Defining the chromatin and transcriptional regulation of organ specification
Published in Healthcare & Nursing
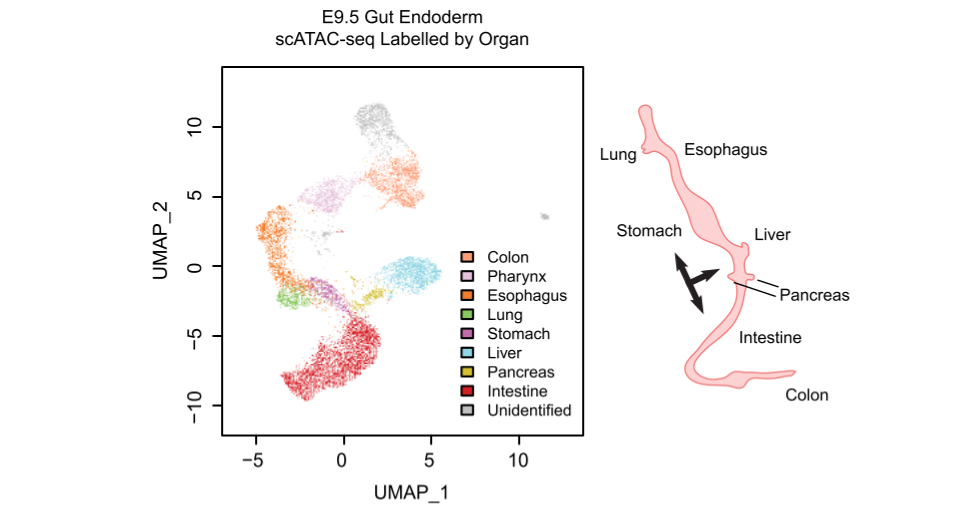
In mammalian embryonic development, the development and differentiation of primitive gut tube has always been a research focus, owing to its role as the precursor to a number of important organs, which include the colon, intestine, stomach, pancreas, liver, lung, esophagus, pharynx, and thyroid1. While mesodermal cells are going to become mesenchymal cells of the gut2, endodermal cells as a single-cell layer inside the primitive gut tube will develop to form the functional epithelial cells in different organs, such as the absorptive enterocytes in the intestine, the acid-secreting parietal cells in the stomach, and the hormone-producing cells of the exocrine lineage in the pancreas3–5. Thus, fate decisions of these endodermal cells have attracted our particular attention. During organogenesis, typically from E9.5 to E13.5, the gut tube starts to form morphological identities of organs. However, the endodermal layer remains uniform over this time, and they only become more differentiated during the regionalization process later1,6.
Nevertheless, previous studies showed that the seemingly identical endodermal cells present different gene expression patterns along the anterior-posterior (AP) axis1,7. With the application of single-cell RNA-seq, recent studies revealed that endodermal cells show organ-specific characteristics in gene expression at the earliest stage of gut development, indicating that organ fates are pre-programmed into the cells8,9. However, it was still unclear how this spatiotemporal control over transcription is regulated, and in particular, which transcription factors (TFs) are involved. Assay for Transposase-Accessible Chromatin using sequencing (ATAC-seq) can measure genome-wide chromatin accessibility and enables the identification of potential regulatory regions and TFs. Recent studies have utilized bulk ATAC-seq on the gut tube at later stages of gut development, between E11.5 and E16.510–12. Considering the organogenesis is well under way at the earliest stages and the insufficient resolution of bulk analyses, a single-cell level epigenetic landscape of gut organ development at an early stage needs to be developed urgently. To bridge this gap, we utilized the recently developed single-cell (sc) ATAC-seq technique to uncover patterns of chromatin accessibility in early gut development (Figure 1).
