IMPROVING THE EFFICIENCY OF CRISPR/CAS SYSTEM, CCExo; a story how coiled-coil supported improvement of genome editing tool was born
Published in Bioengineering & Biotechnology
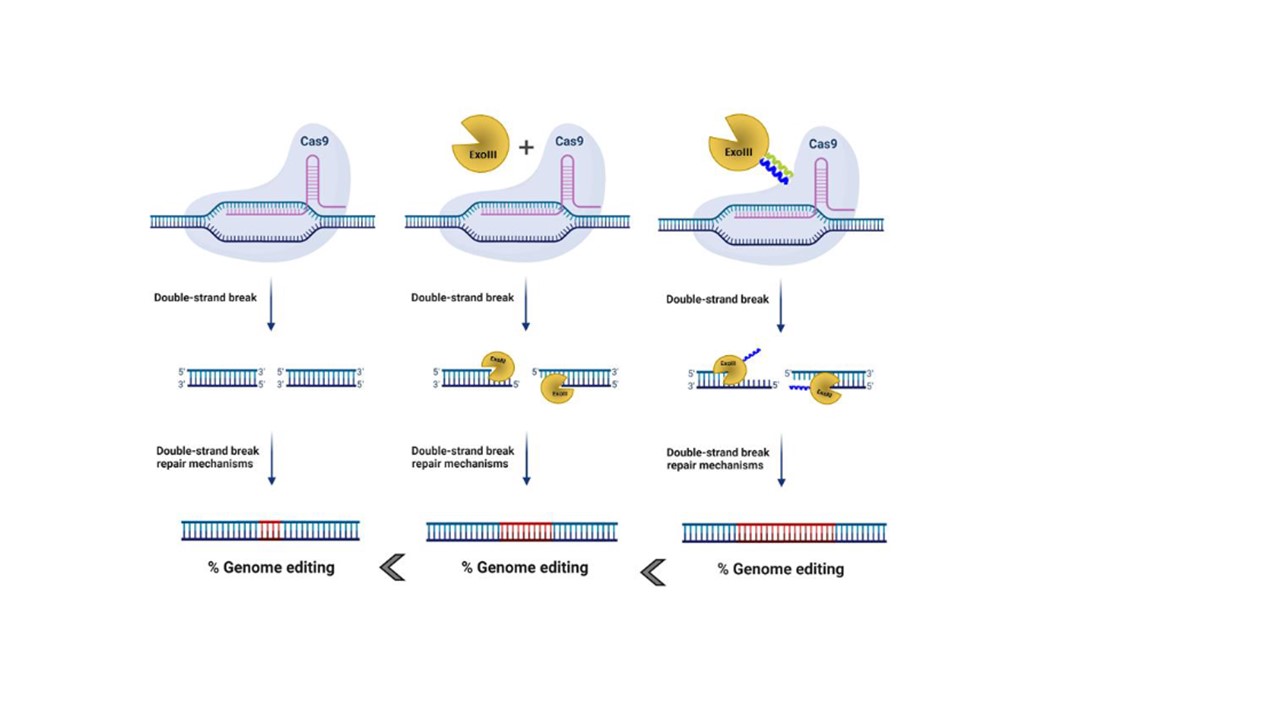
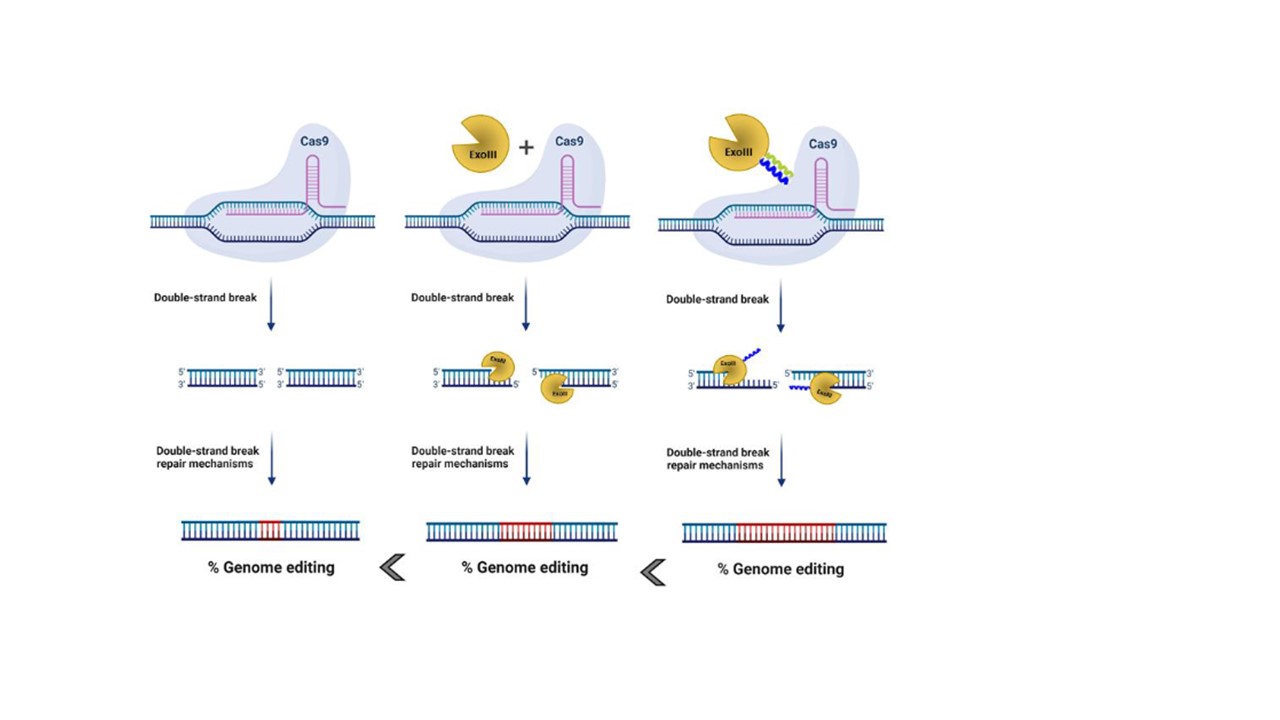 When CRISPR/Cas system first emerged as a powerful tool that enables genome editing in human cells1 I did not realize that this will become the major topic my “scientific life”. My first encounter with modifications of CRISPR was connected to the delivery of CRISPR/Cas system via extracellular vesicles2. In the meantime, we played around with ideas how to improve the efficiency of this already very capable genome editing tool. My supervisor prof. Roman Jerala gave me an idea to explore use of exonucleases that are one of the key players in rearranging genomic integrity to increase the effect of genome modification with CRISPR/Cas system. We chose several 5'→3' or 3'→5' exonuclease to test the idea if co-expression of CRISPR components and exonuclease can augment the mutation rate of CRISPR/Cas system. By first experiments on eGFP reporter cell lines we discovered that overexpression of certain exonuclease, of which ExoIII from E.coli worked best, led to increased desired gene inactivation. As true synthetic biologists that we are, we constructed several fusion proteins, where Cas9 protein and ExoIII enzyme were covalently bound. By our surprise the indel detection did not surpass the co-expression and only the N-termini fusion via 20 amino-acid linker worked. To put it mildly we did not want to be satisfied with those results as there were already some reports where co-expression or fusion proteins of exonucleases and genome editing proteins were published3,4. Then, we conjured a new approach of recruitment of exonucleases to the Cas9 protein. As our lab has substantial knowledge5 in designing heterodimeric peptide pairs that form coiled-coils (CC) based on an interplay of designable electrostatic and hydrophobic interaction that we have already used as building blocks to design CRISPR-based transcriptional regulators6, we decided to use CCs to tether the exonucleases to Cas9 in order to bring them into the close proximity of DNA double-stranded breaks.
