Tick-borne diseases are considered a major risk to human health and are increasing in numbers due to environmental changes that impact the geographic distribution of the tick vectors. Among various pathogens transmitted by ticks, Babesia parasites are particularly important because of their ability to multiply within host erythrocytes and lyse these cells at the end of each cycle thereby causing symptoms similar to those caused by malaria parasites. Several Babesia species have been found to infect humans with B. microti responsible for the majority of cases worldwide. Another Babesia species Babesia duncani has previously been shown to cause severe infection in both immunocompromised and immunocompetent individuals and causes lethal infection in immunocompetent mice and hamsters. The parasite has also been shown to be an excellent model to study intraerythrocytic parasitism and to evaluate the in vitro and in vivo efficacy of new antiparasitic drugs due to its ability to propagate continuously in human red blood cells in vitro and to cause infection in mice. However, the biology of this parasite has only recently started to be elucidated.
In this report (Singh et al., 2023 Nature Microbiology), we report the first detailed sequence, assembly, annotation and 3D structure of the nuclear genome of B. duncani. The data generated from these analyses have shed light on the factors driving its virulence and identified potential drug targets for development of more effective antibabesial therapies and possibly therapies for other intraerythrocytic parasites.
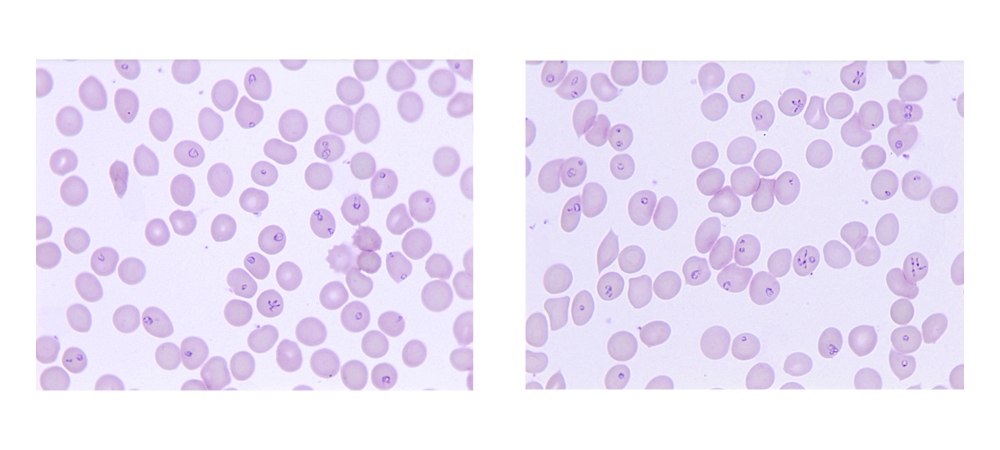
Our analysis revealed that the B. duncani nuclear genome consists of five chromosomes of sizes ranging between 3.13 and 0.35 MB. 4,222 genes were annotated and found to have an average length of 1656 bp. Roughly 60% of the genome was calculated to be protein coding. B. duncani was found to share 842 proteins (20% of the proteome) with other apicomplexan parasites with most conserved proteins involved in DNA replication, transcription, and translation. Another 869 proteins of B. duncani are unique to this parasite and have no predicted function.
This study further identified more than 700 genes that belong to 73 multigene families unique to B. duncani and 105 families belonging to orthologous apicomplexan groups. The expression, localization and epigenetic profiles of these genes were determined by RNAseq, HiC, ChIP-seq and FISH analyses and were similar to those of gene families that encode virulence factors in the human malaria parasite P. falciparum. Collectively, the data point to the possibility of antigenic variation in this parasite and a possible role of these newly identified proteins in B. duncani virulence.
Metabolic reconstitution and drug target mining identified several enzymes and functions that could be targeted by available or newly developed antiparasitic drugs. The conservation of these targets among different apicomplexan pathogens makes B. duncani a perfect model organism for other parasites to pursue future medicinal chemistry efforts to identify molecules that could be rapidly evaluated in vitro and in mice to assess their efficacy and safety.
The multi-omics analysis presented in this paper provides unprecedented insight into the biology, pathogenesis, and drug susceptibility of B. duncani. The data reported in this study have been made available to the scientific community and will stimulate efforts to develop better and more sensitive diagnostic assays and better treatment strategies to treat human babesiosis.
Follow the Topic
-
Nature Microbiology

An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in