Identification of DHX40 as a candidate susceptibility gene for colorectal and hematological neoplasia
Published in Cancer
Hereditary susceptibility to colorectal cancer has been recognized for at least a century [1], whereas for hematological neoplasia, inherited molecular determinants have been observed only recently [2]. Familial colorectal cancer type X (FCCTX) refers to families that fulfil the diagnostic criteria for Lynch syndrome but lack the hallmark molecular features of the disease [1]. While the genetic etiology of FCCTX is mostly unknown, the inactivation of RPS20 (ribosomal protein S20) was recently linked to FCCTX predisposition [3] and to Diamond Blackfan anemia [4], connecting ribosomopathies to colorectal and hematological carcinogenesis. The recent recognition of inherited variants in DDX41, a DEAD box RNA helicase gene, causing susceptibility to myelodysplastic syndrome and myeloid leukemias [2] provides further proof of dysregulation of ribosome biosynthesis and RNA processing in neoplasia.
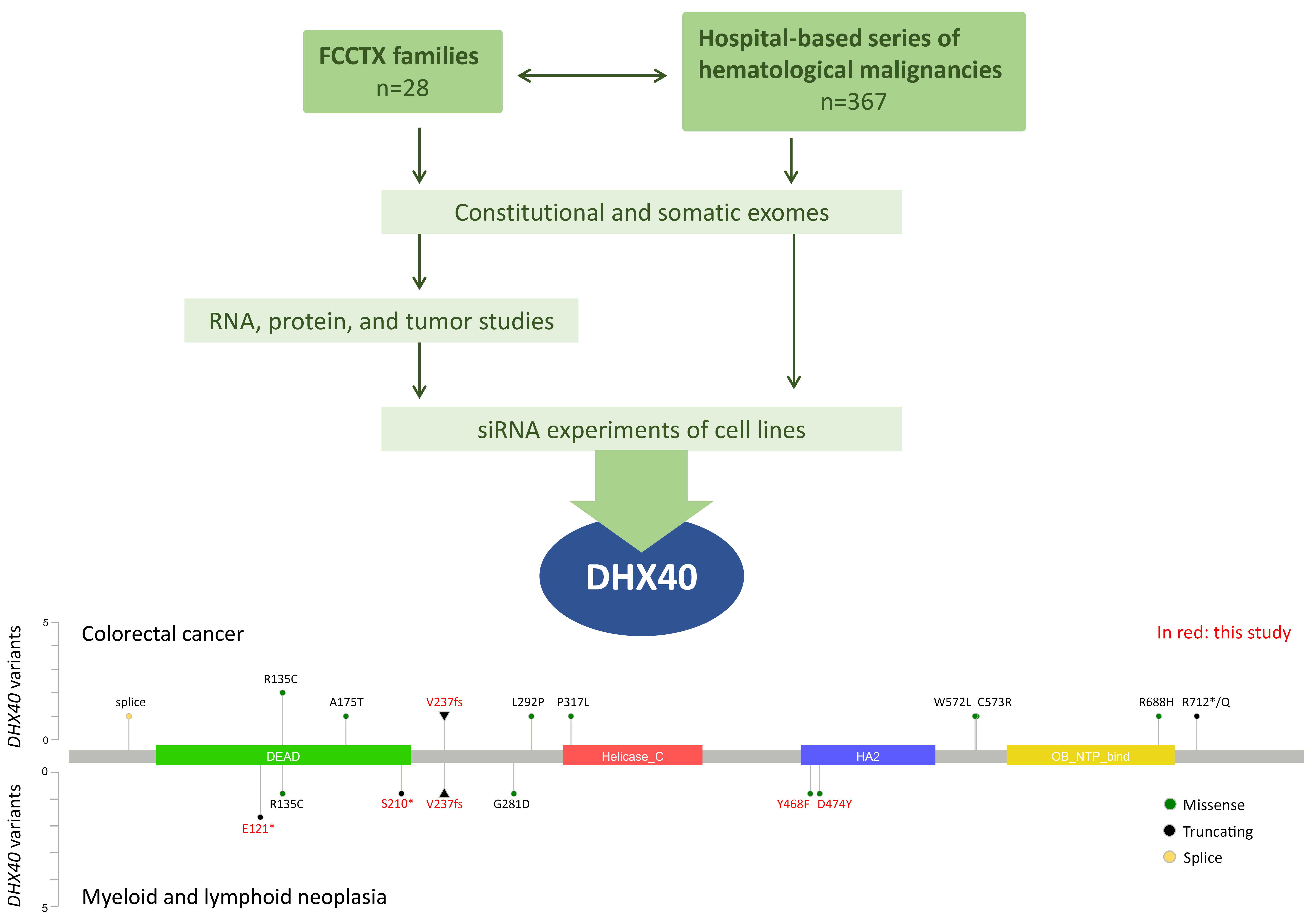
We conducted exome sequencing on 28 unexplained FCCTX families to identify their predisposing genes (Figure 1). A family with myelodysplastic syndrome/acute leukemia and colorectal cancer caught our attention. After thorough variant analysis, a heterozygous truncating variant affecting the DEAH-box RNA helicase gene DHX40 was pinpointed, showing excellent segregation with neoplasia. The variant is predicted to lead to the deletion of the helicase domain and all downstream C-terminal portion of DHX40. Analyses on healthy and tumor tissues supported the idea that loss of function of the constitutionally mutant allele alone was sufficient for tumorigenesis, without a second hit (haploinsufficiency). Knock-down of DHX40 in cell lines by siRNA, followed by RNA-sequencing, revealed enrichment of genes functioning in RNA metabolism among differentially expressed transcripts shared by all cell lines. Moreover, DHX40-siRNA treatment was associated with significant elevation of novel splice events, especially exon skipping. Our findings of haploinsufficiency and aberrant splicing as characteristic features of DHX40 inactivation resemble those previously reported for DDX41 [2, 5].