When CRISPR/Cas system first emerged as a powerful tool that enables genome editing in human cells1 I did not realize that this will become the major topic my “scientific life”. My first encounter with modifications of CRISPR was connected to the delivery of CRISPR/Cas system via extracellular vesicles2. In the meantime, we played around with ideas how to improve the efficiency of this already very capable genome editing tool. My supervisor prof. Roman Jerala gave me an idea to explore use of exonucleases that are one of the key players in rearranging genomic integrity to increase the effect of genome modification with CRISPR/Cas system. We chose several 5'→3' or 3'→5' exonuclease to test the idea if co-expression of CRISPR components and exonuclease can augment the mutation rate of CRISPR/Cas system. By first experiments on eGFP reporter cell lines we discovered that overexpression of certain exonuclease, of which ExoIII from E.coli worked best, led to increased desired gene inactivation. As true synthetic biologists that we are, we constructed several fusion proteins, where Cas9 protein and ExoIII enzyme were covalently bound. By our surprise the indel detection did not surpass the co-expression and only the N-termini fusion via 20 amino-acid linker worked. To put it mildly we did not want to be satisfied with those results as there were already some reports where co-expression or fusion proteins of exonucleases and genome editing proteins were published3,4. Then, we conjured a new approach of recruitment of exonucleases to the Cas9 protein. As our lab has substantial knowledge5 in designing heterodimeric peptide pairs that form coiled-coils (CC) based on an interplay of designable electrostatic and hydrophobic interaction that we have already used as building blocks to design CRISPR-based transcriptional regulators6, we decided to use CCs to tether the exonucleases to Cas9 in order to bring them into the close proximity of DNA double-stranded breaks.
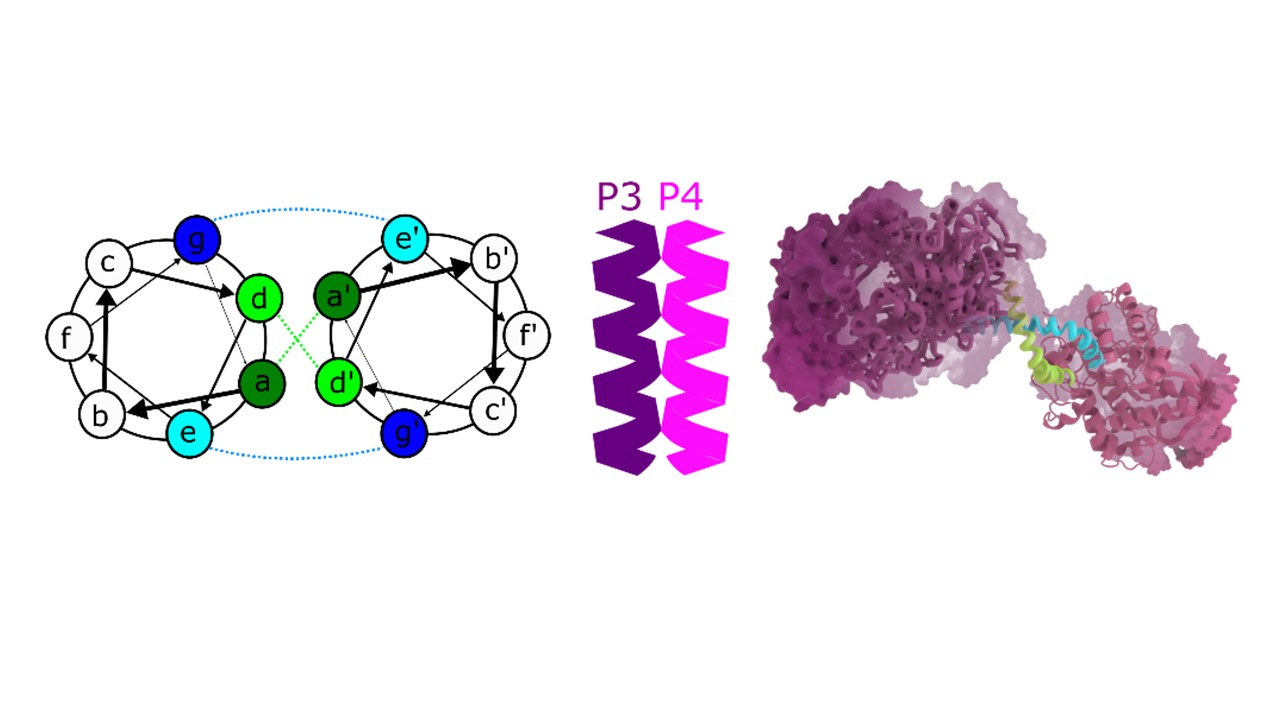
Already first experiments on HEK293 cells revealed to our delight that CCExo (CRISPR-Coiled-coil--Exonuclease) approach significantly increased genome modification on several tested genes and for all tested exonucleases, where again ExoIII showed best efficiency compared to conventional CRISPR system, co-expression or fusion proteins. Based on different sequences of CC forming peptide pairs, CC heterodimers with tunable binding affinities can be produced. We wondered if switching of CC with various binding affinity could reflect in the efficiency of genome editing.
