Metabolic kinases in the renal cancer road
Published in Cancer
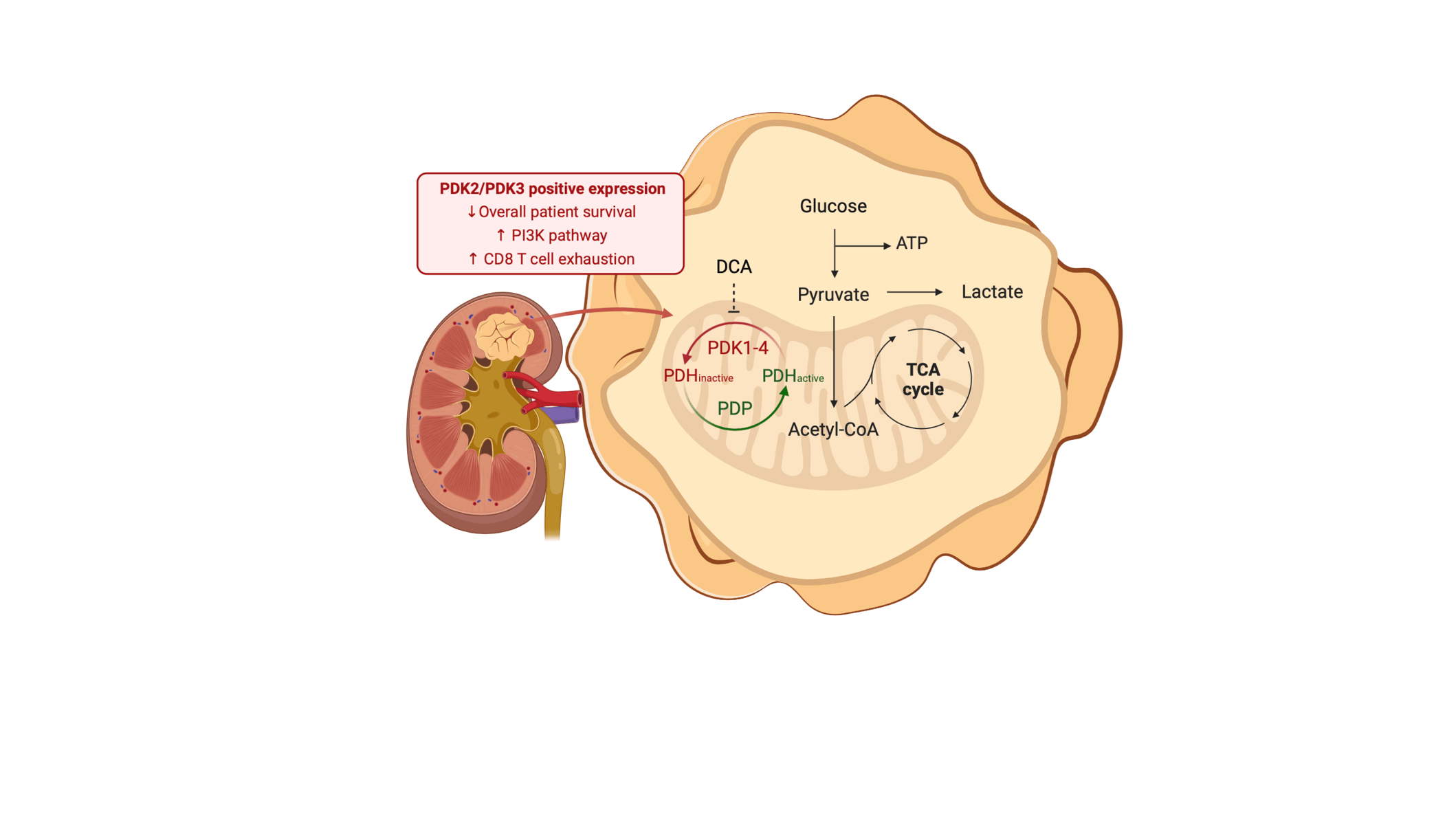
Renal cancer cells constitute a paradigm of tumor cells with a glycolytic reprogramming which drives metabolic alterations favoring cell survival and transformation (1). During glycolysis, pyruvate is converted by the pyruvate dehydrogenase complex into Acetyl-CoA before it enters into the TCA/Krebs cycle. The pyruvate dehydrogenase complex (PDH) is a holoenzyme composed of three catalytic subunits assembled into a multimolecular complex, and its regulation is tightly controlled by phosphorylation/dephosphorylation by pyruvate dehydrogenase kinases (PDK; inactivate PDH) and pyruvate dehydrogenase phosphatases (PDP; activate PDH) (2). The inhibitory targeting of PDK metabolic kinases constitutes a potential therapy in the treatment of cancer, including renal cancer (3).
