In vitro reconstitution of the Deinococcus radiodurans Nucleotide Excision Repair system – a new tool to decipher the functions of the Uvr proteins.
Published in Microbiology
In this paper, we report the successful in vitro reconstitution of a bacterial Nucleotide Excision Repair (NER) pathway. NER is a repair pathway capable of removing of very diverse DNA lesions such as pyrimidine–pyrimidone (6–4) photoproducts (6-4-PP), cyclobutane pyrimidine dimers (CPD) and a wide range of helix-distorting lesions including DNA adducts. Here, we built this repair system using a doubly labeled oligonucleotide and purified UvrABC proteins from the radiation resistant bacterium Deinococcus radiodurans.
When conceiving this new repair assay, we decided to make several important changes to the system that differentiate it from earlier NER systems. First, we made use of an oligonucleotide labeled with a centered fluorescein conjugated to a thymine, a listed substrate of NER, but also an additional fluorophore on the 5’ end of the lesion-containing strand. These two labels allowed us to track the various products resulting from the incision by the Uvr proteins. Secondly, the proteins were all purified from one single organism D. radiodurans, a mesophilic bacterium. With these very stable proteins, we were able to monitor the assay on longer time courses and work on each component of our system. These two characteristics enabled us to test many conditions in order to optimize the in vitro incision assay.
One of the first difficulties that we encountered during this study was the variable activity of the purified UvrC protein. Although all three proteins (UvrA1, UvrB and UvrC) are essential to the system, the inactivity of UvrC can be directly observed as a reduced incision activity, since both endonuclease domains are carried by UvrC. While the purification of the proteins UvrA1 and UvrB were reproducible, the purification of UvrC was more delicate and the quality of the purified protein was more variable. We thus had to optimize all steps of the purification to ensure a stable, robust and reproducible activity of the endonuclease domains at the end of the purification process. After each purification, the protein was thus tested in the assay and compared to a “reference” batch that had shown activity and stability in previous experiments.
The optimization of the incision assay itself was the longest journey as we tried to isolate each parameter and study how it affected the incision efficiency of the system. Different electrophoresis protocols were tested to allow a clear visualization and quantification of each fragment resulting from the incision of the substrate. Different concentrations and ratios of the three proteins were assessed for a defined concentration of substrate. After completing this step, we monitored the incision efficiency under different temperature, buffer and salt conditions. We also tested up to seven cofactors with different ranges of concentrations to determine which ones were needed to reach the highest incision activity. These divalent ions are known to be determining actors in this enzymatic assay together with the nucleotide (ATP) that is also necessary to drive this system.
Once we set the conditions needed for an efficient system in which we could properly identify the products resulting NER incision, we used Matrix-Assisted Laser Desorption/Ionization-Time Of Flight (MALDI-TOF) to establish the sites of cleavage of the substrate by UvrABC. We were able to detect unambiguously the exact sequence of the fragment resulting from the repair along with its size corresponding to a 12 mer oligonucleotide. This step also allowed us to conclude on the sites of cleavage of two other substrates that we tested in the in vitro assay. The latter substrates were used to explore the substrate specificity of our NER system. With one substrate, we kept the fluorescein conjugated thymine as a lesion and changed the surrounding DNA sequence to analyze the potential effect of the sequence on the incision efficiency. With the second substrate, we prepared a bulkier substrate by coupling biotin conjugated thymine with streptavidin to see whether this bulkier lesion would be a better substrate for NER.
The incision assay presented in this paper will undoubtedly be a useful tool for further studies of the bacterial NER pathway. The fact that it has been fully optimized will now enable us to dissect the contribution of each component to this repair reaction. Building on this work, we thus hope to shed light on some of the complex mechanisms underlying NER in the near future.
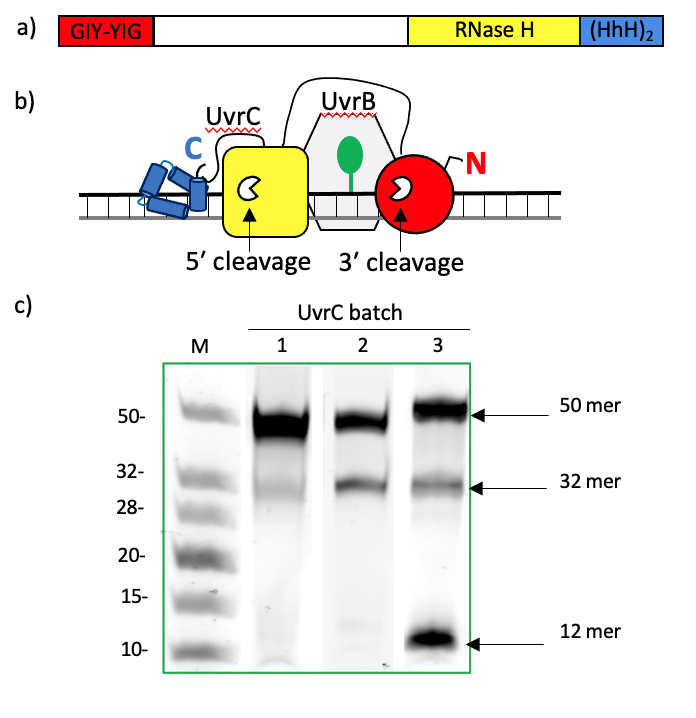
Figure 1: a) Schematic diagram of UvrC, a multidomain protein composed of an N-terminal GIY-YIG endonuclease domain (red), a C-terminal RNase H endonuclease domain (yellow) and two terminal helix-hairpin-helix (HhH) motifs (blue). b) Illustration of the assembly of UvrB (grey) and UvrC, colored as in (a), on double-stranded DNA containing a fluorescein-conjugated thymine (green), a known substrate of the NER, in the central position of the top strand. In this configuration, UvrC can incise the DNA on both the 5’ and 3’ sides of the lesion using its respective C- and N-terminal endonuclease domains. c) TBE-polyacrylamide urea gel analyses of the drUvrABC incision activity using different batches of purified drUvrC protein. Reactions were performed for 1 hour at 37°C using 25 nM F26-seq1 substrate, 1 μM drUvrA1, 0.5 μM drUvrB and 2 μM drUvrC in the presence of 10 mM Mg2+ and 4 mM ATP. Lane 1: incision by a partially cleaved drUvrC still containing its N-terminal His-tag and some residual nucleic acid contamination. Lane 2: incision by drUvrC purified on Ni-NTA resin which was visibly stripped of its nickel after elution of drUvrC. Lane 3: incision by drUvrC after optimization of the purification protocol. The gels were visualized with the green filter to detect fluorescein-labelled bands.
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
DNA repair and human disease
Publishing Model: Hybrid
Deadline: Oct 31, 2026
Cell death and inflammatory signalling
Publishing Model: Hybrid
Deadline: Oct 28, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in