Explore the Research

Origin and Evolution of RAS Membrane Targeting - Oncogene
Oncogene - Origin and Evolution of RAS Membrane Targeting
Why we started the study.
HRAS, NRAS and KRAS have been a focus of cancer biologists for more than 40 years because they are the most frequently mutated oncogenes in cancer. Around 20% of human cancers harbor mutations in these genes. The interest in RAS genes has recently intensified as attempts to directly target their gene products with drugs have finally born fruit. The three RAS genes code for four proteins: KRAS4B and KRAS4A, generated by the alternative usage of the last two coding exons of the KRAS gene, HRAS, and NRAS. Until recently, KRAS4A has been considered less relevant in cancer than KRAS4B because of its relatively low expression. However, recent findings by Philip Lab and others have shown specific molecular interactions and equal expression of KRAS4B and KRAS4A in colorectal tumours [1-4]. Importantly, KRAS4A, but not KRAS4B, has been found to directly regulate hexokinase 1 on the outer mitochondrial membrane, an interaction that uncovered a unique metabolic vulnerability in tumors expressing relatively high levels of oncogenic KRAS4A, which could have therapeutic potential [2, 5]. These observations led Mark to be interested in the evolutionarily origins of KRAS4A.
The scenario.
The products of the RAS genes, also known as classical or canonical RAS proteins, consist of two functionally distinct regions: catalytic GTP/GDP binding domains (G-domains) with a highly conserved structure that act as binary molecular switches, and an unstructured C-terminal hypervariable region (HVR) of 22-23 residues that anchor the GTPases to cellular membranes by using different combinations of membrane targeting motifs (Fig. 1). Evolutionarily, HRAS, NRAS, and KRAS genes form