Danger, pathogen may explode: revealing how specialized bacterial cells release toxic payloads
Published in Chemistry, Microbiology, and Cell & Molecular Biology

Bacteria have mastered the art of survival and adaptation, sometimes to the detriment of their hosts. In order to resist mechanical forces that may remove them from host surfaces, pathogenic bacteria first attempt to establish a foothold by using specialized adhesion molecules to anchor themselves to specific host receptors. Once attached, bacteria deploy an arsenal of virulence factors to breach the host's defenses. These include toxins, enzymes, and other proteins that manipulate the host's cellular machinery and immune responses.
Toxin complex (Tc) toxins are a potent type of secreted virulence factor primarily employed by Gram-negative insect pathogens. One property that makes them special is their extremely large size, which can rival that of ribosomes. Another is their modularity: the pentameric “A-component” responsible for binding to target cells and cytotoxin injection can combine with a range of “BC-cocoons”, each of which can carry a different type of cytotoxin. This plug-and-play functionality enables Tc toxins to simultaneously attack several different intracellular targets, improving their odds of killing host cells.
Like other secreted toxins of Gram-negative bacteria, Tc toxins are synthesized in the bacterial cytoplasm and must cross multiple barriers on the way out of the cell. These include the phospholipid inner membrane, peptidoglycan sacculus, and lipopolysaccharide outer membrane. Most other (smaller) toxins of Gram-negative bacteria achieve this either by being injected directly into host target cells or ejected into extracellular space by specialized secretion systems, with the use of each secretion system introducing unique requirements to the toxin such as being unfolded during translocation or containing a signal sequence that directs it to the correct export apparatus. Tc toxins however lack such signal sequences and appear to be far too large to fit the physical dimensions of secretion systems established for smaller toxins.
So how do they then manage to get out?
To determine this, we turned to Yersinia entomophaga, a distant relative of the Black Death pathogen. The remarkable efficiency with which Y. entomophaga kills insects is dependent on its sole Tc toxin YenTc, which it abundantly secretes into growth media in laboratory conditions. After initially establishing controllable YenTc secretion conditions by modulating growth media pH, we knocked out all conventional Y. entomophaga secretion systems one by one to see which of them was responsible for YenTc secretion. To our surprise, none of these knockouts stopped secreting YenTc into the growth media. Moreover, mass spectrometry revealed that the secreted proteome was enriched not only in YenTc, but also in many other toxins and virulence factors, most of which also lacked known signal sequences for secretion. One enriched protein – an endolysin – caught our eye since endolysins are a core component of the very recently established family of type 10 secretion systems (T10SS). A look into the Y. entomophaga genome confirmed that the endolysin in question was indeed located in a T10SS operon, which also contained a holin, i-spanin and o-spanin as core components. Knocking these out stopped Y. entomophaga secretion in its tracks – and meant that we finally found the machinery responsible for export of Tc toxins as well as the other virulence factors not assigned to a known secretion pathway!
