Mass die-offs of marine birds and mammals in Peru sound the alarm on the spread of highly pathogenic avian influenza (H5N1) viruses throughout South America
Published in Microbiology

Reader discretion advised: This post contains images of deceased animals that some readers may find distressing.
It began with the pelicans. Early in 2023 Peru’s pristine beaches became littered with thousands of bird carcasses, as more than 40% of Peruvian pelicans (Pelecanus thagus) succumbed to the new and highly lethal H5N1 avian influenza strain that was quickly moving down the coast. The virus caused a systemic infection that spread throughout the body, eventually reaching the brain. The birds appeared disoriented, jerked and twisted their long necks, and often walked in circles before dropping dead. But the pelicans were only the start. The virus was spreading fast.

South American sea lions (Otaria flavescens) were next. These are highly social animals that live in large boisterous colonies along the South American coastline, from Peru to Argentina, nuzzling pups and tussling on the shorelines. O. flavescens had no prior history of infection with avian influenza. For decades, South America’s geographic isolation had buffered its wildlife from the highly pathogenic flu viruses that routinely ravaged Europe, Asia, and occasionally North America. But the new H5N1 that emerged in Eurasia in 2021 was unusually transmissible and lethal. Wild birds quickly spread the virus from Europe to North America, where it is still causing the largest animal disease outbreak in US history. After ravaging US poultry farms, the virus cropped up in grizzly bears, bobcats, racoons and many other wild animals that had never reported influenza before.
