Familial endemic Burkitt lymphoma in Uganda
Published in Cancer
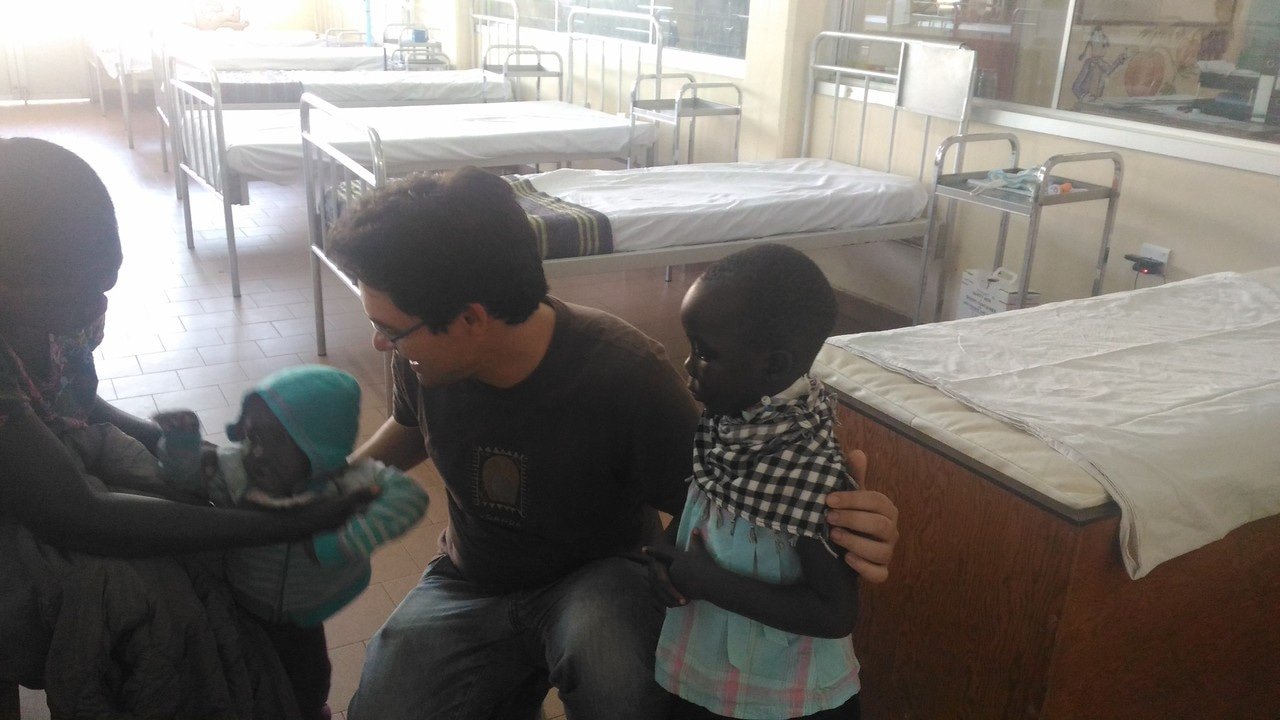

Burkitt lymphoma (BL) is a rare cancer first described in African children by Denis Burkitt in 1958. Its rapidly progressive course and high incidence in equatorial Africa attracted interest of researchers searching for a viral cause of cancer and led to discovery of Epstein-Barr virus (EBV), the first human virus linked to a human cancer, which also paved the way for discovery of other cancer-causing viruses. Epidemiological reports of BL co-distribution with Plasmodium falciparum infection led to the malaria hypothesis of BL etiology. Tumor studies led to discovery of chromosomal abnormalities, strongly focused attention on the genetic basis of cancer and led to the discovery of MYC as a key molecule involved in carcinogenesis. Interestingly, isolated reports of BL diagnosed in relatives in the 1960s and 70s in different areas, including in Bwamba County in Uganda and Shirati District in Tanzania, and outside Africa triggered excitement with many expecting that study of BL might lead to discovery of germline DNA variants that may predispose to BL. However, civil disturbance in Africa interrupted this research [1].
In 2008, Dr. Joseph Fraumeni Jr., Director of the National Cancer Institute’s (NCI) Division of Cancer Epidemiology and Genetics approved a plan for a large population-based study to investigate comprehensively the infectious, immunological, and genetic risk factors for BL.
