy. Finally, we generated a map of putative 'T to T' receptor ligand interactions that could help to identify previously understudied nodes of communication between T cells in the lung cancer TME.
These findings coalesce with other reports in the field12–15 and throughout the TRACERx collection, most notably those in Joshi et al NMED 201916, which demonstrates a relationship between clonal TCR expansion and TMB, and Rosenthal et al Nature, 20198 that details the extent of neoantigen-directed immune escape in the TRACERx cohort. Taken together, these data elucidate a pervasive axis of neoantigen surveillance in NSCLC that may synchronize tumor evolution and T cell dynamics, revealing key circuits potentially amenable to therapeutic intervention.
Whether re-direction of neoantigen-associated T cell differentiation pathways via antibody mediated immunomodulation or during manufacture of cell therapies will yield therapeutic benefit is yet to be determined. Moreover, it remains unclear how neoantigen type (e.g. single nucleotide variant vs insertion-deletion-derived), the mechanism of antigen presentation (e.g. lymph node vs tumor), or other features of the mutational architecture will sculpt the fate of neoantigen specific T cells. These and other outstanding questions remain the subject of ongoing investigation in the extended TRACERx cohort.
References
1. Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science (2015). doi:10.1126/science.aaa4971
2. Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non. 348, 1–6 (2016).
3. McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science (80-. ). (2016). doi:10.1126/science.aaf1490
4. Simoni, Y. et al. Bystander CD8+T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature (2018). doi:10.1038/s41586-018-0130-2
5. Gros, A. et al. Prospective identification of neoantigen-specific lymphocytes in the peripheral blood of melanoma patients. Nat. Med. 22, 433–438 (2016).
6. Tran, E. et al. T-Cell Transfer Therapy Targeting Mutant KRAS in Cancer. N. Engl. J. Med. 375, 2255–2262 (2016).
7. McGranahan, N. et al. Allele-Specific HLA Loss and Immune Escape in Lung Cancer Evolution. Cell 171, 1259-1271.e11 (2017).
8. Rosenthal, R. et al. Neoantigen-directed immune escape in lung cancer evolution. Nature 567, 479–485 (2019).
9. Jamal-Hanjani, M. et al. Tracking the Evolution of Non–Small-Cell Lung Cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
10. Day, C. L. et al. PD-1 expression on HIV-specific T cells is associated with T-cell exhaustion and disease progression. Nature 443, 350–354 (2006).
11. Zajac, A. J. et al. Viral Immune Evasion Due to Persistence of Activated T Cells Without Effector Function. J. Exp. Med. 188, 2205–2213 (1998).
12. Schietinger, A. et al. Tumor-Specific T Cell Dysfunction Is a Dynamic Antigen-Driven Differentiation Program Initiated Early during Tumorigenesis. Immunity 45, 389–401 (2016).
13. Utzschneider, D. T. et al. T Cell Factor 1-Expressing Memory-like CD8+ T Cells Sustain the Immune Response to Chronic Viral Infections. Immunity 45, 415–427 (2016).
14. Zhou, X. et al. Differentiation and Persistence of Memory CD8+ T Cells Depend on T Cell Factor 1. Immunity 33, 229–240 (2010).
15. Chen, Z. et al. TCF-1-Centered Transcriptional Network Drives an Effector versus Exhausted CD8 T Cell-Fate Decision. Immunity 1–16 (2019). doi:10.1016/J.IMMUNI.2019.09.013
16. Joshi, K. et al. Spatial heterogeneity of the T cell receptor repertoire reflects the mutational landscape in lung cancer. Nat. Med. (2019). doi:10.1038/s41591-019-0592-2
Follow the Topic
-
Nature Cancer
This journal aims to provide a unique forum through which the cancer community will learn about the latest, most significant cancer-related advances across the life, physical, applied and social sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Cancer Neuroscience: from mechanisms to therapy
Publishing Model: Hybrid
Deadline: Jan 30, 2027



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in