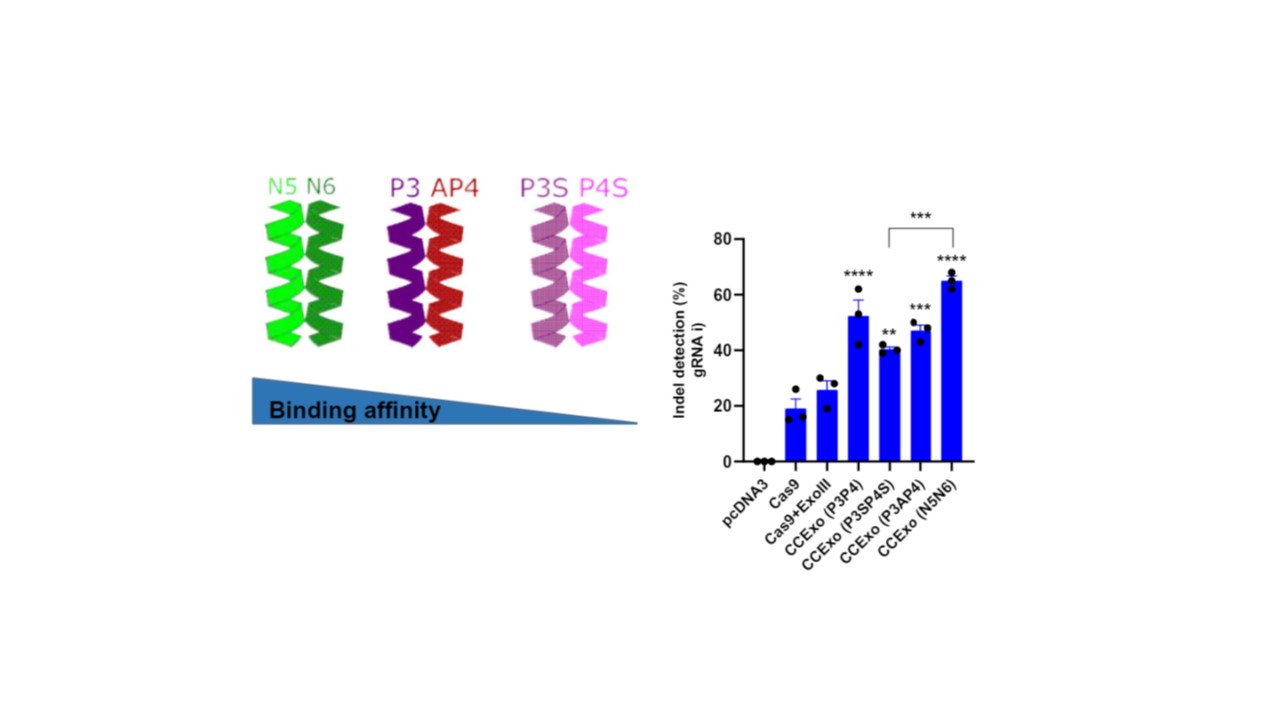
Indeed, we discovered that CCs with high binding affinity we also produced highest percentage of indel mutations and also by using high binding affinity CC pair N5:N6 we were able to obtain genome editing for gRNAs that lacked activity in the conventional CRISPR/Cas system. This was also showed for several gRNAs targeting different genes in several cell lines. We did not stop at S.pyogenes Cas9. We also showed that CC-mediated tethering of ExoIII can augment the activity of Cas12a system. By comprehensive study on different genes we discovered that not only the percentage of indel mutations was increased but also that the length of indels are substantially increased when using CCExo thus offering the possibilities to more reliably achieve knock-outs at higher degrees.