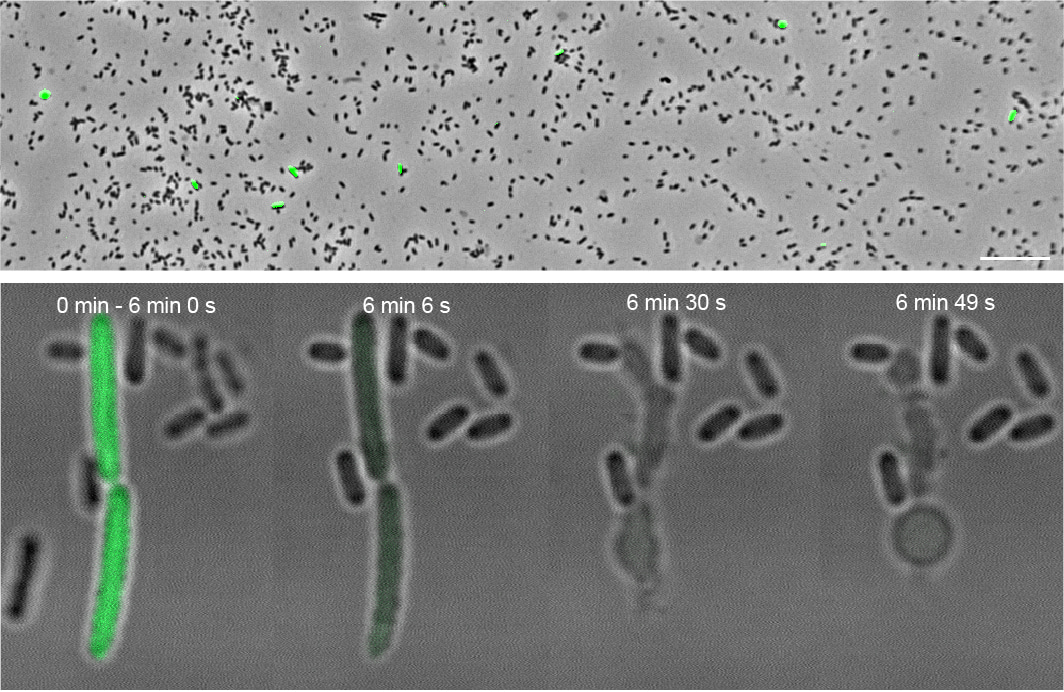
This was not the only surprise discovery that awaited us. We wanted to visualize the process of T10SS-mediated secretion, which had so far not been investigated in detail, and therefore inserted a green fluorescent protein (GFP) into the YenTc operon to assist with this. To our amazement, only a small subpopulation of these genetically identical cells expressed the Tc toxin at any given time. Furthermore, these cells were much larger than their YenTc-nonexpressing brethren and upon exposure to secretion-inducing conditions, they explosively burst and released the cytoplasmically accumulated YenTc into the environment – not at all a typical secretion mechanism! We named this specialized subpopulation of kamikaze toxin-producers “soldier cells”. The suicidal nature of this protein release mechanism simultaneously explains why only a small percentage of cells show a soldier cell phenotype.
Eventually, we were able to uncover the genetic switch that converts normal cells into soldier cells, and found that it is also responsible for coupling the production of the T10SS to the many toxins and virulence factors that soldier cells exclusively produce. This ensures that a soldier cell will not sacrifice itself without releasing its deadly payload and, conversely, that precious cellular resources are not wasted to produce toxins that the cell would have no way of exporting. We also found that this genetic switch is inactive in environments that poorly mimic an insect host, preventing futile production of soldier cells.
Using these findings, we generated chemically inducible soldier cell strains which had the T10SS either intact or lacking certain core components. By subjecting these strains to secretion-inducing conditions, we were able to either let them undergo the full T10SS-mediated protein release or to block this process at different stages where the missing T10SS components were needed. After vitrifying these cells by plunge freezing and removing excess material using a focused ion beam, we visualized them on an electron microscope using cryo-electron tomography, a revolutionary structural biology technique capable of delivering detailed 3D images of proteins in the context of their native cellular ultrastructure. Using this approach, we were able to visualize the step-by-step action of a T10SS in unprecedented detail, exposing the drastic changes occurring in the bacterial cell wall under the influence of the various T10SS components, and to track the subcellular localization of the fully assembled YenTc toxins from the bacterial cytoplasm to extracellular space.