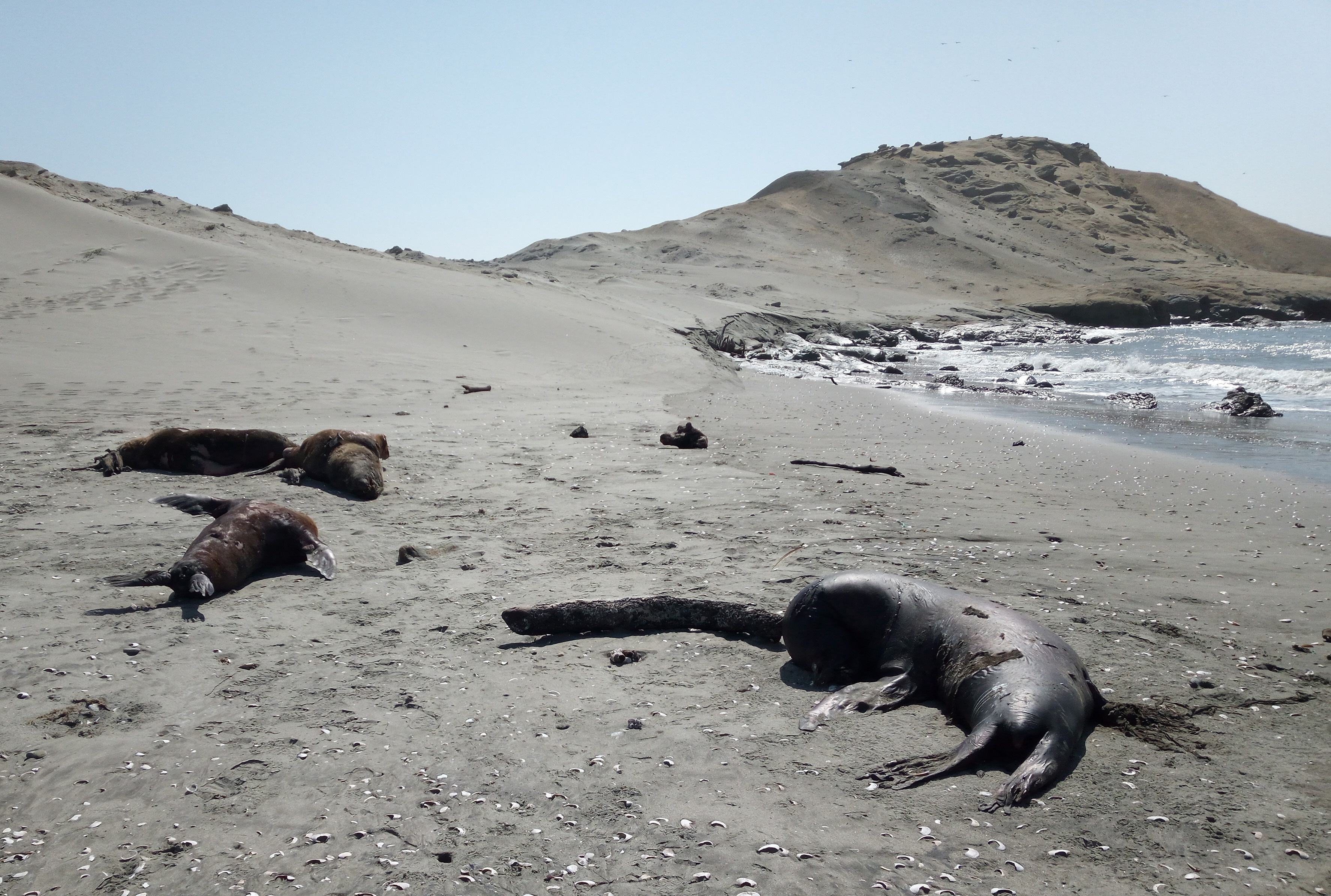

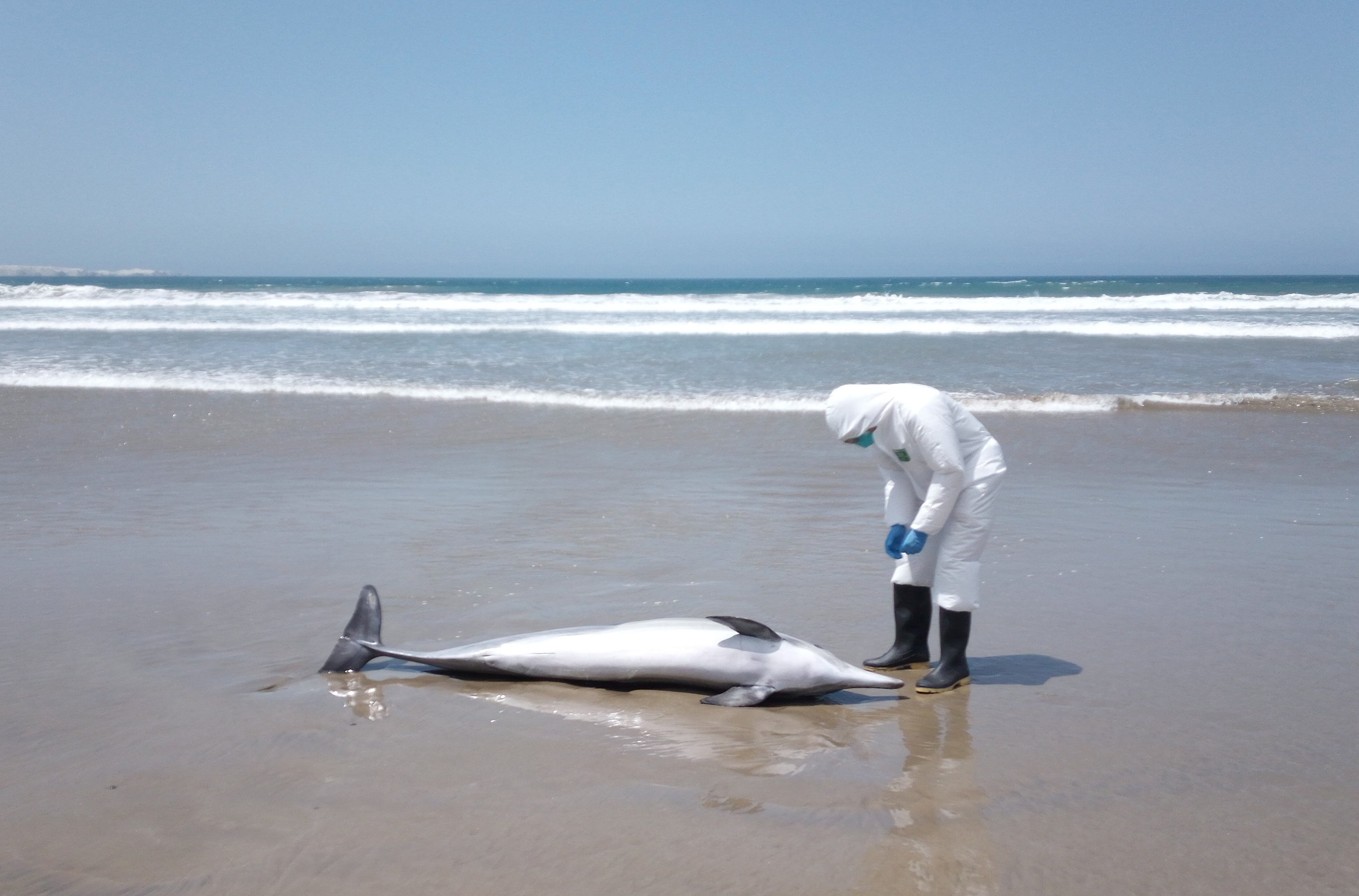
Still, it was shocking when thousands of sea lions started to wash onto Peru’s beaches, many already dead, while others were displaying neurological abnormalities like those observed in the pelicans. Even a dolphin washed up dead on shore. Samples from marine mammals and sea birds were collected by Peruvian authorities and sent to scientists, who worked around the clock to establish the cause of death. In some cases it was difficult to extract enough virus RNA for genetic sequencing, as the animals tested had been dead for a while and their tissues exhibited extensive degradation. However, complete genetic sequences of H5N1 viruses from three species of sea birds (sandpiper, pelican, cormorant) and two species of mammals (South American sea lion and common dolphin) were obtained, providing enough data to begin to piece together the story.

The H5N1 viruses in Peru were genetically similar to the H5N1 viruses that had circulated earlier in wild birds in North America, which supported the conclusion that the virus was introduced to Peru by migratory birds arriving from North America to overwinter in warmer climates. But there was a twist. The Peruvian virus was a “reassortant.” Half of its genome was related to the Eurasian H5N1 virus that originally spread through wild birds from Europe