High affinity team creates spatial biology resources for construction of a Human Reference Atlas
Published in Protocols & Methods
Coming together is a beginning, staying together is progress, and working together is success. ~Henry Ford
Our story begins at the National Institutes of Health (NIH) and the Human Cell Atlas (HCA) joint meeting held March 30-March 31, 2020, weeks into a global pandemic that would claim nearly 7 million lives and last more than 3 years. The scientific goals of the meeting were to: 1) Build stronger international collaborations, 2) Encourage best practices related to cutting-edge technologies, and 3) Create synergies across atlas-building consortia. Several scientists from the Human BioMolecular Atlas Program (HuBMAP), an international consortium focused on creating a multiscale atlas of the human body at single cell resolution1,2, were in attendance. In one breakout session, members from 16 international consortia joined forces to author tables detailing the anatomical structures, cell types, and biomarkers of diverse human tissues3. Dr. Katy Börner and her team would go on to create a powerful visualization tool for querying and comparing datasets in support of a Human Reference Atlas .
In another breakout session, scientists united around a shared purpose: increasing the reproducibility and rigor of multiplexed antibody-based imaging studies. The energy was electric and, at times, therapeutic as researchers shared their struggles with validating antibodies, building panels, and benchmarking spatial biology methods. It was evident to everyone on the call that we had something. Nearing the close of the breakout session, Dr. Neil Kelleher announced, “Rita Strack, the editor of Nature Methods, is giving us the greenlight to write a review!” Months of collaborative work, all virtual, resulted in a primer outlining guidelines and resources for the adoption of multiplexed antibody-based imaging methods4.
None of us is as smart as all of us. ~Ken Blanchard
Since the publication of our Nature Methods Perspective, we have connected to overcome the time, cost, and expertise required to construct highly multiplexed imaging panels. We achieved this goal by creating Organ Mapping Antibody Panels (OMAPs): combinations of antibodies that define cell populations and anatomical structures reproducibly in diverse tissues of healthy human origin5. Each antibody included in an OMAP is associated with an Antibody Validation Report (AVR), providing an overview of the antibodies validated for each protein marker, details on the characterization process, and representative images. In the HuBMAP Nature Collection, OMAPs were used to collect spatial data for the human skin6 and intestine7, furthering our understanding of tissue structure and organization.
From the beginning, collaboration has been foundational to our success. The Affinity Reagent Imaging and Validation Working Group (Figures 1-3) formed a strategic alliance with the ASCT+B Working Group and partnered with the HuBMAP Integration, Visualization and Engagement (HIVE) Infrastructure and Engagement Component (HIVE IEC) team to standardize metadata reporting for ingest and display of imaging data. We leveraged long-standing relationships with commercial partners and welcomed new members who brought their creativity, passion, and interdisciplinary talents to the team. Dr. Diane Saunders immersed herself in the data standards initially established by Dr. Andrea Radtke and Ellen Quardokus and lent her artistic gifts and Excel prowess to the display items included in the manuscript. Dr. Elizabeth (Liz) McDonough’s passion for antibody validation and multiplexed imaging shines in her leadership of the AVR effort with Ellen Quardokus and the HIVE IEC team. Dr. Michael Caldwell coordinated communications, scheduled meetings, and handled logistics in the critical role of Project Manager previously held by Dr. Jeannie Camarillo of the Kelleher group. Dr. Christopher Werlein, a dedicated pathologist, devoted his time and expertise to reviewing OMAPs for their accuracy and completeness in anatomical structure and cell phenotype assignments. Christopher Werlein faithfully tuned into meetings at 8 pm local time (Hannover, Germany) and was soon joined by the indefatigable Dr. Werner Müller (Cologne, Germany), who like Dr. Anne Wiblin (Cambridge, UK), established a perfect attendance record. Nine time zones away, Dr. Michael Snyder (Stanford, CA) regularly provided senior leadership experience, encouragement, and domain expertise related to atlas construction.