As we wanted to investigate the translational potential of this technology we wondered if somatic cells in the living organism can be modified to greater extent using CCExo. We therefore hydrodynamically delivered CCExo coding plasmids into the liver of mice that resulted again in higher genome modification compared to basic CRISPR system.
As we have been quite with plasmid-based approach, the next logical step was producing recombinant proteins to form RNPs (after we showed the lack of toxicity in several cells and in animals). Luckily for us, the production and isolation of Cas9-N5 and N6-ExoIII went smoothly. As we had RNPs in our hands, we wanted to determine the mechanism of CCExo that also provided some new insights and properties of Cas9. By doing countless of in vitro digestion assays we discovered that CCExo prevents scarless re-ligation of the Cas9 cleaved DNA template. Cas9/gRNA/DNA complex has been known to be very stable with slow dissociation even after DNA cleavage. By increasing the concentration of N6-EXOIII we tested an interesting hypothesis that tethering of N6-ExoIII to Cas9-N5 influences faster Cas9 dissociation from the DNA template, thus increasing Cas9 processivity. This may be the reason for increased efficiency of CCexo.
As RNPs can be used to genetically modify therapeutically relevant cells we engineered human CD3 cells by disrupting PD1 gene thus providing another use of CCExo, which is particularly relevant for the CAR-T cell immunotherapy, and this is the target of several ongoing clinical trials.
Thanks to the comments of the reviewers we introduced CIRCLE-seq method and bioinformatics analysis at a higher level in our laboratory. By determination of the potential off-target sites on at the genome-wide scale we concluded that despite substantial increase in genome editing we did not observe increased off-target activities.
Finally, we wanted to investigate another therapeutically relevant CCExo approach. As a model for cancer-specific genomic signature we chose well-known BCR-ABL fusion resulting from the genome translocation that is responsible for the development of chronic myelogeneous leukemia (CML). With SCID mice-based xenograft model we showed that CCExo can be used to treat cancer better than the conventional CRISPR/Cas system.