to North America in late 2021. The other half came from less pathogenic viruses that circulated in wild birds in the Americas, and was likely picked up while the H5N1 virus raced across the US, before it arrived in Peru. One can imagine that adding new genes from a less pathogenic virus might have attenuated the virus to make it less deadly. However, that did not happen. Rather, the altered genome appears to have opened new pathways for H5N1 evolution, and potentially new ways to adapt to mammals.

Bird-to-mammal host switches require a collection of mutations in key sites that help the virus bind to different host cell receptors and replicate its genome using host cell machinery. Two polymerase mutations known to help avian viruses replicate in mammalian cells (D701N and Q591K) were found in several H5N1 viruses obtained from sea lions in Peru. The H5N1 story in the Americas continues to evolve, and since the time of our Peru study, similar double-mutant reassortant H5N1 viruses have been found in one human and several sea lions in Chile. Furthermore, shortly after the Peruvian outbreak, Chile also reported mass die-offs of marine mammals and penguins. By late August, Argentina followed suit with sea lion die-offs that were also traced to H5N1. While many hoped the H5N1 situation in South America would stabilize when the migratory birds returned to North America for summer breeding, the continued mass die-offs suggest otherwise, and raise concerns for a re-surgency of circulating highly pathogenic avian influenza viruses in the region.