Think big, act bold, start simple, and iterate fast. ~John R. Patrick
The operational heart of our working group is steady progress through teamwork and iteration. In the beginning, there was a vision and a template but no plan for publishing our work. Ellen Quardokus was instrumental in creating the infrastructure for publishing OMAPs and AVRs. We are grateful for the wise counsel of senior leaders, particularly Dr. Neil Kelleher who advised us to, “Start with one OMAP. Get it solid. Then expand to more OMAPs.” Our first OMAP was an IBEX panel for the human lymph node authored by Dr. Andrea Radtke based on tissue-specific panels described in earlier work8. Upon expanding to include other multiplexed imaging methods, we realized the need for living, versioned documents to allow for method specific amendments. The evolving suite of ASCT+B tables, a parallel effort within HuBMAP, provided an excellent roadmap for expanding our offerings and coordinating publication with Human Reference Atlas releases. Within one year, we published 6 additional OMAPs, updated our templates and support documents, and included detailed reports for each antibody included in an OMAP. Additional information on the 7 inaugural OMAPs are found in our Nature Methods article included in the HuBMAP Nature paper package. In June 2023, we celebrated the completion of 6 additional OMAPs for a total of 13 OMAPs with hundreds of AVRs to be featured in a redesigned site in 2023.
Alone we can do so little. Together we can do so much. ~Helen Keller
To quote Dr. Katy Börner, “Cells live their lives in 3D spatial neighborhoods. Good working relationships with neighboring cells are key to delivering a well-defined function, together. To build an atlas of the healthy human, we need to understand the multi-scale structure and function of cellular neighborhoods in health, during aging, and when disease strikes.”
The parallels between healthy tissues and optimally functioning teams are many but the lesson is simple: We, like the cells we study, achieve more together!
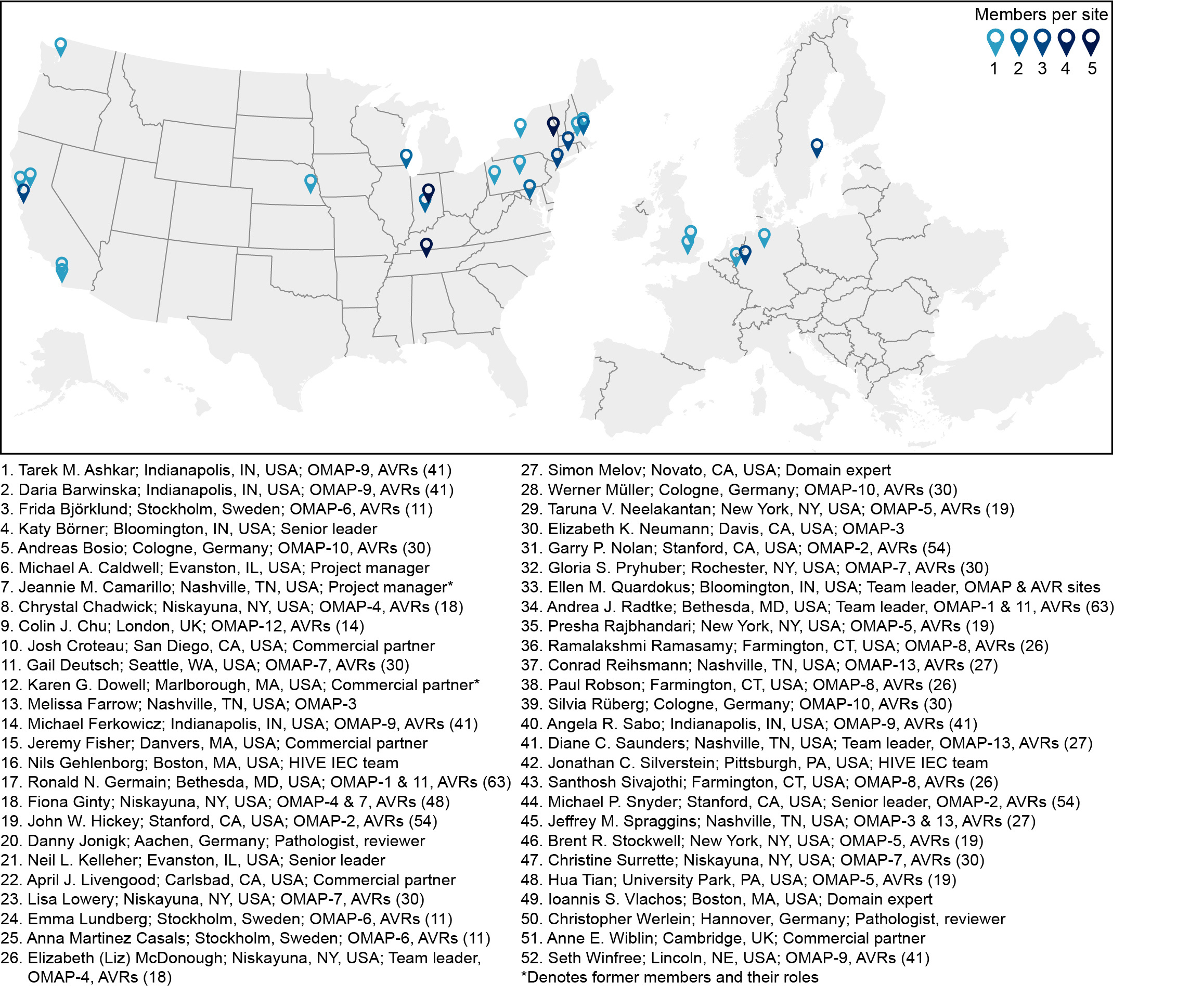