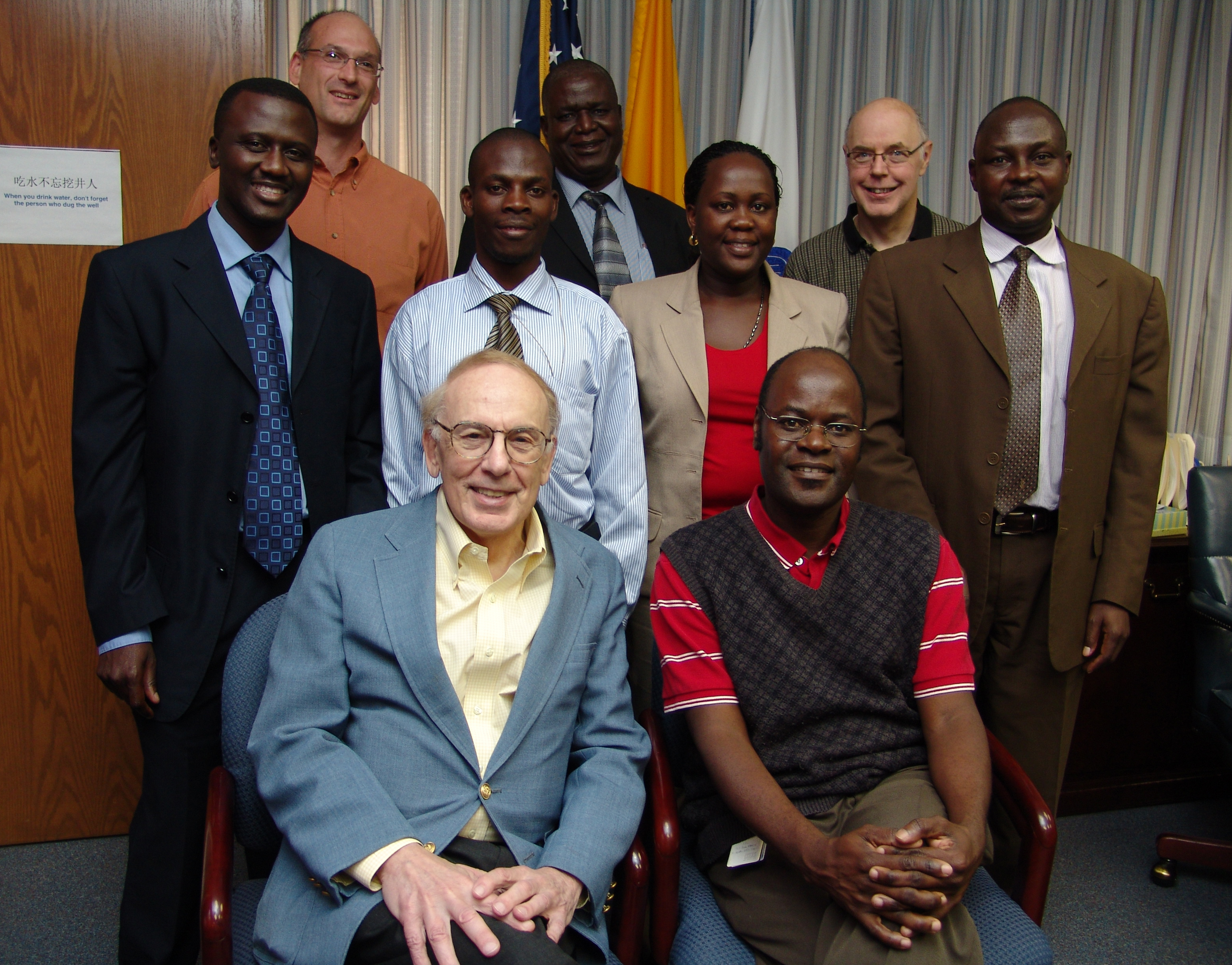

The study, called EMBLEM (https://emblem.cancer.gov/), was designed to enroll children and minors (0-15 years old) with BL in six regions in Uganda, Kenya and Tanzania (2 per country).
