LAMAs: controlling proteins with small molecules
Published in Protocols & Methods

Explore the Research

Chemogenetic Control of Nanobodies - Nature Methods
Engineered nanobody allows reversible control of activity in cells through the binding of small molecules.
Biological systems are dynamic. With the advent of methods to visualize the dynamics of biological systems, there is also a need for tools that can perturb these biological systems in a dynamic manner. Let me introduce you to a new molecular tool: ligand-modulated antibody fragments, or LAMAs, which can selectively sequester and release proteins inside cells using small-molecules.
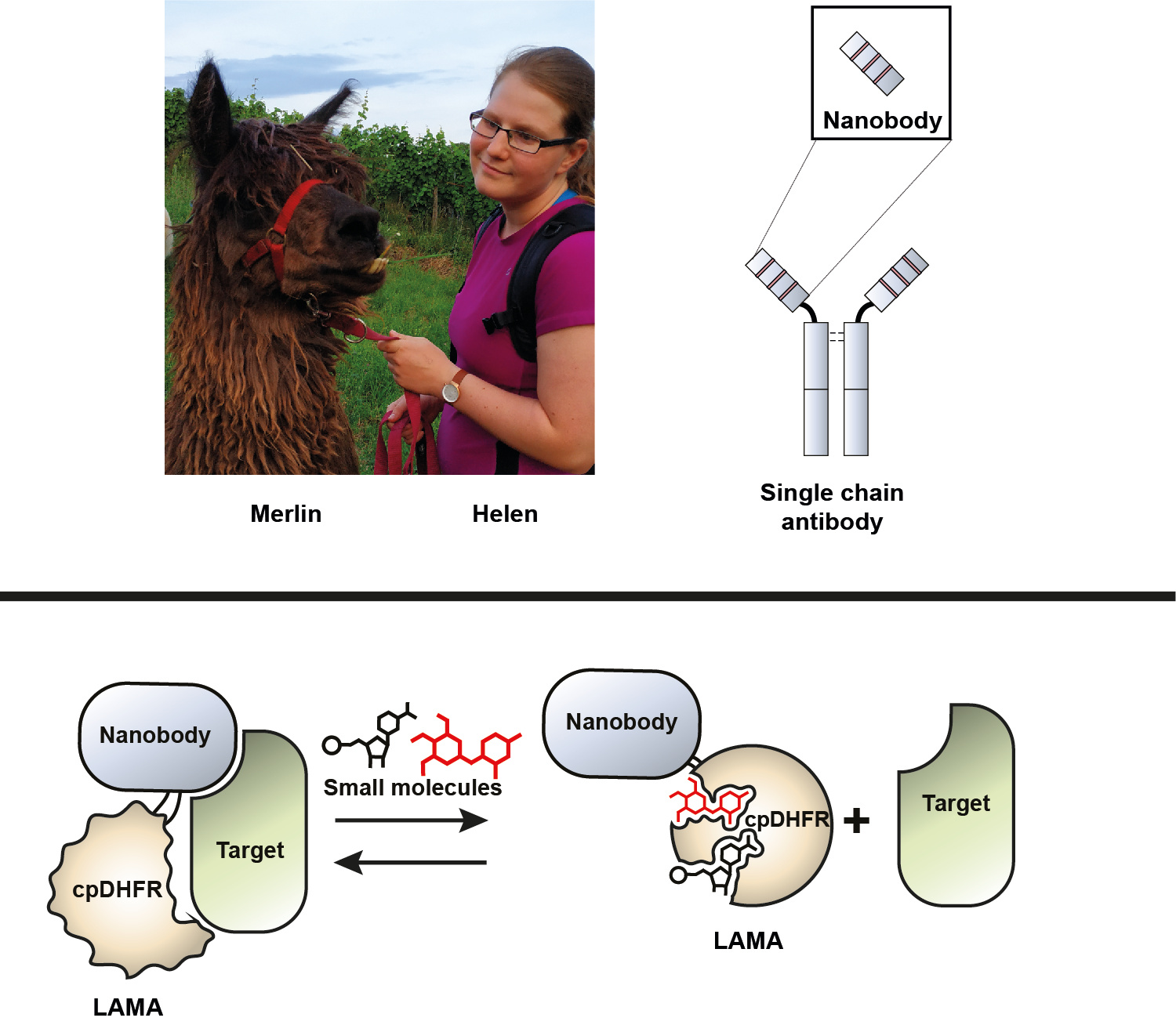
LAMAs were born when we were searching for dynamic protein switches that could be controlled on fast time scales inside of cells.
Nanobodies are derived from a part of antibodies produced by the Camelide family [1]. They provide a way to selectively and tightly bind targets, but this binding is almost irreversible. To develop protein switches dependant on an external stimuli we needed to append a “switching” domain. A circular permutated form of bacterial dihydrofolate reductase (cpDHFR) turned out to be exactly what we were looking for. We inserted cpDHFR into the nanobody, and named the resulting protein chimeras LAMAs (Figure 1). Interestingly, in contrast to the often used chemical inducers of dimerization, the affinity of LAMAs is high in the absence, but decreases on the addition of small-molecules that bind DHFR.