We have studied the expression of regulators of energy metabolism in renal cancer by immunohistochemistry on renal tumor samples, and observed a selective expression of the pyruvate dehydrogenase kinases (PDK1-4), responsible for phosphorylating and inactivating the pyruvate dehydrogenase complex (4). High PDK1 expression associated with good prognosis, whereas high PDK2 and PDK3 expression associated with lower overall survival of renal cancer patients (Figure 1). This suggests differential contribution of the distinct PDK in the outcome of renal cancer. In our study, PDK2 showed a predominantly nuclear localization in renal cancer tumors, whereas PDK1, PDK3 and PDK4 displayed a major cytoplasmic localization. A similar PDK subcellular localization pattern was previously observed in prostate tumors (5), suggesting different spatial functions associated to the distinct PDK, including nuclear functions. Whether TCA cycle enzymes play functional roles in the nucleus of cancer cells is a current open issue that deserves to be investigated. In addition, the possibility that specific PDK exert catalytic activities in cancer beyond their role as pyruvate dehydrogenase kinases is also worth to explore.
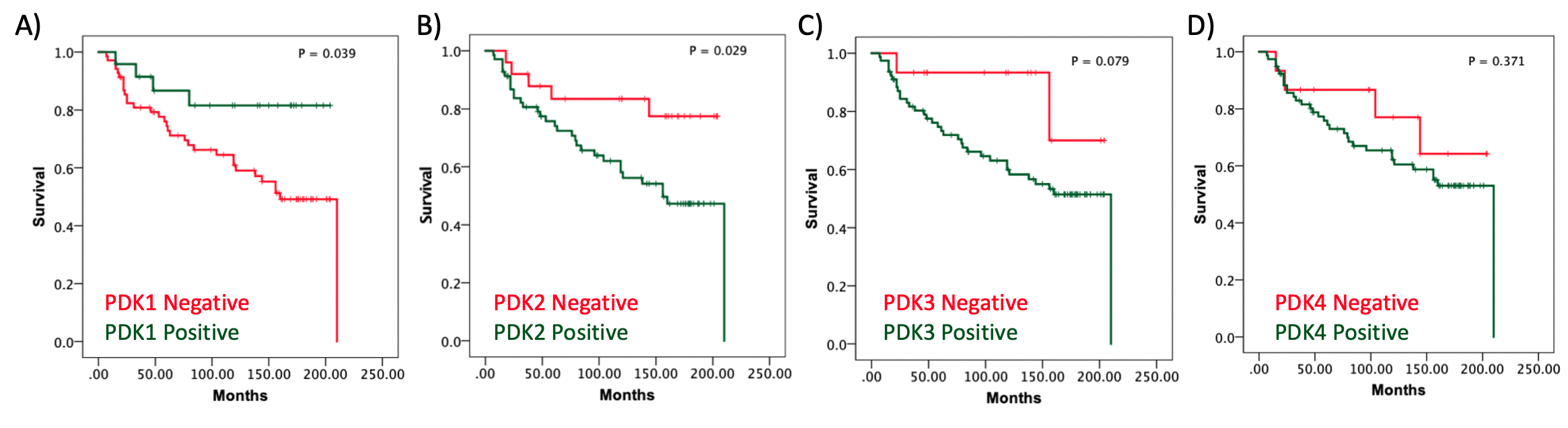
Figure 1. Kaplan-Meier survival curves of clear cell renal cell carcinoma (ccRCC) patients according to PDK1-4 tumor expression. Kaplan–Meier curves of overall survival based on PDK1 (A), PDK2 (B), PDK3(C), and PDK4 (D) protein expression in the ccRCC cohort.