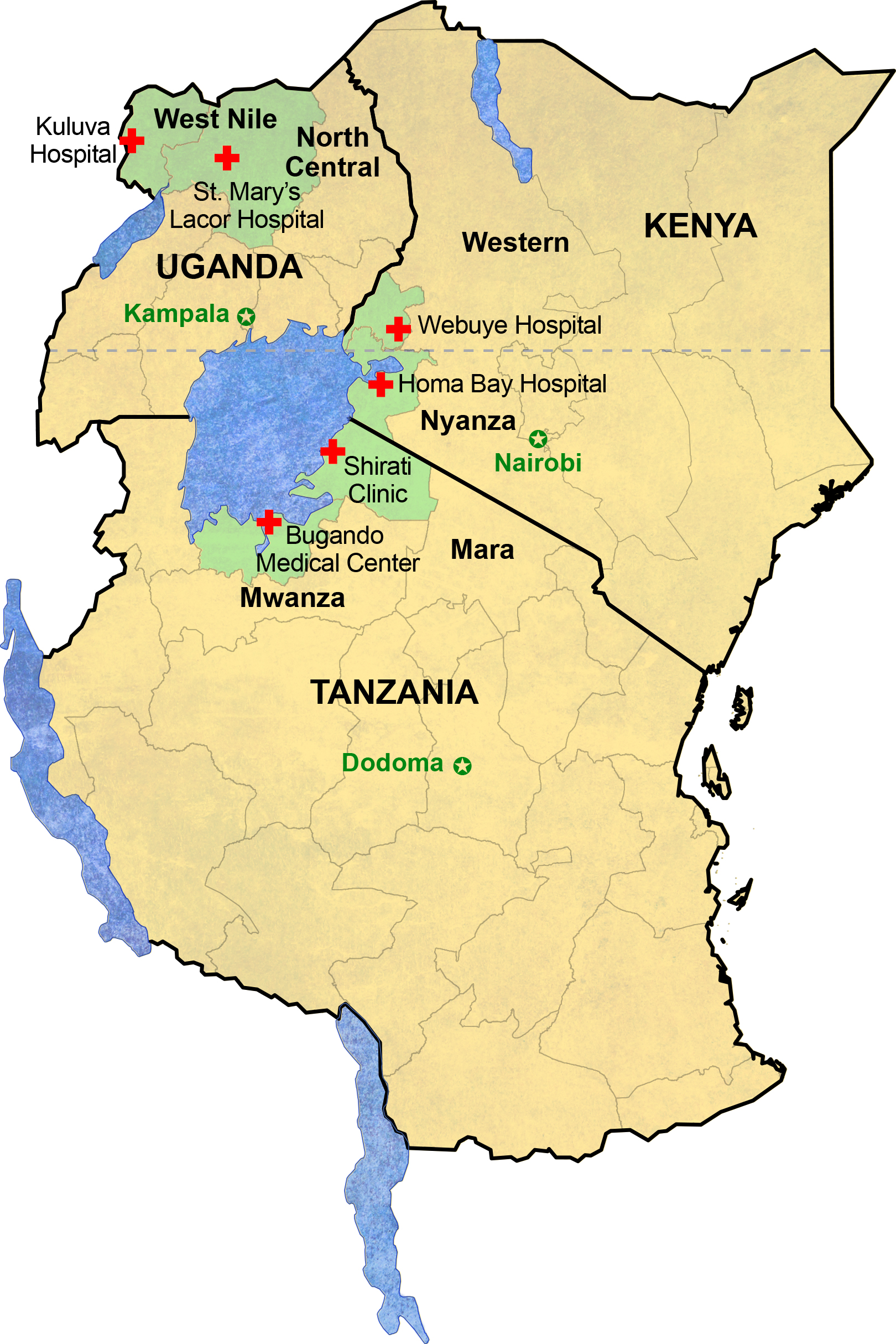
However, the plans to enroll cases almost collapsed when it was discovered that the state of local pathology in the region had not kept pace with that practiced in developed countries and that the quality clinical and epidemiological data would be undermined [2]. Our collaborator, Prof. Leona Ayers, at Ohio State University, traveled to Uganda and gave a refresher training for hospital technicians to improve the quality of histopathology and formalin fixed paraffin embedded tumor blocks.
