Leveraging 3-D models and immunometabolic profiling to understand how vaginal bacteria may promote cancer development
Published in Cancer

The association of specific bacteria with other malignancies are well established. For example, Helicobacter pylori is strongly associated with gastric cancer; Fusobacterium nucleatum, enteropathogenic Escherichia coli, and Bacteroides fragilis are associated with colon cancer. Numerous studies have investigated the mechanisms underlying the associations of these species with malignancy. In contrast, a paucity of research has been conducted into the pro-carcinogenic mechanisms of bacteria that have been associated with cervical cancer and other cancers of the female reproductive tract (FRT). Next-generation sequencing by 16S ribosomal profiling of women suggests that specific bacterial genera, such as Fusobacterium, Atopobium, and Sneathia, are associated with cervical cancer. We recently published a report in NPJ Biofilms and Microbiomes detailing the immmunometabolic profiling with these organisms (Laniewski et al., 2021). However, other genera, such as Peptoniphilus and Porphyromonas, have been associated with endometrial cancer and little data exists on tumorigenic mechanisms. These organisms are also associated with cervicovaginal dysbiosis and bacterial vaginosis. As such, we embarked on our study to investigate the immunometabolic mechanisms of select understudied BV-associated bacteria (BVAB: Fusobacterium gonidiaformans, F. nucleaum, Peptoniphilus lacrimalis, Porphryomonas uenonis, and Lancefieldella parvula) known to be associated with gynecologic malignancies. To achieve our goals, we coupled the use of our robust three-dimensional (3-D) human cervical cell model with an -omics approach using immunoassays and untargeted global metabolomics.
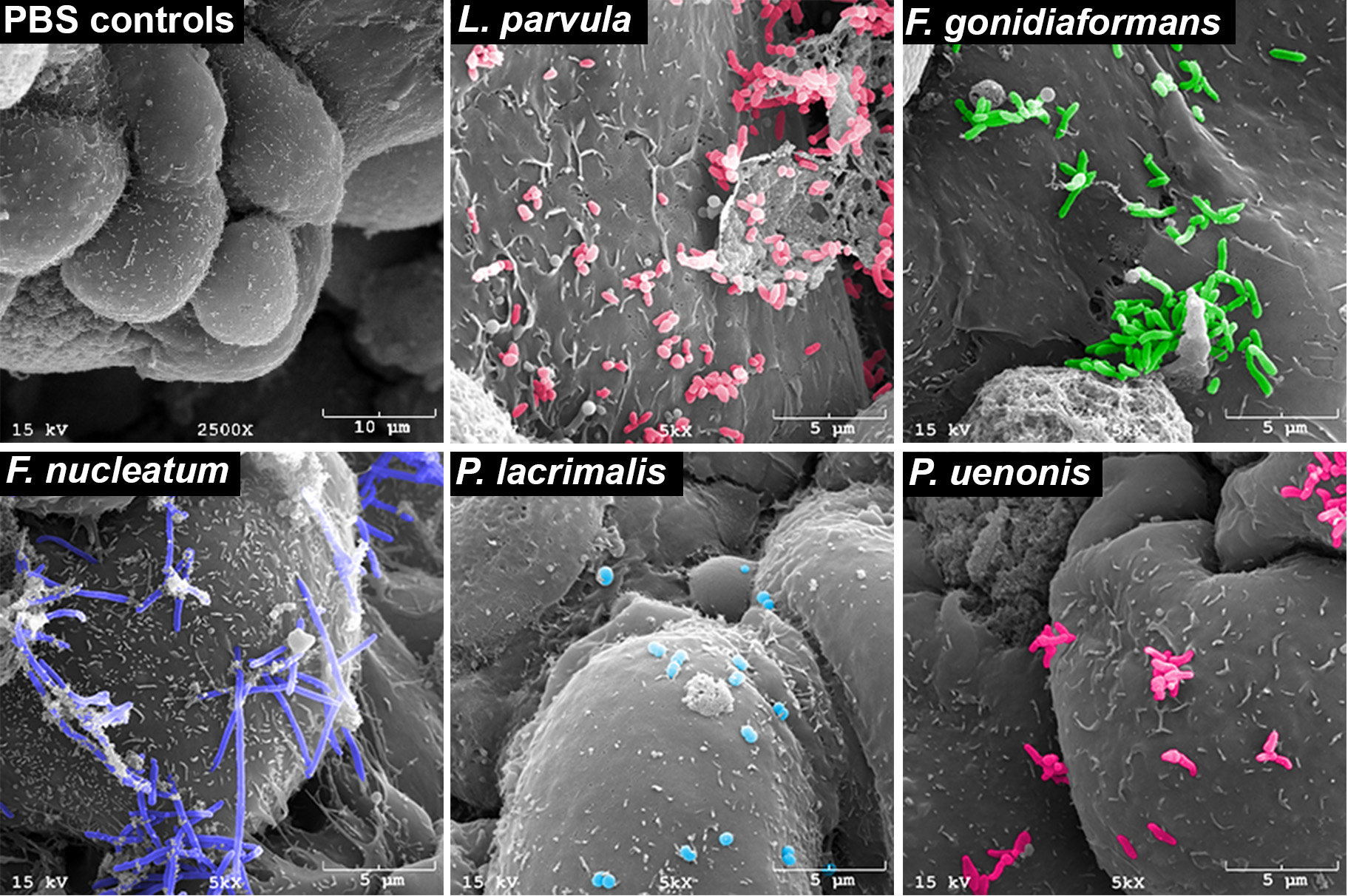