We isolated RNA from a subgroup of whole tumor tissue sections (PDK2/PDK3 positive vs. PDK2/PDK3 negative), and using nanoString technologies we analyzed gene expression profiles in pan cancer and immune exhaustion panels. Gene expression analysis revealed molecular association of PDK2 and PDK3 expression with PI3K signaling pathway, as well as with T cell infiltration and exhausted CD8 T cells (Figure 2).
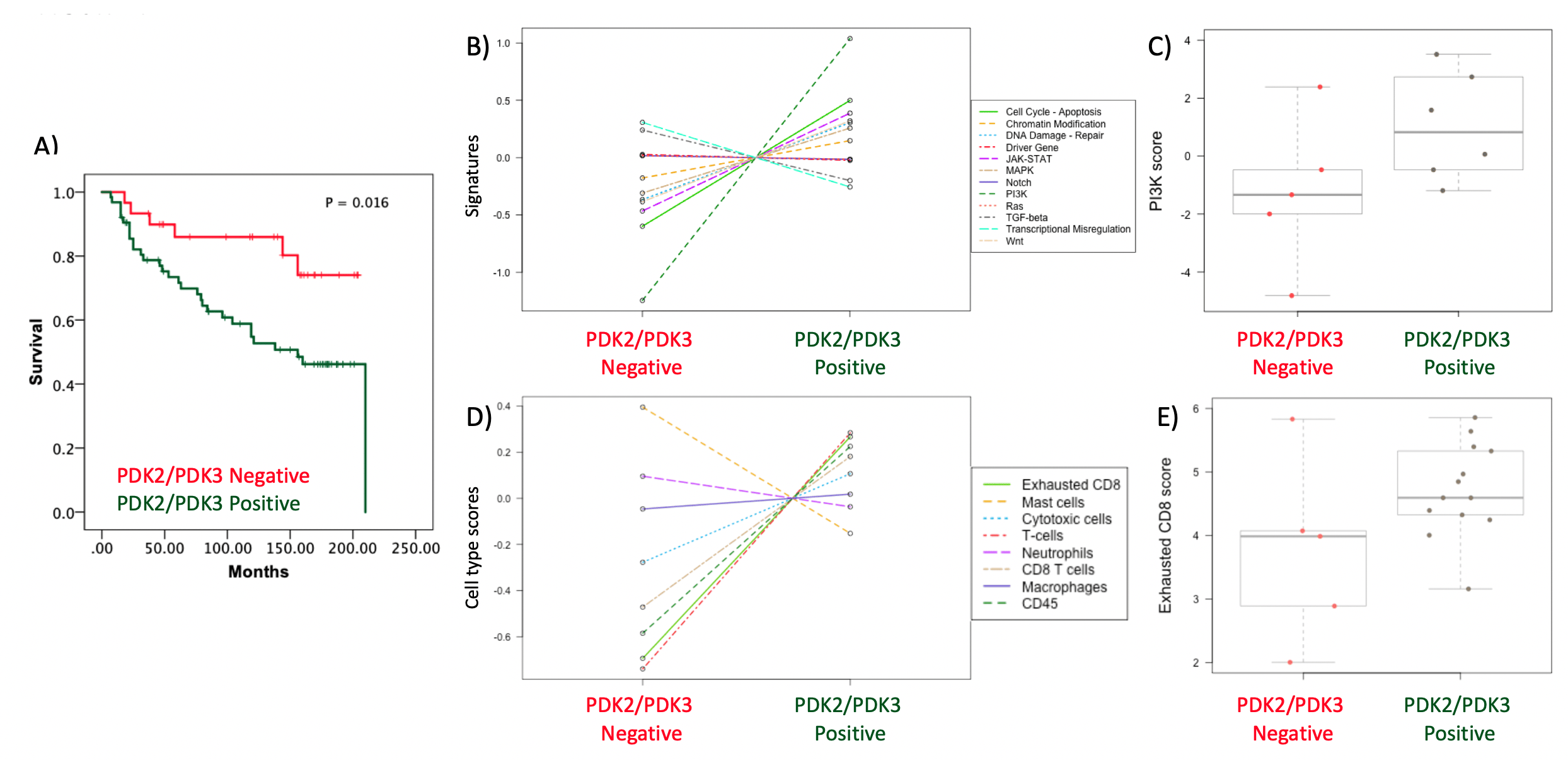
Figure 2. Molecular analysis of ccRCC patients according to PDK2 and PDK3 tumor expression. A) Kaplan–Meier curves of overall survival based on PDK2/PDK3 expression in the ccRCC cohort. Patients with high PDK2/PDK3 protein expression and patients with low PDK2/PDK3 protein expression had significantly different time to recurrence. B) PanCancer Pathway signature in ccRCC tumor samples. Note PI3K pathway as most differentially expressed signaling pathway in high PDK2/PDK3 protein expression vs low PDK2/PDK3 protein expression tumors. C) Box plot of PI3K pathway score in high PDK2/PDK3 protein expression vs low PDK2/PDK3 protein expression tumors. D) Cell type score in ccRCC tumor samples using Immune Exhaustion panel. Note CD8 T cell exhaustion cell type as one of the higher enrichments in high PDK2/PDK3 protein expression vs low PDK2/PDK3 protein expression tumors. E) Box plot of exhausted CD8 score in high PDK2/PDK3 protein expression vs low PDK2/PDK3 protein expression tumors.
PDK can be inhibited by dichloroacetate (DCA), which has been proposed as a therapeutic approach in several cancer types, including renal cancer (6). In our analysis, treatment of human renal cancer cell lines with DCA resulted in lower cell viability, which was accompanied by an increase in pAKT. The identity of the PDK inhibited by DCA in renal cancer cells needs to be explored, as well as the functional relation of PDK activities with the PI3K/AKT survival pathway. In addition, the use of alternative PDK inhibitors will be helpful to validate PDK inhibition as a therapeutic approach in renal cancer.