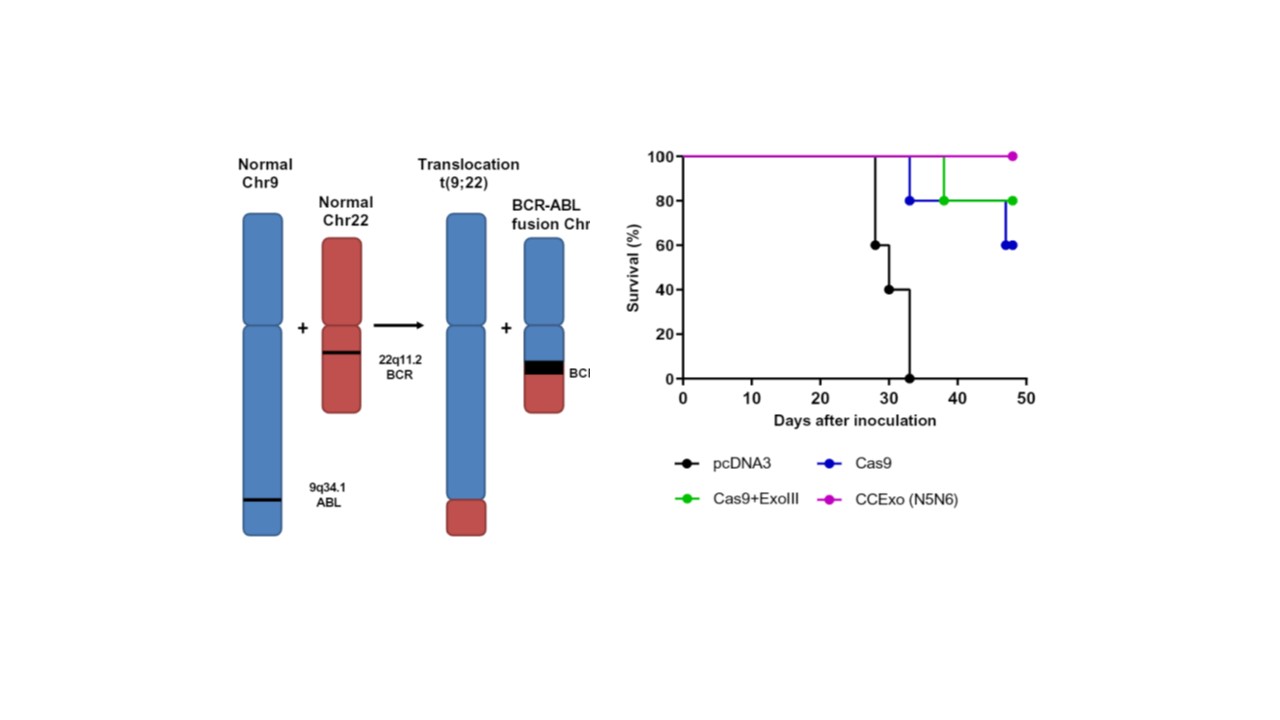 We established a fruitful collaboration with clinicians at the Department of Hematology of the University Medical Centre that provided us CML patient cells on which CCExo anti-cancer properties was determined. IT was very satisfying to observe that CCExo RNP electroporation in CML cancer cells resulted in augmented cell death compared to controls.
We established a fruitful collaboration with clinicians at the Department of Hematology of the University Medical Centre that provided us CML patient cells on which CCExo anti-cancer properties was determined. IT was very satisfying to observe that CCExo RNP electroporation in CML cancer cells resulted in augmented cell death compared to controls.
At the end I would like to say that we are very happy to share our results with others and that we made a small but potentially an important contribution in the genome modification technology. We are confident that the CCExo platform can be used not only for more efficient knock-outs but its variants could be used for treating many diseases with genetic etiology.
The described paper:
Lainšček, D., Forstnerič, V., Mikolič, V. et al. Coiled-coil heterodimer-based recruitment of an exonuclease to CRISPR/Cas for enhanced gene editing. Nat Commun 13, 3604 (2022). https://doi.org/10.1038/s41467-022-31386-1
References
- Jinek, M. et al. A Programmable Dual-RNA – Guided DNA Endonuclease in Adaptive Bacterial Immunity. Science (80-. ). 337, 816–822 (2012).
- Lainšček, D. et al. Delivery of an Artificial Transcription Regulator dCas9-VPR by Extracellular Vesicles for Therapeutic Gene Activation. ACS Synth. Biol. 7, 2715–2725 (2018).
- Mashimo, T. et al. Efficient gene targeting by TAL effector nucleases coinjected with exonucleases in zygotes. Sci. Rep. 3, 1–6 (2013).
- Clements, T. P., Tandon, B., Lintel, H. A., McCarty, J. H. & Wagner, D. S. RICE CRISPR: Rapidly increased cut ends by an exonuclease Cas9 fusion in zebrafish. Genesis 55, 1–6 (2017).
- Ljubetič, A. et al. Design of coiled-coil protein-origami cages that self-assemble in vitro and in vivo. Nat. Biotechnol. 35, (2017).
- Lebar, T., Lainšček, D., Merljak, E., Aupič, J. & Jerala, R. A tunable orthogonal coiled-coil interaction toolbox for engineering mammalian cells. Nat. Chem. Biol. (2020). doi:10.1038/s41589-019-0443-y
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in