Next, we analyzed exome sequencing data from ~400 patients with hematological malignancies for possibly pathogenic variants in genes from the DEAD/H box family (Figure 1). Sixty-six individuals of 367 (18%) carried constitutional variants fulfilling our selection criteria. None of the variants affected DHX40. Somatic variants in the DEAD/H box family genes meeting our selection criteria were observed in 122 bone marrow samples out of 432 (28%), including DHX40 variants in four (two with AML and two with myeloma). The bottom part of Figure 1 summarizes all DHX40 variants of suspected pathogenicity reported to date in individuals with colorectal cancer or hematological neoplasia, highlighting in red those we discovered.
Of the almost 60 genes belonging to the DEAD/H box family genes, surprisingly many were affected by either constitutional or somatic variants in our hospital-based hematological series. Our hematological series was unselected for features of familial cancer, and no samples for co-segregation studies were available. While several DEAD/H box family genes pinpointed in this series showed significant association to hematological and colorectal neoplasia according to public databases, our observations are tentative, and the true significance of the findings remains to be determined by future studies.
In summary, we describe a rare truncating germline variant of DHX40 in a multi-generation family with hematological and solid malignancies. Moreover, our findings from a large hospital-based patient series suggest the involvement of the broader DEAD/H box gene family in hematological malignancies. Our findings add DHX40 as a new candidate to the growing list of RNA metabolism-related genes that may underlie predisposition to FCCTX [3, 6] and myeloid disorders [2].
REFERENCES
- Olkinuora AP, Peltomäki PT, Aaltonen LA, Rajamäki K. From APC to the genetics of hereditary and familial colon cancer syndromes. Hum Mol Genet. 2021;30:R206-r224.
- Polprasert C, Schulze I, Sekeres MA, Makishima H, Przychodzen B, Hosono N, et al. Inherited and Somatic Defects in DDX41 in Myeloid Neoplasms. Cancer Cell. 2015;27:658-670.
- Nieminen TT, O’Donohue M-F, Wu Y, Lohi H, Scherer SW, Paterson AD, et al. Germline Mutation of RPS20 , Encoding a Ribosomal Protein, Causes Predisposition to Hereditary Nonpolyposis Colorectal Carcinoma Without DNA Mismatch Repair Deficiency. Gastroenterology. 2014;147:595-598.e595.
- Bhar S, Zhou F, Reineke LC, Morris DK, Khincha PP, Giri N, et al. Expansion of germline RPS20 mutation phenotype to include Diamond-Blackfan anemia. Hum Mutat. 2020;41:1918-1930.
- Chlon TM, Stepanchick E, Hershberger CE, Daniels NJ, Hueneman KM, Kuenzi Davis A, et al. Germline DDX41 mutations cause ineffective hematopoiesis and myelodysplasia. Cell Stem Cell. 2021;28:1966-1981.e1966.
- Bellido F, Sowada N, Mur P, Lázaro C, Pons T, Valdés-Mas R, et al. Association Between Germline Mutations in BRF1, a Subunit of the RNA Polymerase III Transcription Complex, and Hereditary Colorectal Cancer. Gastroenterology. 2018;154:181-194.e120.
Follow the Topic
-
Leukemia
This journal publishes high quality, peer reviewed research that covers all aspects of the research and treatment of leukemia and allied diseases. Topics of interest include oncogenes, growth factors, stem cells, leukemia genomics, cell cycle, signal transduction and molecular targets for therapy.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in