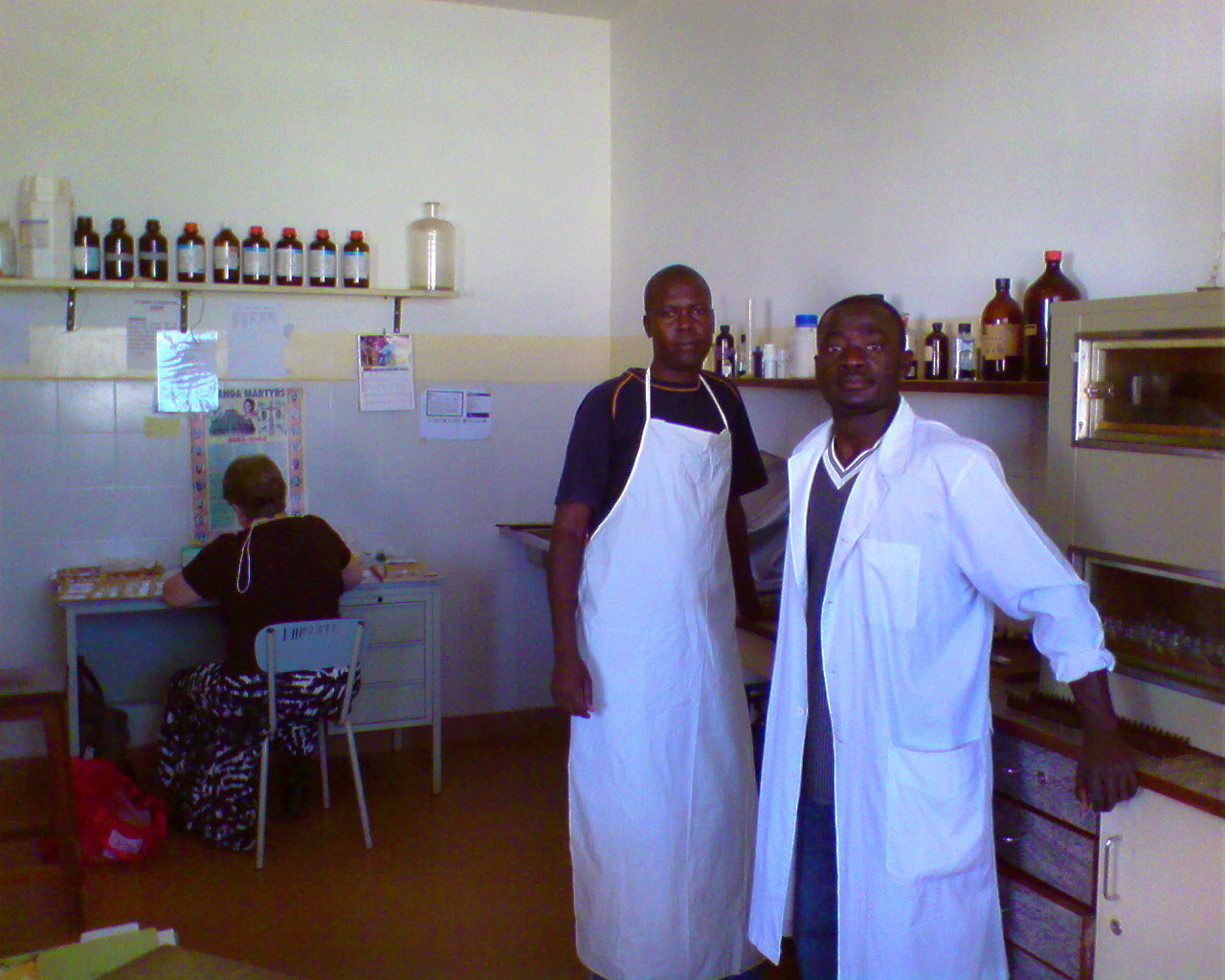

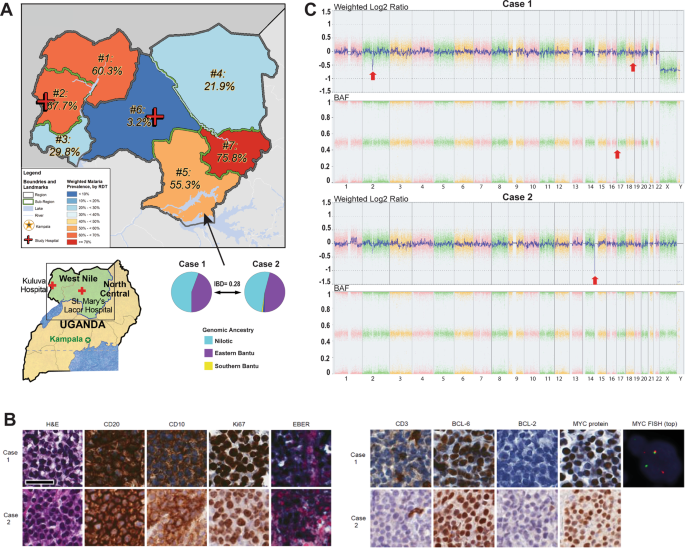
This training put case enrollment back on track (2010-2016). In 2016, we genotyped the first 198 cases (two-thirds confirmed by pathology) enrolled in Northern Uganda at > 4 million loci using the Illumina 5M array. Our challenge to find a dedicated analyst to sift through the extensive genomic data was disappeared when our paths crossed with Mateus H. Gouveia (in poster image), a population geneticist from Brazil, who was studying African ancestry in the diaspora in Brazil and was looking to add samples from populations in Africa. Mateus visited Gulu in Uganda in 2016 and was intrigued by populations living in the River Nile Basin. He embarked on the first population genetics study of admixture, gene flow and natural selection in eBL belt populations (here)[3] alongside his prior interest in the "origin and admixture of African ancestry populations" [4]. To our surprise, we discovered based on genomic data this first set that two of the cases (one boy and one girl) were second-degree relatives. At first, we considered this to be a good anecdote, but Dr. Gouveia, after hearing about the isolated familial BL cases, urged action. Thanks to improvements in pathology, both cases had tumor tissue; so the local diagnosis was confirmed by outside review at NIH and Ohio State University, and OncoScan SNP-array analysis at the University of Ulm, making these the first African BL cases in relatives to be robustly confirmed. Next, we performed careful, coordinated, extensive exome sequence analysis was done, and the results compared with exome data from BL and non-BL cases in Tanzania, US, Germany, and Guatemala. We found in both children’s germline DNA genetic variants in TCF4 and CHD8. TCF4 has been implicated in the ID3/TCF3 pathway in BL based on somatic genetic data by the German ICGC MMML-Seq consortium [5] . CHD8 was recently found to be recurrently mutated in BL in the BL Genome Sequencing Project [6].
Good research is responsive to patient and community needs and spawns more research [1]. Our research was responsive to patient diagnostic and treatment needs, as well as to the community ethical concerns and perceived needs, expressed by patients, parents, community leaders and representatives on the Community Advisory Board.
