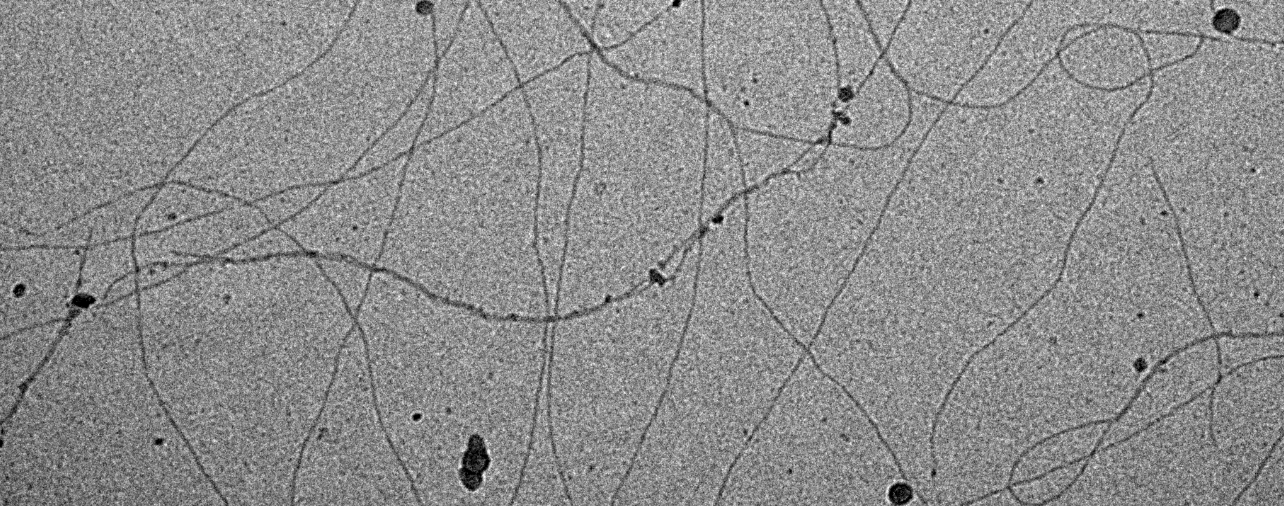
Text books tell us that type IV pili –proteinaceous extracellular appendages, which look like cables and that are made by a wide range of bacteria– are mainly involved in some kind of attachment, i.e. surface motility and biofilm formation or conjugation and DNA acquisition. But why would purely planktonic organisms such as marine cyanobacteria, which grow in environments where surfaces are rarely encountered and that are poor at taking up DNA, produce these nitrogen-expensive structures? In our paper we define a new function for these fascinating appendages: Pili allow planktonic cyanobacterial cells to retain an optimal position in the water column as well as evade predation by grazers (Fig 1).
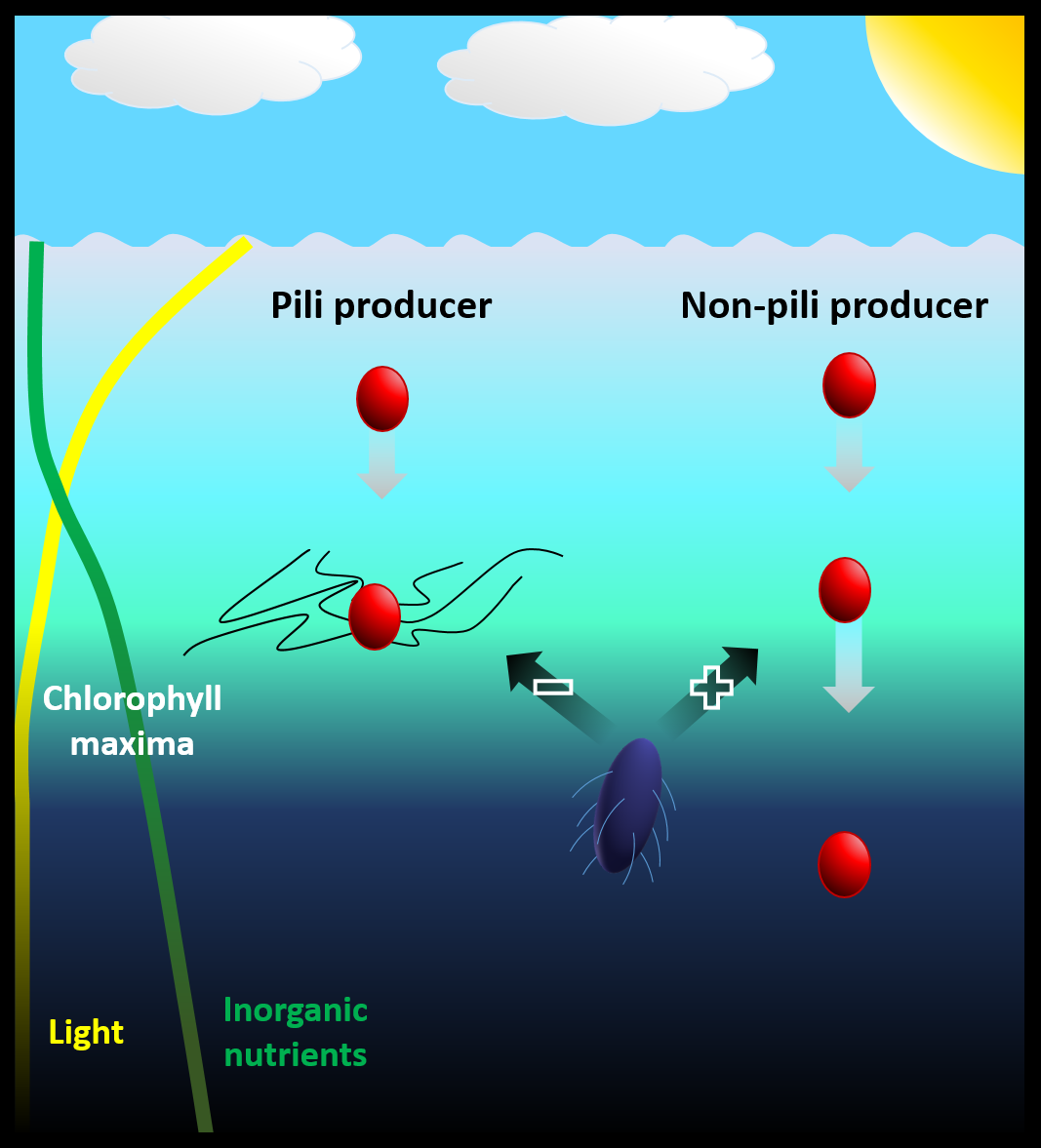
Fig 1 | Graphical abstract representing the biological advantages type IV pili represent to marine cyanobacteria, helping cells to increase their drag and position themselves in the water column as well as evade protozoan grazing.
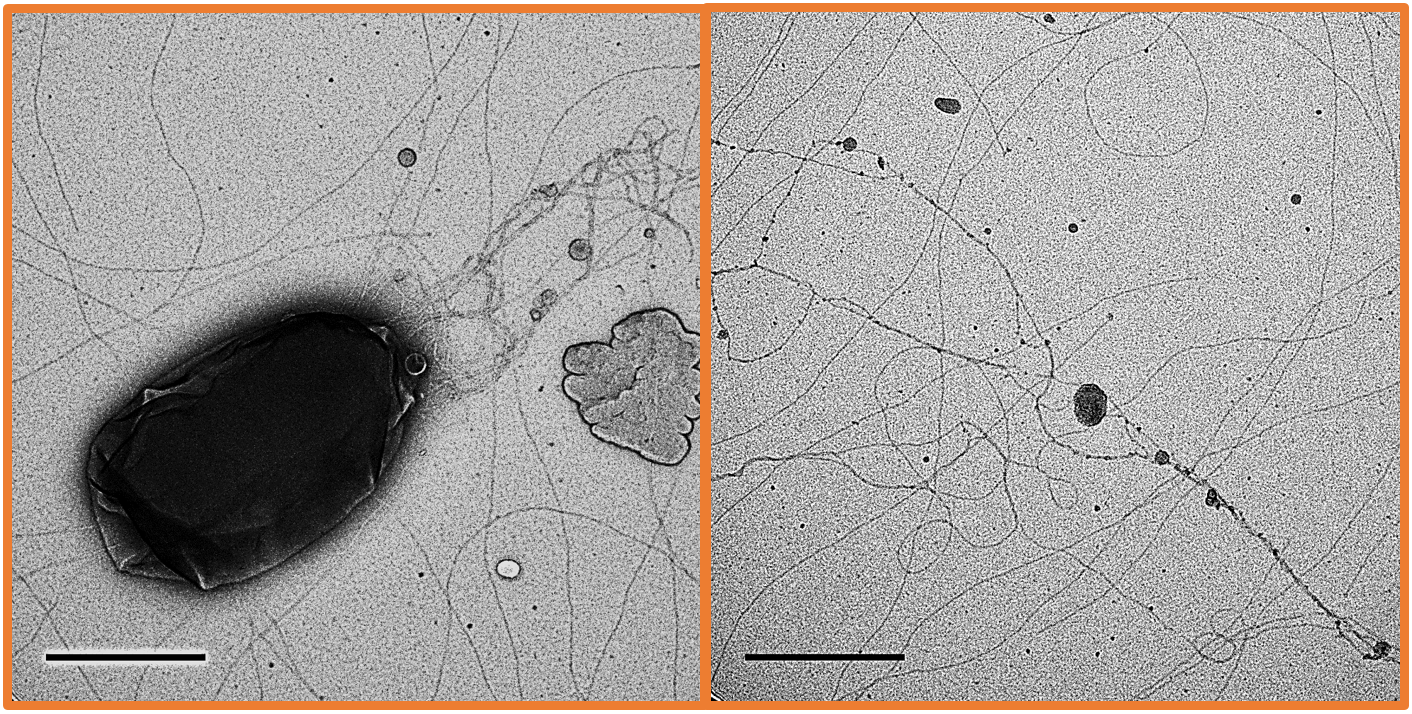