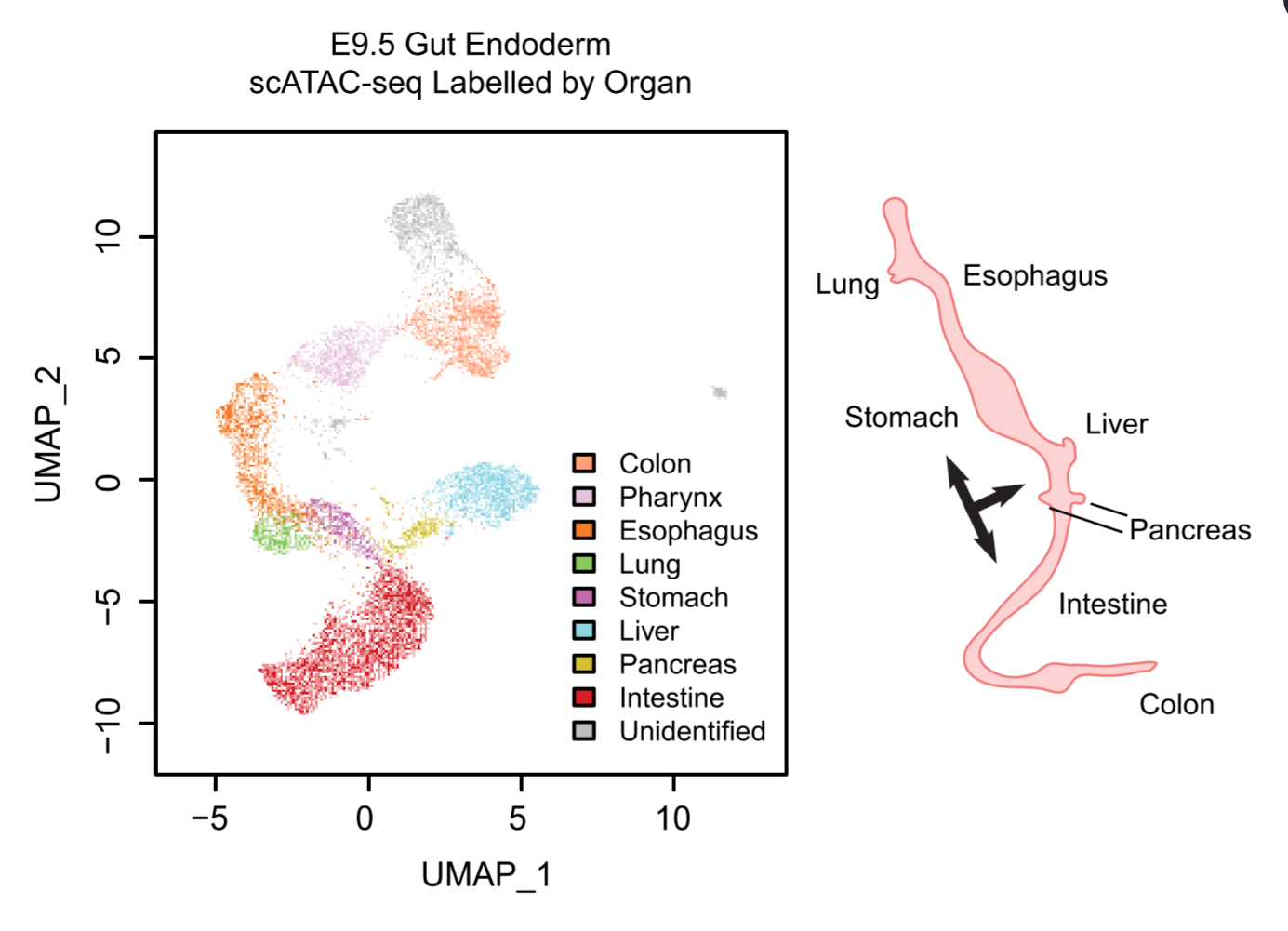
Figure 1
Single-cell scatter plot under UMAP dimensionality reduction demonstrating the clustering and organ assignment of cells. The organ assignment is conducted by comparing the gene score of known marker genes of organs. Each dot represents an individual cell and is colored by the assigned organ type. The spatial distribution pattern of organs on the UMAP resembles the actual anatomic positions of the organs on the right diagram. Black arrows label organ specification.
