CRISPR-based gene-drives: from eukaryotes to prokaryotes
Published in Bioengineering & Biotechnology
Active genetics technology greatly biases transmission of genetic traits, bypassing traditional constraints of Mendelian inheritance and spreading rapidly through wild populations (1). Molecularly defined gene drive constructs in Saccharomyces cerevisiae were copied at efficiencies exceeding 99% when mated to wild yeast (2). Using active genetics to bias inheritance of desired alleles in laboratory mice appears to enable rapid assembly of otherwise impractical genotypes involving multiple homozygous genes (3). In another example, vector mosquito Anopheles stephensi was engineered to carry Cas9, a gRNA and antimalarial gene cassette and might be introduced to block propagation of the malaria parasite Plasmodium falciparum (4).
Could we expand the versatility of active genetics from eukaryotes to prokaryotes? “Very unlikely considering the haploid nature and the single copy of each gene in a prokaryote cell context”, we thought initially. But what about a bacterial strain harboring a plasmid with multiple copies of the target gene, where the gene drive could propagate and spread within a single cell? Definitely worth a try. We brought together the expertise of the Bier lab in CRISPR-based gene drives with the microbiology research focus of the Nizet Lab to establish proof-of-principle of the first bacterial gene-drive system, dubbed Pro(karyotic)-Active Genetics or “Pro-AG”.
Our main motivation has been to explore this revolutionary genetic platform as a weapon in the fight against antibiotic resistance, a critical public health problem that has been found in all regions of the world. Some estimates hold that if dramatic actions are not taken soon, antibiotic resistance could cause 10 million deaths each year by 2050, crippling modern medicine and triggering catastrophic economic damage. We contemplated if modifying a standard bacterial CRISPR editing system by incorporating an active self-copying mechanism could enhance efficiency in scrubbing antibiotic resistance in prokaryotes.
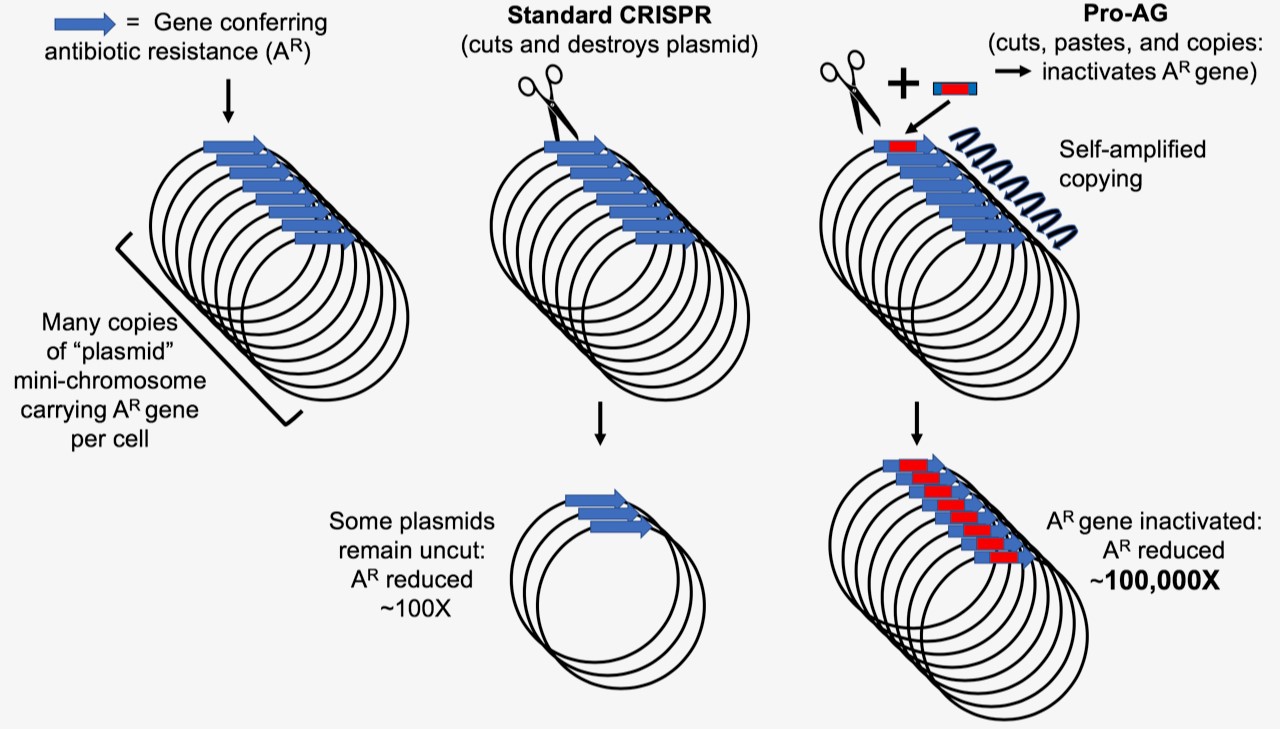
We designed an analogous CRISPR-based gene-drive system for the bacterium Escherichia coli that efficiently copies a gRNA cassette and any adjacent cargo that are flanked with sequences homologous to the targeted gRNA/Cas9 cleavage site. We observed that Pro-AG functionally inactivates an antibiotic resistance marker on a high copy number plasmid with ~100-fold greater efficiency than standard CRISPR-based methods, reflecting an amplifying positive feedback loop due to increasing gRNA dosage. In effect, each insertional editing event expands the pool of functional gRNA donor scaffolds with extended homology arms in their newly copied plasmid context, initiating a “chain reaction” accelerating further insertional events. Remarkably, the high efficiencies achieved with Pro-AG copying system in bacteria are comparable to those attained by CRISPR-based gene-drive systems in diploid organisms.

CAPTION: Andrés Valderrama, PhD examining high efficiency Pro-Ag editing of E. coli to replace a target gene with a functional GFP gene.
Pro-AG can likewise effectively edit large plasmids, single-copy genomic targets or introduce functional cargo genes. We hope that multiple genome-engineering applications could ultimately benefit from incorporation of the Pro-AG platform, including elimination of bacterial virulence factors carried by diverse episomal elements and resistance determinants in commensal bacteria or prokaryotic pathogens, targeting antibiotic resistance genes from bacteria in the environment, livestock, inland fish farms, or sewage treatment ponds, reprogramming genetic circuits impacting bacterial physiology, or tailoring interactions among the microbiota in environmental or host niches. To take Pro-AG to the next level for these biomedical and/or biotechnological applications, it will be necessary to couple Pro-Ag to efficient delivery systems such as phage vectors or conjugation mechanisms for spread throughout bacterial populations.
- Gantz, V. M. & Bier, E. The mutagenic chain reaction: a method for converting heterozygous to homozygous mutations. Science 348, 442–444 (2015).
- DiCarlo, J. E., Chavez, A., Dietz, S. L., Esvelt, K. M. & Church, G. M. Safeguarding CRISPR-Cas9 gene drives in yeast. Nat. Biotechnol. 33, 1250–1255 (2015)
- Grunwald, H. A. et al. Super-Mendelian inheritance mediated by CRISPR-Cas9 in the female mouse germline. Nature 566, 105–109 (2019).
- Gantz, V. M. et al. Highly efficient Cas9-mediated gene drive for population modification of the malaria vector mosquito Anopheles stephensi. Proc. Natl. Acad. Sci. U.S.A. 112, E6736-43 (2015).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Jun 01, 2026
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in