a branch in the tree of the RAS family of small GTPases, which in humans comprise 40 members [6, 7]. The RAS family itself is a subset of a much larger group of evolutionarily related small GTPases known as the RAS Superfamily. Although, the broad evolutionary history of the RAS superfamily has been the subject of several studies, some aspects of the evolution of the canonical RAS proteins (HRAS, NRAS, KRAS4B, and KRAS4A) have not been fully addressed, such us the diversification of the RAS genes, the origin of the HVR sequences, and the generation of the KRAS4A splice variant.
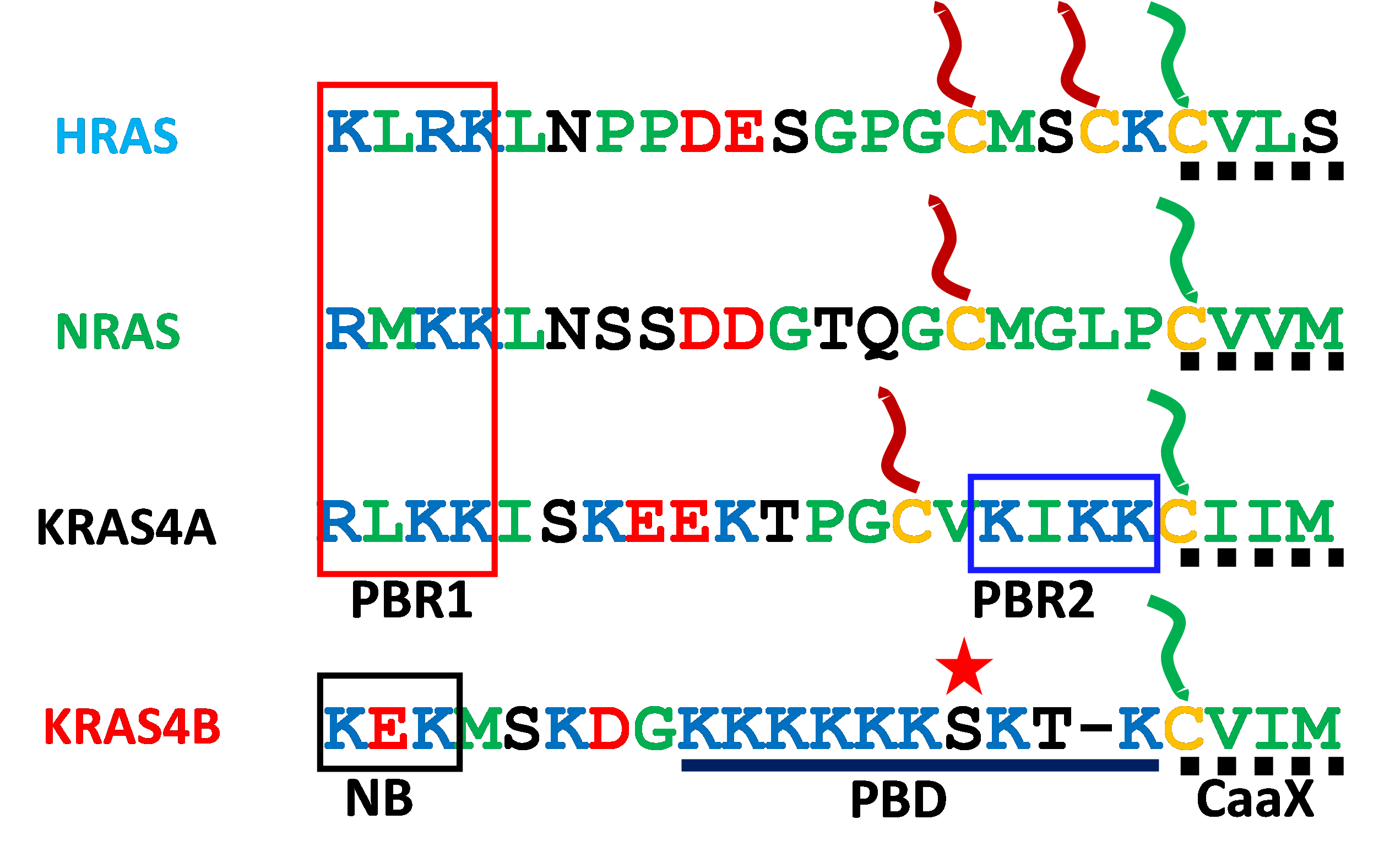
Fig. 1. Canonical RAS proteins have specific combinations of membrane targeting motifs in their hypervariable regions (HVRs). The ubiquitous C-terminal CaaX motif (dotted line) is shown with its cysteine (yellow) modified with a farnesyl lipid (green). The other HVR cysteines are shown modified with palmitate (red). The strong polybasic domain (PBD) of KRAS4B is shown with lysines in blue with an intervening serine 181 starred to indicate the phosphorylation site. Polybasic region 1 (PBR1) is shared by HRAS, NRAS and KRAS4A. Polybasic region 2 (PBR2) is unique to KRAS4A. A neutralized basic motif (NB) is boxed in KRAS4B.
The methodology
We first compiled representative RAS proteins from key major groups of eukaryotes extending over 1000 million of years (MY) of evolutionary distance, from single celled amoebozoans to mammals. When we began the study newly sequenced genomes of lower vertebrates such as jawless fishes like lampreys and hagfish were available to pinpoint the diversification of the RAS genes. We analysed the similarities and characteristics of the protein and DNA sequences, the gene structures (exon/intron patterns), and the evolutionary conservation of neighbourhood genes (synteny).