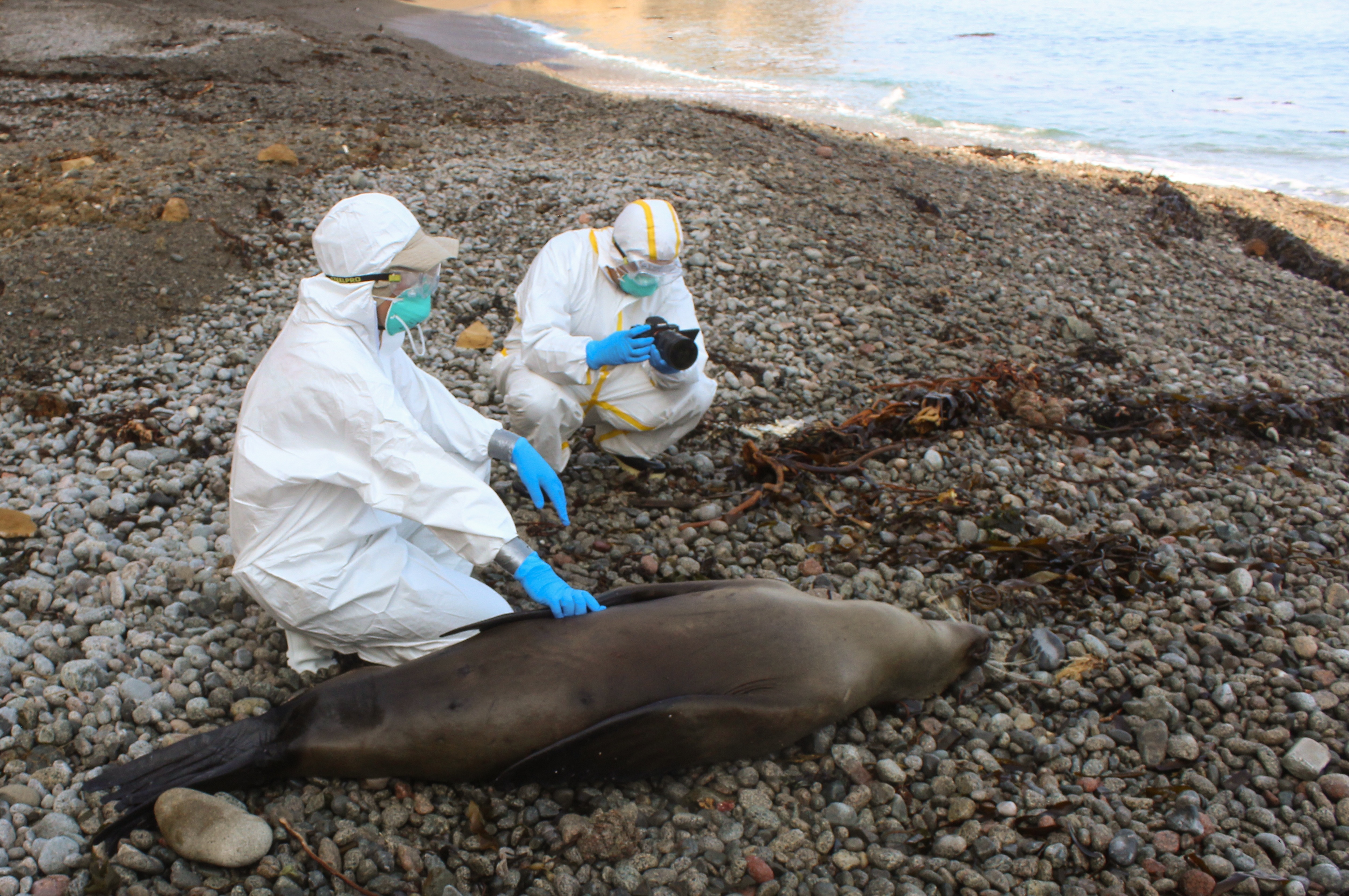

Scientists have been left scratching their heads, wondering how exactly sea lions and other marine mammals in South America are getting infected –perhaps by contaminated beaches or sea water, or by scavenging of deceased birds? A crucial question that remains is whether the virus ever spread mammal-to-mammal. In theory, mammal-to-mammal transmission can be spotted on phylogenetic trees based on the clustering patterns of H5N1 sequences gathered from mammalian hosts. But large numbers of background sequences from wild birds are needed in these trees to ensure that apparent mammalian clusters are not sampling artifacts, or arising by chance. During the COVID-19 pandemic many countries, including in the global South, increased high-throughput genomic sequencing capacity to sequence SARS-CoV-2 viruses. But large-scale sequencing did not translate into sequencing of other respiratory viruses, and most definitely has not occurred for an H5N1 wildlife outbreak that has yet to pose obvious imminent danger to humans. To date, there are fewer H5N1 global sequences available from all wild bird, poultry, mammalian and human samples combined during 2021-2023 (n = 6,982) than for SARS-CoV-2 viruses sequenced last year from Delaware, USA (population 1 million).