The study met NCI’s goal to introduce genomic methods to the study of the genetic architecture of BL. The TCF4 and CHD8 genetic variants are new clues which warrant more investigation. We show the potential to identify families with heightened risk of BL who can be studied for genetic susceptibility (This article is available online at: https://doi.org/10.1038/s41375-020-01052-w).
References:
- Mbulaiteye SM, Talisuna AO, OgwangMD, et al. African Burkitt's Lymphoma: Could Collaboration with Hiv-1 and Malaria Programmes Reduce the High Mortality Rate? Lancet 2010; 375:1661-3.DOI: 1016/S0140-6736(10)60134-1
- Ogwang MD, Zhao W, Ayers LW, Mbulaiteye SM. Accuracy of Burkitt lymphoma diagnosis in constrained pathology settings: importance to epidemiology. Arch Pathol Lab Med. 2011 Apr; 135(4):445-50. DOI: 1043/2009-0443-EP.1
- Gouveia MH, Bergen AW, Borda V, et al. Genetic signatures of gene flow and malaria-driven natural selection in sub-Saharan populations of the “endemic Burkitt Lymphoma Belt” PLoS Genet. 2019 Mar 8;15(3):e1008027. doi: 10.1371/journal.pgen.1008027.
- Gouveia MH, Borda V, Leal TP, et al. Origins, Admixture Dynamics, and Homogenization of the African Gene Pool in the Americas. Mol Biol Evol. 2020 Jun 1;37(6):1647-1656. doi: 10.1093/molbev/msaa033.
- Richter J, Schlesner M, Hoffmann S, Kreuz M, Leich E, Burkhardt B et al. Recurrent mutation of the ID3 gene in Burkitt lymphoma identified by integrated genome, exome and transcriptome sequencing. Nat Genet 2012; 44: 1316–1320. DOI: 10.1038/ng.2469
- Grande BM, Gerhard DS, Jiang A, et al. Genome-wide discovery of somatic coding and non-coding mutations in pediatric endemic and sporadic Burkitt lymphoma. Blood. 2019. doi: 10.1182/blood-2018-09-871418.
Follow the Topic
-
Leukemia
This journal publishes high quality, peer reviewed research that covers all aspects of the research and treatment of leukemia and allied diseases. Topics of interest include oncogenes, growth factors, stem cells, leukemia genomics, cell cycle, signal transduction and molecular targets for therapy.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in