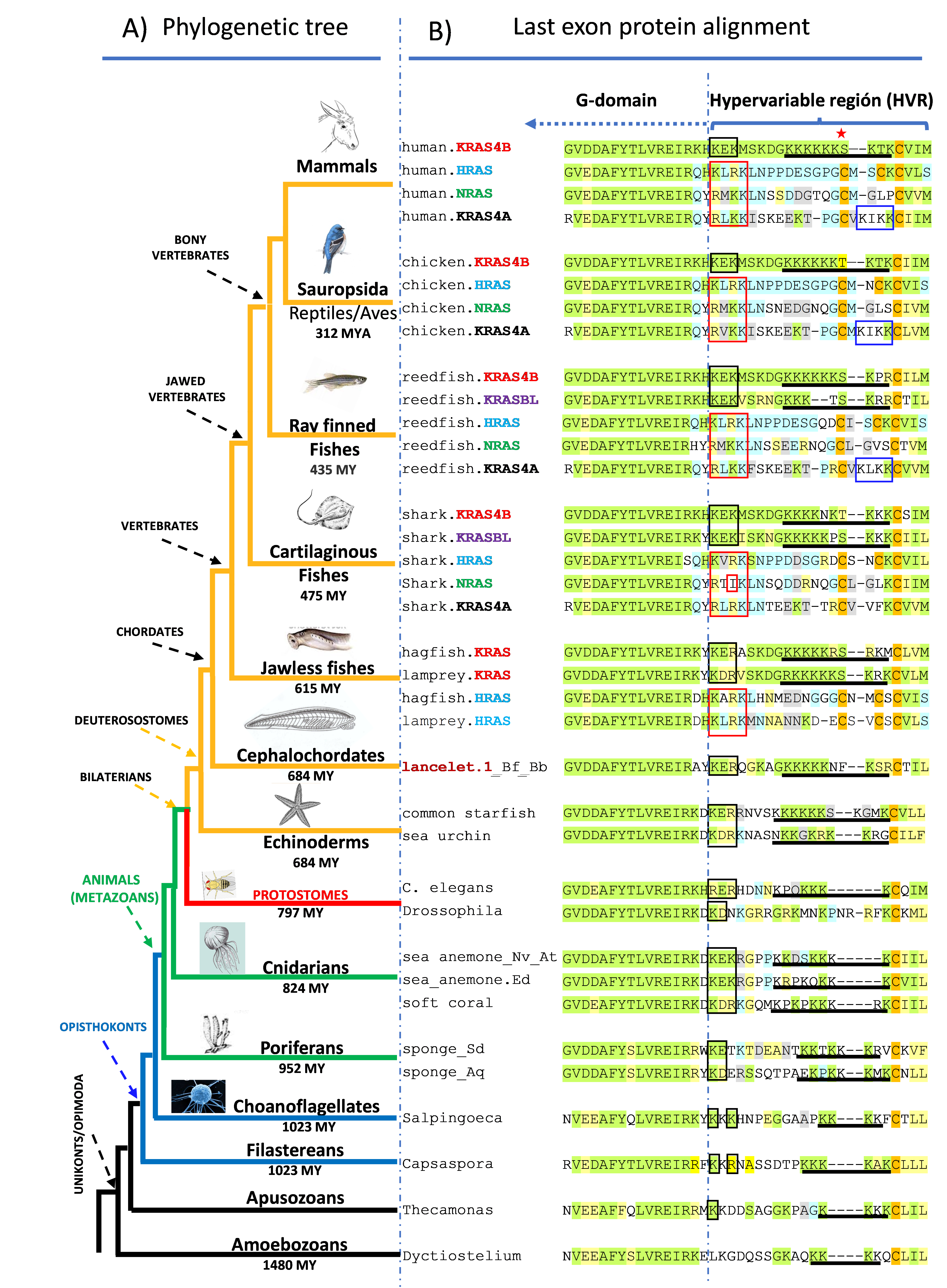
Fig. 2. Simplified phylogenetic tree of eukaryotic organisms with the protein alignment of the HVR regions (bracket). Membrane targeting motifs are depicted as in in Fig 1.
What we found.