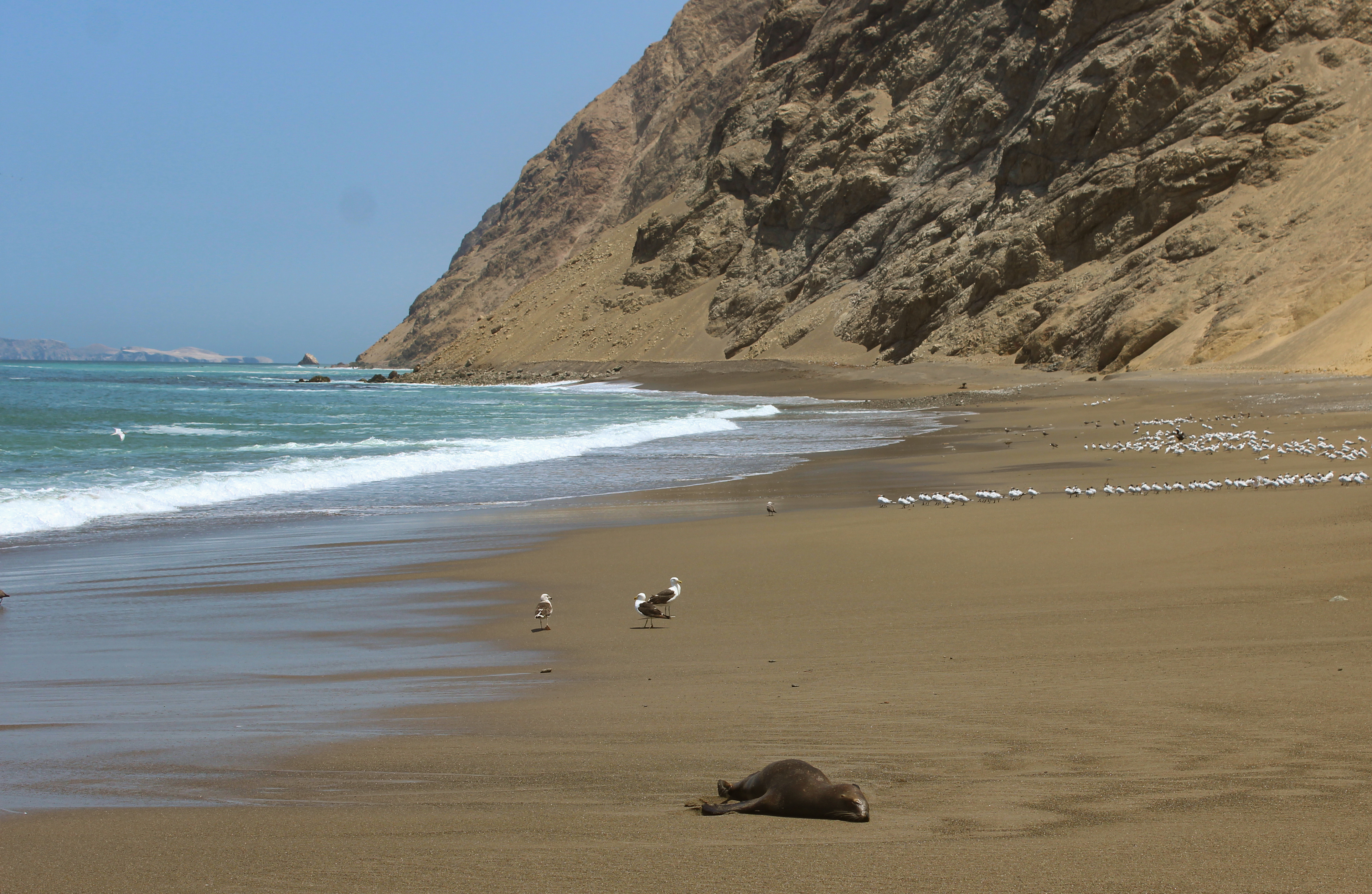

Despite these gaps in data, scientists must advise policymakers on how to respond to new and unfamiliar threats. While Asia has a long history of vaccinating poultry against influenza, vaccination remains controversial in the Americas because of economic trade implications. While the US began vaccinating endangered Californian Condors against H5N1, such a costly and logistically complicated campaign is unlikely to be extended to Peru’s endangered wildlife, like the Humboldt penguin. Climate change, habitat loss, and other stressors have already put many wildlife species on the brink. The COVID-19 pandemic was a warning that rapid planetary changes are creating new disease threats, and we must address the underlying drivers of disease emergence before they spiral into additional global pandemics.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in