Do endodermal cells of the primitive gut tube show organ-specific chromatin patterns in early development?
We isolated Epithelial Cell Adhesion Molecule (EPCAM) expressing endodermal cells from the primitive gut tube through fluorescence-activated cell sorting at E9.5 for scATAC-seq. In total, we obtained 12,067 cells with high quality ATAC-seq data. After clustering cells based on their genome-wide chromatin accessibility patterns, we were intrigued to find that cluster-specific accessible regions are enriched around the transcription start sites of some well-known organ marker genes, indicating the lineage fate of each cell cluster. Based on this, we successfully assigned cell clusters to organs they are going to form in the future. We were very excited to find that the organ projection patterns on the scATAC-seq UMAP plot matches the anatomical patterns of the organs to be established at later stages (Figure 1).
What can multi-organ data tell us?
Two questions that came to our minds were how the development of different organ precursors are governed in different ways and whether the regulatory mechanism is dynamic during the whole process of organ development. We addressed these questions with cross-organ and temporal analyses covering 4 major organs (lung, stomach, pancreas, intestine). These analyses reveal that each organ in the developing gut endoderm uses its own suite of regulatory machinery to push forward its developmental trajectory. Within each organ, chromatin accessibility shows dynamics during development, implying different TFs dominating the regulation at different stages. In late organ development, such stage-specific regulatory suites determine further branching of organs, for example, the development from early-stage intestine to colon and small intestine.
How do SOX2 and CDX2 impact lineage fate decisions by regulating chromatin accessibility?
We then turned our attention to TFs that are critical for maturation and differentiation of epithelial cells in the developing gut. SOX2 and CDX2 are two TFs regulating development of the foregut (stomach, lung esophagus, pharynx) and hindgut (small intestine, colon), respectively in a mutually exclusive way7. We therefore wanted to better understand how they interact with chromatin to regulate organ fates in the developing gut. Utilizing the published bulk ATAC-seq data from Sox2 knockout stomach and Cdx2 knockout intestinal samples, we were able to investigate how these TFs help to determine lineage fate decisions by summarizing relationships between the chromatin accessibility changes associated with Sox2/Cdx2 knockout and the organ-specific chromatin patterns of the gut tube. Finally, we show that Sox2 knockout stomachs will experience a fate shift from the stomach to neighboring organs such as the lung and, to a smaller extent, the intestine, while Cdx2 knockout leads to a fate shift from the intestine to the foregut.
Will SOX2 expression change lineage fate decisions alone?
Loss of key lineage-specific TFs in the stomach and intestine alters patterns of chromatin accessibility and gene expression, ultimately leading to changes in cell identity10–15. We next sought to answer whether Sox2 expression alone at earlier developmental stages could sufficiently alter organ identity. By generating a genetic mouse model (Pdx1Cre/+;R26Sox2-IRES-GFP ) targeting Sox2 expression to the hindstomach, the intestine and the pancreas, we were able to track gene expression and organ identity changes upon Sox2 overexpression. As a result, we found that misexpression of Sox2 in gut precursors does alter organ identity. Specifically, both the pancreas and intestine lost their original fates and obtained foregut-like features.
What about the misexpression of SOX2 in cancers?
SOX2 has been shown to be abnormally activated in colon and pancreatic cancers16,17. By conditionally activating Sox2 expression in intestine epithelial cells (mouse model: Villin CreERT2/+;R26Sox2-IRES-GFP), we found prolonged Sox2 expression is sufficient to impede intestinal epithelial differentiation in adult homeostasis. Through a similar experiment on the pancreas (mouse model: Mist1 CreERT2/+;R26Sox2-IRES-GFP), a dramatic shift of a number of SOX2 expressing cells from acinar cells to ductal cells was observed, which is consistent with the acinar-ductal metaplasia (ADM) process in pancreatic cancer18. Together with the fact that these cells are hyperproliferative, our data provide evidence that abnormal Sox2 expression in adult pancreatic epithelial cells may alter lineage fate decisions involved in tumorigenesis.
