Molecular traffic across the biological membrane is a tightly regulated process, key to maintain adequate cellular concentrations. Import of essential nutrients, changes in ion concentrations and export of cytotoxic molecules is mediated by specialized transmembrane proteins, such as transporters. These proteins are highly dynamic entities. Transporters twist, bend and rearrange dramatically to provide VIP access for their substrate through the surrounding lipid bilayer. All these complex movements are happening within the lipid environment of the membrane, and it is becoming increasingly clear that the lipids are not passive bystanders. Membrane lipids are directly involved in the transport mechanism.
How this fundamental transport process happens at molecular level is not yet understood. How does a transporter change conformation? What role do lipids play in these structural rearrangements? In this work, Argyris Politis and his team and collaborators provide answers to this puzzling molecular enigma. We combined hydrogen-deuterium exchange mass spectrometry (HDX-MS) and molecular dynamics (MD) simulations on transporters reconstituted in different lipid environments to gain insight in the elusive transport process.
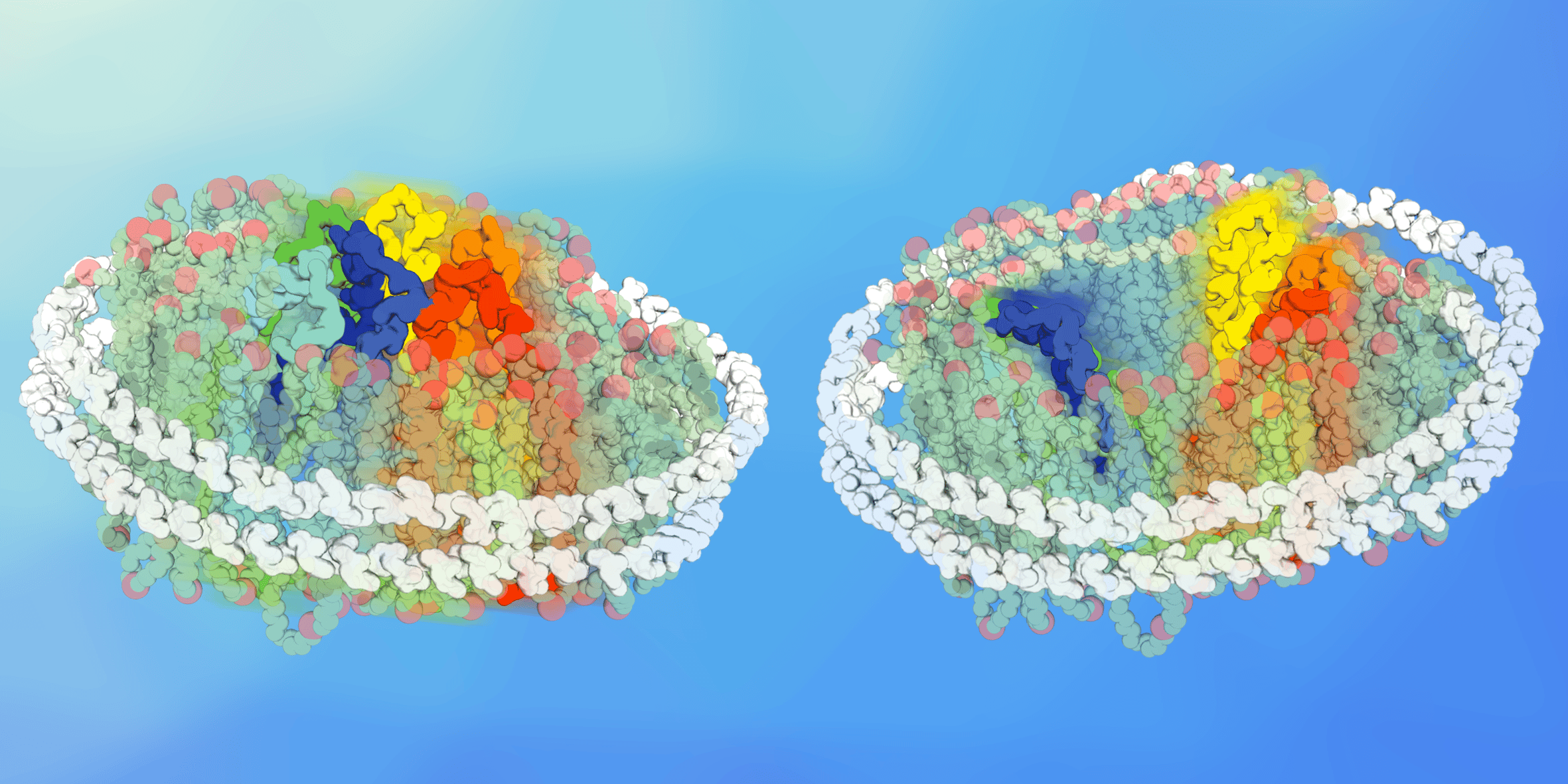
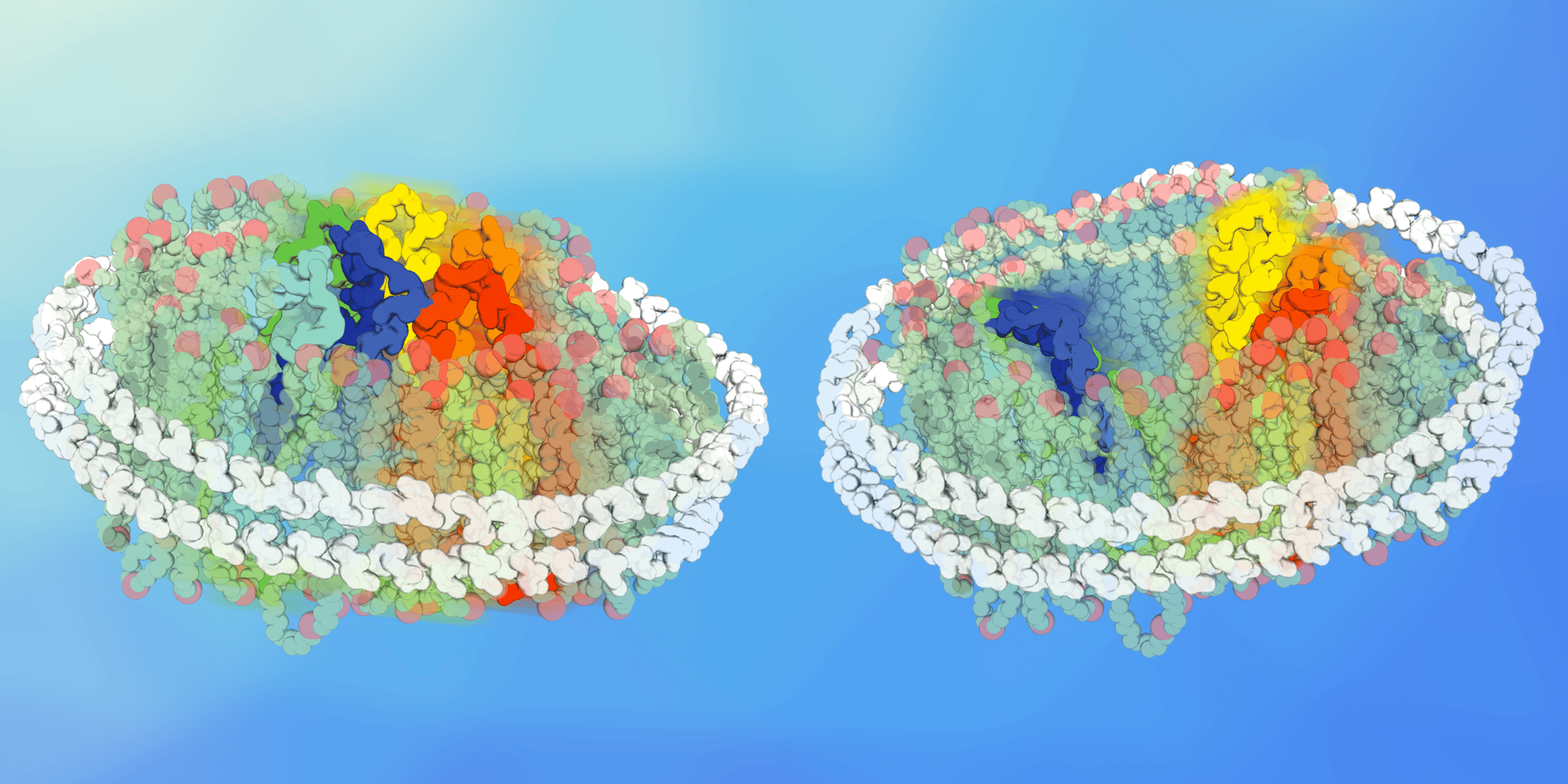
Figure 1: Membrane transporters reconstituted in nanoscale lipid bilayers of tuneable composition, the so-called nanodiscs.
Like many projects, this one came out from the meeting of people with converging interests. Argyris and Chloe were both convinced that the role of lipids in the mechanism of membrane proteins had been overlooked. We used our expertise in mass spectrometry and membrane protein biochemistry to study a series of homologous transporters and see whether we could highlight general principles of lipid-mediated regulation of transport. We decided to look at the bacterial transporters XylE, LacY and GlpT, partly because they were well characterized at the structural level and mostly because these proteins were used by the Booth group who happened to be next door.