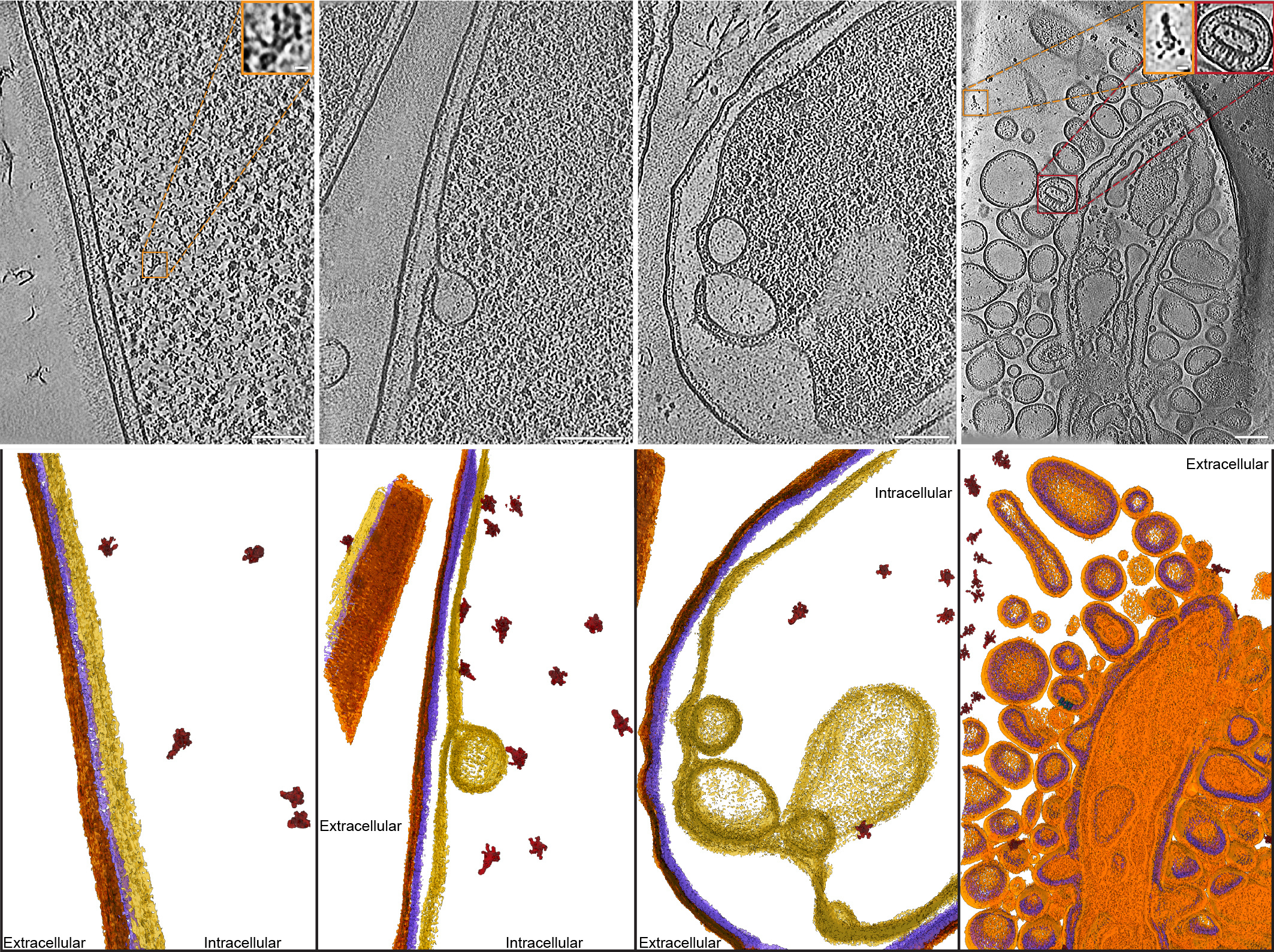
What are the most important takeaways of this study?
First, it establishes a pathway for Gram-negative pathogens to rapidly release virulence factors in a way that bypasses the usual stringent cargo prerequisites of using canonical secretion systems. Second, it identifies a new specialized subtype of bacterial cells which are sacrificed in order to devastate the invaded host – and thus ensure the rest of its bacterial brethren benefit. Finally, it offers the tantalizing insight that bacterial pathogenicity may in some cases rely on the presence of a soldier cell minority. If this turns out to also be true for certain human pathogens, then specifically targeting such soldier cells may represent a promising new treatment approach.
Follow the Topic
-
Nature Microbiology

An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.
Related Collections
With Collections, you can get published faster and increase your visibility.
The Clinical Microbiome
Publishing Model: Hybrid
Deadline: Mar 11, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in