Figure 1. HuBMAP gets the job done across nine time zones. The number of members per geographical location is depicted with color shading of symbols as indicated. Members, their locations, and some of their contributions are described. Listed individuals are either authors on our Nature Methods article or contributed an OMAP and AVRs. See Figures 2 and 3 for additional working group members.
.jpg)
.jpg)
Acknowledgments
Banner image created by artist Norman Luo. Helpful suggestions and edits from Dr. Katy Börner.
References
1 . Snyder, M. P. et al. The human body at cellular resolution: the NIH Human Biomolecular Atlas Program. Nature 574, 187-192, doi:10.1038/s41586-019-1629-x (2019).
2. Jain, S., et al. Advances and prospects for the Human BioMolecular Atlas Program (HuBMAP). Nat. Cell Biol., doi: 10.1038/s41556-023-01194-w (2023).
3. Börner, K. et al. Anatomical structures, cell types and biomarkers of the Human Reference Atlas. Nat. Cell Biol., 23, 1117-1128, doi:10.1038/s41556-021-00788-6 (2021).
4. Hickey, J. W. et al. Spatial mapping of protein composition and tissue organization: a primer for multiplexed antibody-based imaging. Nat. Methods, doi:10.1038/s41592-021-01316-y (2021).
5. Quardokus, E.M. et al. Organ Mapping Antibody Panels: a community resource for standardized multiplexed tissue imaging. Nat. Methods, doi: 10.1038/s41592-023-01846-7 (2023).
6. Ghose, S. et al. 3D reconstruction of skin and spatial mapping of immune cell density, vascular distance and effects of sun exposure and aging. Commun. Biol. 6, 718, doi: 10.1038/s42003-023-04991-z (2023).
7. Hickey, J.W. et al. Organization of the human intestine at single-cell resolution. Nature 619, 572–584, doi: 10.1038/s41586-023-05915-x (2023).
8. Radtke, A. J. et al. IBEX: an iterative immunolabeling and chemical bleaching method for high-content imaging of diverse tissues. Nat. Protoc., doi:10.1038/s41596-021-00644-9 (2022).
Follow the Topic
-
Nature Methods
This journal is a forum for the publication of novel methods and significant improvements to tried-and-tested basic research techniques in the life sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Methods development in Cryo-ET and in situ structural determination
Publishing Model: Hybrid
Deadline: Jul 28, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in