We all know how high-throughput ‘omic techniques are particularly good at flagging interesting features that generate a myriad of hypotheses that need experimental testing. And this is how this story started. When analysing exoproteomes from marine bacteria, we systematically observed an exceptionally high production of pili-like proteins in the milieu of the marine cyanobacterium Synechococcus sp. WH7803. During shotgun proteomics, where 100s or 1000s of proteins are detected, when you observe one single protein representing up to 25% of the proteome… it is definitely something worth further investigation! As I was once told, detecting the main subunit of a protein structure in a proteomic dataset does not mean the structure is being assembled. Hence, we checked whether PilA1 –the main pili subunit swamping the proteomic datasets– was in fact assembled into pili structures. This was done by transmission electron microscopy (TEM; Fig 2). It was surprising to see the abundance and length of these structures (over 10 µm). It is in fact not easy to prepare piliated cells on TEM grids because: i) centrifugation to concentrate cells shears off the pili, and ii) cells with pili do not seem to ‘stick’ well to the grids. Is this the reason why they were not observed to date?
At the time, we were looking for a good gene target for knocking out in order to try an improved protocol to generate mutants in marine cyanobacteria. Anyone that has worked with these organisms knows how difficult it is to grow them and, moreover, how complicated it is to genetically manipulate them. They do not grow well forming colonies on plates and this is aggravated when searching for a mutant cell amongst millions of dead ones. Considering that the plating efficiency of marine cyanobacteria is low, and the combined frequency of conjugation and recombination during mutagenesis is around 1 in 109, a cell that has successfully undergone mutation needs to grow amongst billions of dead cells. We formulated a protocol by which we implemented a ‘helper’ strain that helps detoxify the environment from the hoard of organic matter originating from dead cells –which is known to kill Synechococcus [Christie-Oleza et al. 2017]. This ‘helper’ stain contains sacB and, hence, can be eliminated by the addition of sucrose to redeem an axenic Synechococcus culture. The protocol can be found in Supplementary Note 1 of our paper. It takes over a month for Synechococcus to grow colonies in sloppy agar, so you can imagine how lengthy the entire process can be. But it worked!