Figure 1. Scanning electron microscopy demonstrating bacterial vaginosis-associated bacteria adhering to three-dimensional human cervical cells (from Maarsingh, Laniewski, & Herbst-Kralovetz, Communications Biology, 22;5(1):725, 2022)
First, we found that infection with P. lacrimalis and L. parvula elicited a robust innate immune response from 3-D cervical cells. These species elicited secretion of pro-inflammatory immune mediators, such as interleukin (IL)-1α, IL-1β, IL-6, and IL-8. F. nucleatum and F. gonidiaformans also stimulated significant secretion of IL-1β, IL-6, and IL-8 by 3-D cervical cells, though the immune response was more moderate relative to P. lacrimalis and L. parvula. Infection with P. uenonis dampened 3-D cervical cell chemotactic responses, indicated by significant downregulation of RANTES, IP-10, and fractalkine. This suggests that P. uenonis may evade immune-mediated recruitment clearance mechanisms. Next, we measured cancer-associated protein biomarkers by 3-D cervical cells after infection with BVAB. L. parvula and F. gonidiformans significantly upregulated expression of sFasL and TRAIL, both of which have been shown to be associated with cervical neoplasms. These data suggest that L. parvula and F. gonidaformans, and possibly other BVAB, may promote a pro-carcinogenic cervicovaginal microenvironment by influencing cervical cell expression of anti-tumor immune mediators.
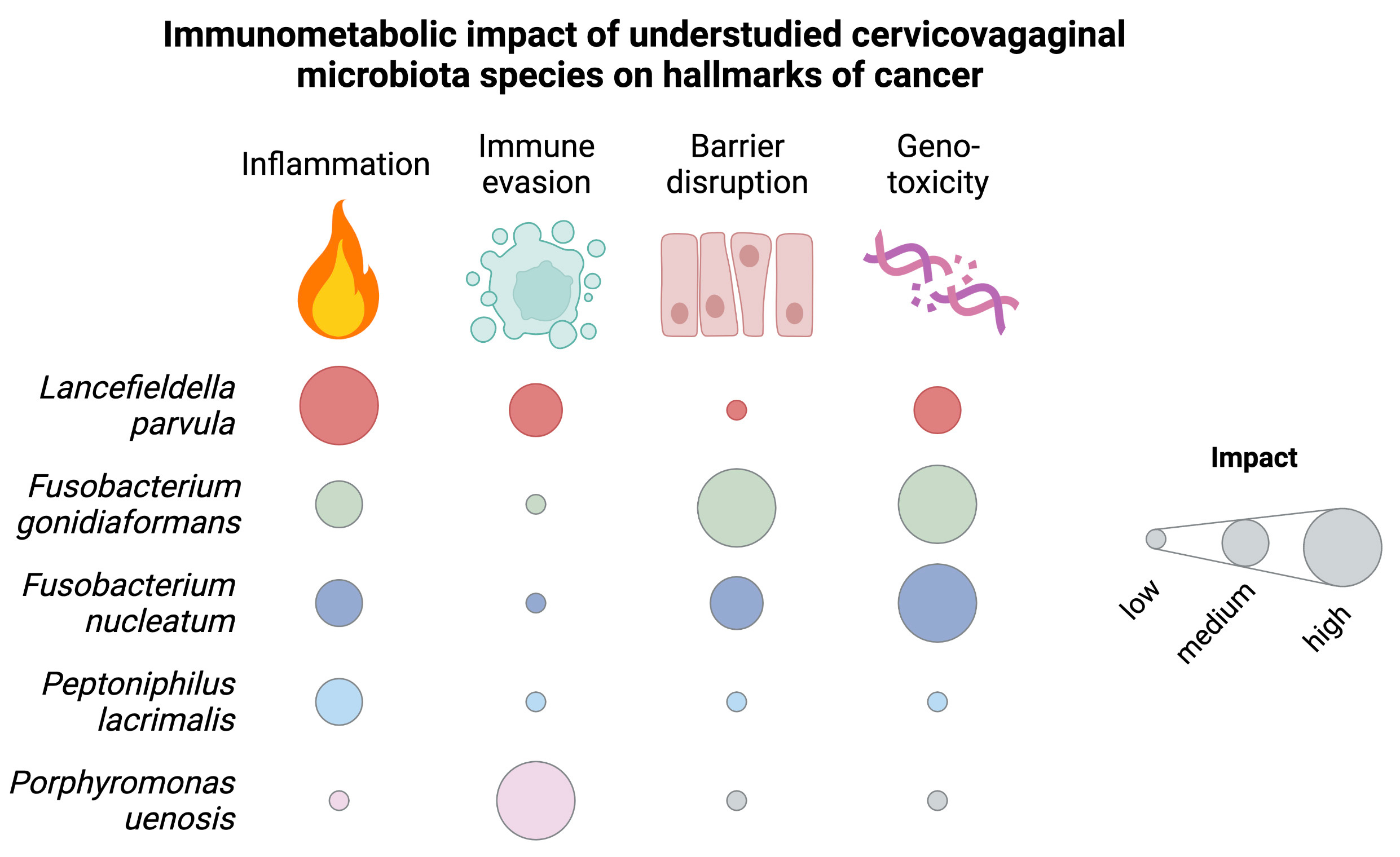
Summary Figure 2. Summary of the putative pro-carcinogenic immunometabolic mechanisms employed by bacterial vaginosis-associated bacteria of the lower female reproductive tract (from Maarsingh, Laniewski, & Herbst-Kralovetz, Communications Biology, 22;5(1):725, 2022)