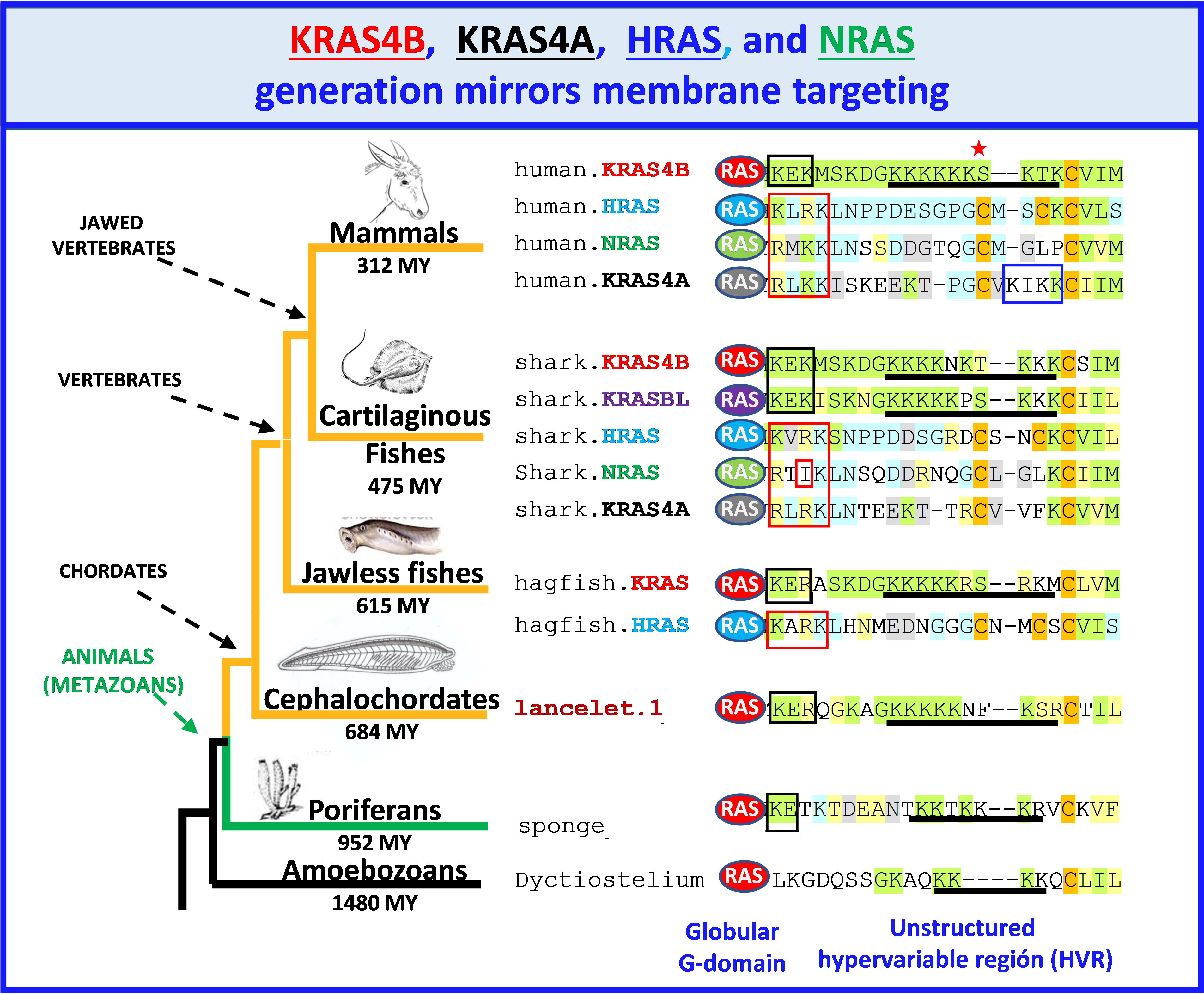
We found the presence of HRAS, NRAS, KRAS4B and KRAS4A, in all vertebrates except the most basal jawless fishes (lampreys and hagfish) that only have HRAS and KRAS4B (Fig. 2). In addition, all vertebrates except mammals, birds, and jawless fishes, have an additional, previously unrecognized RAS GTPase we have designated KRAS4B-like (KRASBL) because its high similarity with KRAS4B. In contrast to vertebrates, with very few exceptions we found only one RAS protein-related sequence in invertebrates, fungi and single celled eukaryotic organisms.