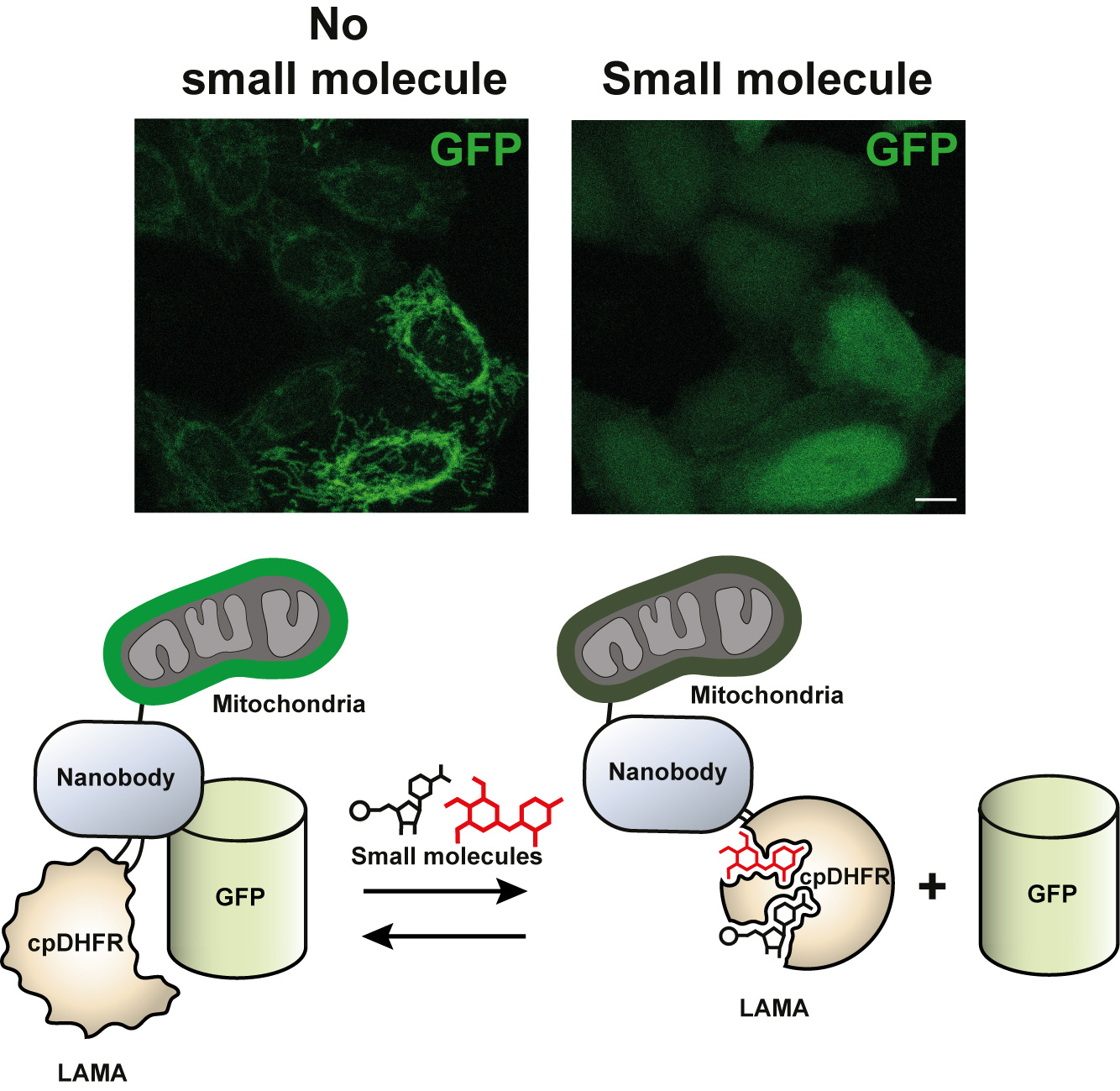
We first focused out attention on turning a nanobody against GFP [2] into a LAMA (Figure 2). GFP-fusion proteins are omnipresent in the life sciences. The Ellenberg group at EMBL for example, have used GFP-fusion proteins to track protein locations and concentrations during the cell cycle [3]. Using the LAMAs we could dynamically delocalize one such protein, and follow the outcome of disrupted mitosis. As many research groups have GFP-fusions of their favourite protein we are excited to see how the GFP-LAMAs can be used in other scenarios that benefit from dynamic control of localisation
Considering the potential of dynamically controlling systems of interest, we next asked: can all nanobodies be turned into LAMAs? The answer is currently no. Not all nanobodies allowed the insertion of cpDHFR and could still bind to the target. Nonetheless, LAMAs could be generated for a number of different nanobodies with different binding modes to their targets (Figure 3).