In summary, our study generated a comprehensive map of epigenomic changes throughout gut development, filling the knowledge gap in this field. We demonstrated how TFs and chromatin accessibility regulate transcription in an organ-specific and dynamic way during the gut endoderm development. We further uncovered the lineage fate shifts induced by the dysregulation of Sox2 and Cdx2 from the view of chromatin accessibility. Finally, the abnormal expression of Sox2 in several mouse models was sufficient to change lineage fate decisions in both development and adult homeostasis. Since lineage fate decisions are frequently altered in disease such as cancer, our study would likely help understand the origin and development of certain types of cancer.
References:
- Zorn, A. M. & Wells, J. M. Vertebrate endoderm development and organ formation. Annu. Rev. Cell Dev. Biol. 25, 221–251 (2009).
- Lewis, S. L. & Tam, P. P. L. Definitive endoderm of the mouse embryo: formation, cell fates, and morphogenetic function. Dev. Dyn. an Off. Publ. Am. Assoc. Anat. 235, 2315–2329 (2006).
- Chin, A. M., Hill, D. R., Aurora, M. & Spence, J. R. Morphogenesis and maturation of the embryonic and postnatal intestine. Semin. Cell Dev. Biol. 66, 81–93 (2017).
- Kim, T.-H. & Shivdasani, R. A. Stomach development, stem cells and disease. Development 143, 554–565 (2016).
- Stanger, B. Z. & Hebrok, M. Control of cell identity in pancreas development and regeneration. Gastroenterology 144, 1170–1179 (2013).
- Thompson, C. A., DeLaForest, A. & Battle, M. A. Patterning the gastrointestinal epithelium to confer regional-specific functions. Dev. Biol. 435, 97–108 (2018).
- Sherwood, R. I., Chen, T.-Y. A. & Melton, D. A. Transcriptional dynamics of endodermal organ formation. Dev. Dyn. an Off. Publ. Am. Assoc. Anat. 238, 29–42 (2009).
- Han, L. et al. Single cell transcriptomics identifies a signaling network coordinating endoderm and mesoderm diversification during foregut organogenesis. Nat. Commun. 11, 4158 (2020).
- Nowotschin, S. et al. The emergent landscape of the mouse gut endoderm at single-cell resolution. Nature 569, 361–367 (2019).
- Kumar, N. et al. The lineage-specific transcription factor CDX2 navigates dynamic chromatin to control distinct stages of intestine development. Development 146, (2019).
- Banerjee, K. K. et al. Enhancer, transcriptional, and cell fate plasticity precedes intestinal determination during endoderm development. Genes Dev. 32, 1430–1442 (2018).
- Francis, R. et al. Gastrointestinal transcription factors drive lineage-specific developmental programs in organ specification and cancer. Sci. Adv. 5, 1–13 (2019).
- Chen, L. et al. HNF4 factors control chromatin accessibility and are redundantly required for maturation of the fetal intestine. Development 146, (2019).
- Verzi, M. P. et al. Differentiation-Specific Histone Modifications Reveal Dynamic Chromatin Interactions and Partners for the Intestinal Transcription Factor CDX2. Dev. Cell 19, 713–726 (2010).
- Verzi, M. P., Shin, H., San Roman, A. K., Liu, X. S. & Shivdasani, R. A. Intestinal master transcription factor CDX2 controls chromatin access for partner transcription factor binding. Mol. Cell. Biol. 33, 281–292 (2013).
- Herreros-Villanueva, M. et al. SOX2 promotes dedifferentiation and imparts stem cell-like features to pancreatic cancer cells. Oncogenesis 2, e61–e61 (2013).
- Lundberg, I. V et al. SOX2 expression is associated with a cancer stem cell state and down-regulation of CDX2 in colorectal cancer. BMC Cancer 16, 471 (2016).
- Storz, P. Acinar cell plasticity and development of pancreatic ductal adenocarcinoma. Nat. Rev. Gastroenterol. Hepatol. 14, 296–304 (2017).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in