We found that KRAS is primordial and gave rise to HRAS through a gene duplication >600 million years ago (MYA) in the common ancestor of vertebrates, that NRAS is a duplication of HRAS and that KRAS4A arose 475 MYA with the evolution in the common ancestor of jawed vertebrates (cartilaginous fish and bony vertebrates) not by a whole gene duplication but rather by capture into the third intron of the KRAS locus of exon 4 of NRAS (Fig. 3)
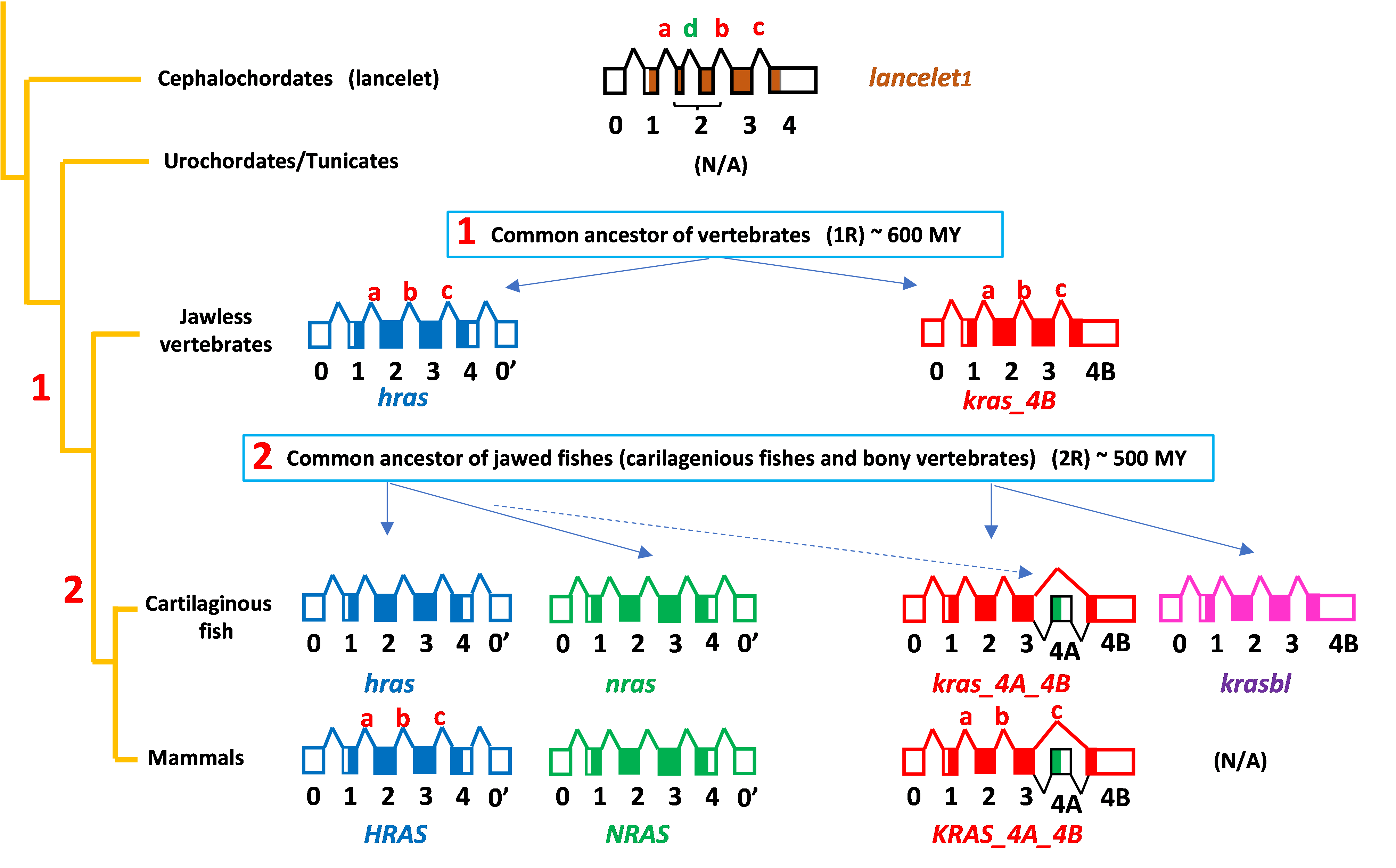
Fig. 3. Scheme of RAS protein expansion from invertebrates (cephalochordates) to mammals. Exon-intron structures are depicted with exons as boxes (1-4 coding, 0 and 0’ non-coding) and introns as lines connecting the exons (a-d).