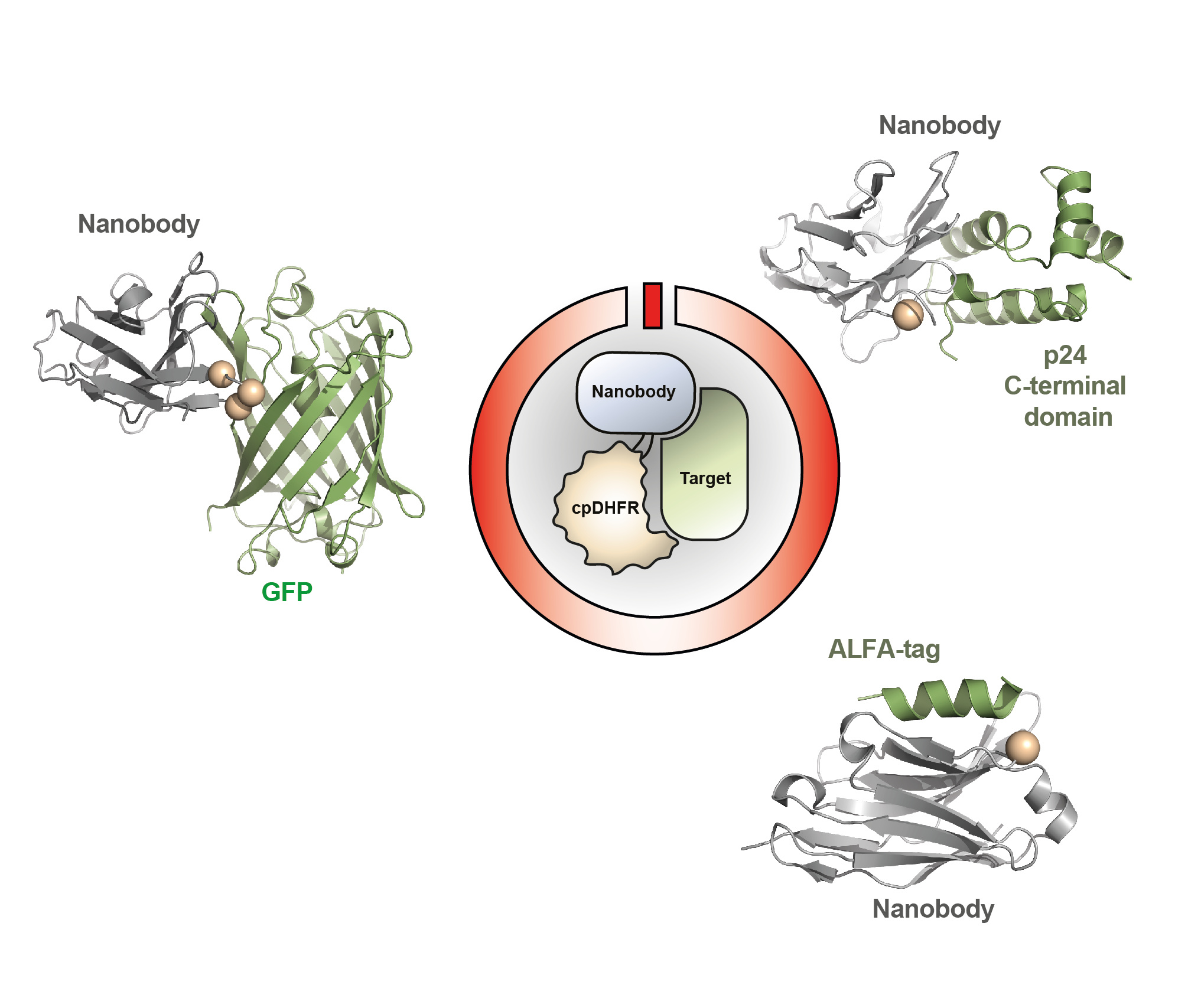
With the growing number of nanobodies made available to researchers, the future of LAMAs looks bright. Dynamic systems require dynamic molecular tools.
Read more about LAMAs in our recent publication: Chemogenetic Control of Nanobodies
1. Hamers-Casterman, C. et al. Naturally occurring antibodies devoid of light chains. Nature 363, 446-448 (1993).
2. Kirchhofer, A. et al. Modulation of protein properties in living cells using nanobodies. Nat. Struct. Mol. Biol. 17, 133-138 (2009).
3. Cai, Y. et al. Experimental and computational framework for a dynamic protein atlas of human cell division. Nature 561, 411-415 (2018).
Follow the Topic
-
Nature Methods

This journal is a forum for the publication of novel methods and significant improvements to tried-and-tested basic research techniques in the life sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Methods development in Cryo-ET and in situ structural determination
Publishing Model: Hybrid
Deadline: Jul 28, 2026
Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in