Dysregulation of metabolism is an active research area in relation to the hallmarks of cancer. We hypothesized that some cervical cancer associated species studied in this report, may modulate a pro-carcinogenic cervicovaginal metabolic microenvironment. To address this hypothesis, we leveraged liquid chromatography/mass spectrometry-based untargeted metabolomics to study the metabolic landscape of culture supernatants of 3-D cervical cells after infection with BVAB. We found that the well-studied oncometabolite 2-hydroxyglutarate was significantly accumulated in culture supernatants after infection with L. parvula, F. gonidiaformans, and F. nucleatum. Using Random Forest, a supervised machine learning approach, we found metabolic signatures of histidine catabolism induced by multiple BVAB. This may implicate disruption of the cervical epithelial barrier by BVAB, specifically involving degradation of the keratinocyte marker filaggrin. Additionally, we found that F. gonidiaformans and F. nucleatum induced robust metabolic signatures of hydrogen sulfide production and oxidative stress. Hydrogen sulfide and oxidative stress metabolites (e.g., superoxide) are genotoxic and persistent infection with BVAB that modulate accumulation these metabolites may promote a pro-carcinogenic state. Finally, we discovered that both F. gonidiaforms and F. nucleatum induced extracellular accumulation of sphingolipids and glycerophospholipids. Lipids are known pro-inflammatory mediators and may be indicative of cellular damage. Furthermore, some sphingolipids induce cancer cell survival mechanisms. Together, we show that BVAB associated with malignancies of the FRT may promote a pro-carcinogenic metabolic microenvironment in a species-specific manner.

Follow the Topic
-
Communications Biology

An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
Signalling Pathways of Innate Immunity
Publishing Model: Hybrid
Deadline: Feb 28, 2026
Forces in Cell Biology
Publishing Model: Open Access
Deadline: Apr 30, 2026

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in