After tedious protocol optimizations and benchmarking experiments, we were confident that HDX-MS was suitable to follow the conformational changes of transporters. We then started the dissection of the molecular mechanism of transport and most interestingly, to try to answer what lipids might be doing to the protein. The first pleasant surprise came from the realisation that all three transporters use a similar mechanism to alternate conformations. They present conserved charge networks, located far from the substrate biding site, which act as molecular switches. The disruption-formation cycle of these networks control the conformational transition from one state to another.
We then set out to understand how lipids change conformations - if they do - and started experiments in nanodiscs. These nanoscale lipid bilayers allows to reconstitute virtually any membrane proteins in a lipid bilayer of chosen composition. We used this tool to detect whether a specific lipid environment could favour a specific conformational intermediate and were pleased to see that indeed, the conformational equilibrium was different in different lipid environments. While presenting these preliminary results at the annual biophysical society meeting in New Orleans in 2017, Chloe was approached by Emad Tajkhorshid. “We think lipids are important too”, he told her. From this mutual understanding started a fruitful collaboration. Mrinal Shekhar from Emad’s group carried out molecular dynamics simulations of XylE and LacY in bilayers of lipid compositions mirroring the ones used in nanodiscs. And that’s when the second good surprise kicked in. In his simulations, Mrinal could see that a specific lipid species was interacting with the charge networks identified as conformational switches. They were disrupting the charge network and thereby switching the equilibrium towards a state open to the cytoplasmic side. The combination of MD simulations and HDX-MS experiments demonstrated clearly that direct and specific lipid-protein interactions modulate the transition between different conformational states.
Together, these findings pave the way for a better description of the mechanism of transporters, and membrane proteins in general. We speculate that the majority of the mechanisms of membrane proteins include lipids. We have to learn to shift the focus of our structural biologist lens to capture the bigger picture.
All the details of our work can be read in our article: “Direct lipid-protein interactions shape the conformational landscape of secondary transporters” in Nature Communications.
Written by Chloe Martens
Figure was created by Andy Lau


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in