We show that the switch 1 and 2 regions of the G-domain are identical throughout evolution, except in fungi and some single celled organisms that have an amino acid changed in Switch II. These sequences thus provide a signature for this subset of RAS superfamily gene products and suggest intense evolutionary pressure to maintain signalling down the MAPK, and perhaps other, pathways.
We found that the PBR1 membrane targeting motif found in HRAS, NRAS and KRAS4A is highly conserved and incorporates a hydrophobic residue interspersed among three basic residues (Fig. 2). This suggests a membrane insertion motif that has not been explored biophysically. We also found that the NB motif consisting of two basic residues flanking an acidic residue is conserved in all KRAS4B including invertebrates (Fig. 2). The function of the NB motif in membrane targeting has not been investigated. We found that all vertebrate KRAS4B and KRASBL sequences include a phosphate acceptor (S/T) three residues upstream of the CaaX motif, suggesting that modulation of the negative charge by phosphorylation is functionally important. Importantly, we found, with the exception of HRAS in cartilaginous fish and KRASBL, that vertebrate CaaX motifs signals for farnesylation rather than geranylgeranylation. In contrast, invertebrate RAS proteins as well as the majority of RAS superfamily small GTPases have CaaX motifs that signal for geranylgeranylation. Thus, it appears that diminishing the hydrophobicity of the prenyl lipid that modifies RAS proteins was essential for their function in vertebrates.
Relevance
Despite highly overlapping regulation and function, the persistence of four RAS isoforms with unique membrane-targeting sequences for >400 MY of evolution suggests important and distinct functions. Targeted therapies for RAS-driven cancers must take into account the differences among the RAS isoforms. Our evolutionary analysis will contribute to a more complete understanding of these differences.
References
1 Tsai FD, Lopes MS, Zhou M, Court H, Ponce O, Fiordalisi JJ et al. K-Ras4A splice variant is widely expressed in cancer and uses a hybrid membrane-targeting motif. Proc Natl Acad Sci U S A 2015; 112: 779-784.
2 Amendola CR, Mahaffey JP, Parker SJ, Ahearn IM, Chen WC, Zhou M et al. KRAS4A directly regulates hexokinase 1. Nature 2019; 576: 482-486.
3 Jing H, Zhang X, Wisner SA, Chen X, Spiegelman NA, Linder ME et al. SIRT2 and lysine fatty acylation regulate the transforming activity of K-Ras4a. Elife 2017; 6.
4 Castel P, Dharmaiah S, Sale MJ, Messing S, Rizzuto G, Cuevas-Navarro A et al. RAS interaction with Sin1 is dispensable for mTORC2 assembly and activity. Proc Natl Acad Sci U S A 2021; 118.
5 Nuevo-Tapioles C, Philips MR. The role of KRAS splice variants in cancer biology. Front Cell Dev Biol 2022; 10: 1033348
6 Colicelli J. Human RAS superfamily proteins and related GTPases. Sci STKE 2004; 2004: RE13.
7 van Dam TJ, Bos JL, Snel B. Evolution of the Ras-like small GTPases and their regulators. Small GTPases 2011; 2: 4-16
Follow the Topic
-
Oncogene
This journal aims to make substantial advances in our knowledge of processes that contribute to cancer by publishing outstanding research.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in