When mutant colonies were retrieved we were already planning a number of experiments to determine the function of these pili as, to this point, we had no idea why marine cyanobacteria would produce them. There were a lot of experiments planed, but while we were growing up the mutants in liquid cultures, the phenotype became pretty obvious (Fig 3)! The mutant cultures consistently sank and they were no longer capable of retaining their planktonic form unless cultures were shaken. As Daniel Sher (@MCE_lab) mentioned in a Tweet, pili in marine cyanobacteria function as pico-parachutes. Apart from increasing drag and avoiding sinking, the rapid extension and retraction of pili allows cells to retain an optimal position in the water column (e.g. nutrient hotspots, optimal light position), a clear advantage versus those that cannot produce pili and randomly drift in the water. We further demonstrated another environmental advantage to producing pili, as these appendages allowed the cyanobacterium to avoid being preyed upon by grazing organisms. Do grazing organisms get entangled in these long pili? How cool would it be if we could image it!
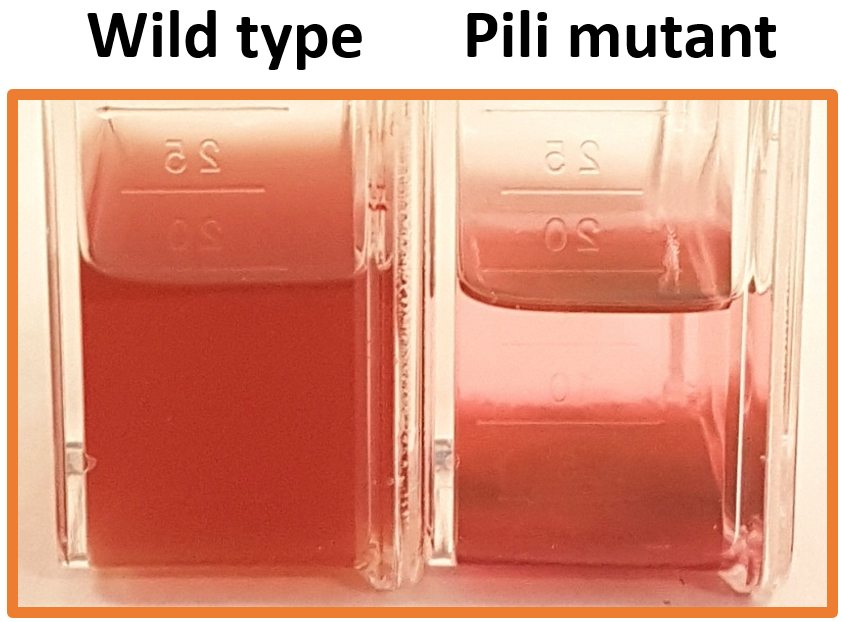
Fig 3 | Non-shaken cultures of wild type and pili mutant Synechococcus sp. WH7803. All cultures of pili mutants are no longer capable of maintaining their planktonic phenotype and have a sinking rate of ~1 cm day-1 which is not caused by cell aggregation.
This discovery opens up an entirely new area of investigation. Why would only one quarter of all marine cyanobacteria in the oceans encode this kind of pili? Alternative methods of flotation most likely exist as strains that do not encode pili (as well as the well-documented absence of flagella or gas vesicles in these marine organisms) are still capable of remaining planktonic. Furthermore, pili have clear ecological trade-offs. Apart from the obvious costly nitrogen investment of producing pili, another is the increased chance of being infected by phage since such pili structures are likely prime phage receptor targets. It is definitely time to use current ‘omic knowledge to go back to the bench and test hypotheses generated from such data. Ultimately, obtaining a mechanistic understanding of processes critical to Earth’s function requires culturing and genetic manipulation of organisms, and hopefully this study highlights the benefits of adopting such a strategy.
Read our paper in….. https://www.nature.com/articles/s41467-021-22152-w
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in