In summary, our work highlights the potential of the proteins of the pyruvate dehydrogenase molecular complex as targets for therapeutic intervention in renal cancer. Together, our findings suggest a differential role for PDK enzymes in ccRCC progression, and highlight PDK as actionable metabolic proteins in relation with PI3K signaling and exhausted CD8 T cells in ccRCC.
References:
(1) Chakraborty, S., Balan, M., Sabarwal, A., Choueiri, T. K. & Pal, S. Metabolic reprogramming in renal cancer: Events of a metabolic disease. Biochim Biophys Acta Rev Cancer 1876, 188559, doi:10.1016/j.bbcan.2021.188559 (2021).
(2) Patel, M. S., Nemeria, N. S., Furey, W. & Jordan, F. The pyruvate dehydrogenase complexes: structure-based function and regulation. J Biol Chem 289, 16615-16623, doi:10.1074/jbc.R114.563148 (2014).
(3) Stacpoole, P. W. Therapeutic Targeting of the Pyruvate Dehydrogenase Complex/Pyruvate Dehydrogenase Kinase (PDC/PDK) Axis in Cancer. J Natl Cancer Inst 109, doi:10.1093/jnci/djx071 (2017).
(4) Nunes-Xavier, C.E., Emaldi, M., Mingo, J. et al. The expression pattern of pyruvate dehydrogenase kinases predicts prognosis and correlates with immune exhaustion in clear cell renal cell carcinoma. Sci Rep 13, 7339, doi:10.1038/s41598-023-34087-x (2023)
(5) Nunes-Xavier, C. E., Mingo, J., Emaldi, M., et al. Heterogeneous Expression and Subcellular Localization of Pyruvate Dehydrogenase Complex in Prostate Cancer. Front Oncol 12, 873516, doi:10.3389/fonc.2022.873516 (2022).
(6) Kankotia, S. & Stacpoole, P. W. Dichloroacetate and cancer: new home for an orphan drug? Biochim Biophys Acta 1846, 617-629, doi:10.1016/j.bbcan.2014.08.005 (2014).
Follow the Topic
-
Scientific Reports
An open access journal publishing original research from across all areas of the natural sciences, psychology, medicine and engineering.
Related Collections
With Collections, you can get published faster and increase your visibility.
Phytochemicals and health
Publishing Model: Open Access
Deadline: Jul 28, 2026
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in