Microbial iron bandits moonlight as Robin Hood: Pyoverdine steals iron from cefiderocol to promote antibiotic cross-protection
Published in Microbiology
Iron is an essential trace nutrient for both humans and microorganisms, playing a crucial role in physiological processes including energy production and DNA synthesis. However, iron availability is limited. Hence, during infections, a tug-of-war for iron between bacteria and the host innate immune system emerges: as the host innate immunity restricts iron availability by secreting iron-binding proteins (e.g., lactoferrin), bacteria have evolved to secrete small molecules with high iron affinity, called siderophores. Due to their iron affinity, siderophores can displace iron from host ferric proteins.
Understanding how bacteria use their siderophore arsenal in the competition for iron enabled the development of cefiderocol, the first trojan-horse antibiotic approved for clinical use. Cefiderocol tricks bacteria into taking up the antibiotic due to the iron-binding component in its structure. Once cefiderocol is transported inside bacterial cells, iron dissociates and cefiderocol is released. Inside the cell, cefiderocol works like other antibiotics that kill bacteria by inhibiting cell wall synthesis. Because bacteria are starved for iron during infections by host iron binding proteins, bacteria express iron transporters on their surface at high levels, which guarantees increased drug uptake and makes cefiderocol very effective against multi-drug resistant microorganisms such as Pseudomonas aeruginosa.
As a new class of antibiotic, mechanisms of cefiderocol resistance are not fully understood. In the published paper “Siderophores promote cooperative interspecies and intraspecies cross-protection against antibiotics in vitro”, we sought to elucidate novel mechanisms of cefiderocol resistance in P. aeruginosa.
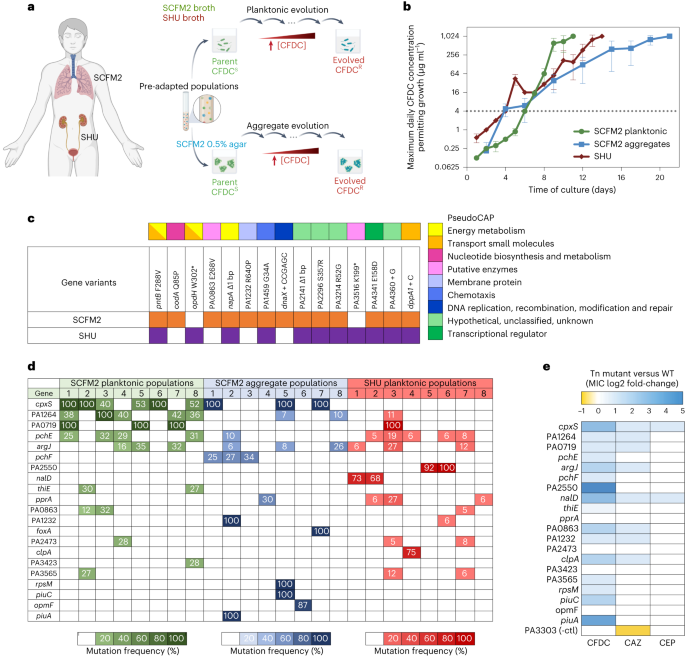
The antibiotic resistance emerges due to natural selection. When bacteria grow they naturally make rare errors in the DNA code, some of these rare DNA variations allow the bacteria to withstand antibiotic pressure and survive treatment while bacteria lacking the variations are killed by the antibiotic. To identify mutations arising under cefiderocol pressure, we modeled the evolution of cefiderocol resistance in our lab. For this, we cultured P. aeruginosa daily in increasing cefiderocol concentrations. When the evolved populations achieved high resistance levels, we sequenced the genomes of the ancestor and evolved populations to compare variations before and after the emergence of cefiderocol resistance. Using this approach, we identified 20 gene variants that were recurrently under selection during cefiderocol pressure. To better comprehend the consequences of evolving cefiderocol resistance, we explored how resistance mechanisms affected bacterial competitiveness.
The DNA variations that promote antibiotic resistance can impose growth disadvantages in antibiotic-free environments. We demonstrated that in drug-free conditions, cefiderocol resistant bacteria revert to a cefiderocol-susceptible state, and cefiderocol resistant populations are outcompeted by the ancestral susceptible strain, indicating that evolved cefiderocol resistance pays a fitness cost. On the other hand, we expected that under cefiderocol pressure, resistant individuals would outcompete the susceptible ones. Surprisingly, we identified cefiderocol resistant bacteria that cross-protected susceptible sibling bacteria, where cefiderocol-resistant evolved bacteria grew in the outermost layers of mixed aggregates, shielding the cefiderocol-susceptible ancestral bacteria from antibiotic killing (Fig 1).
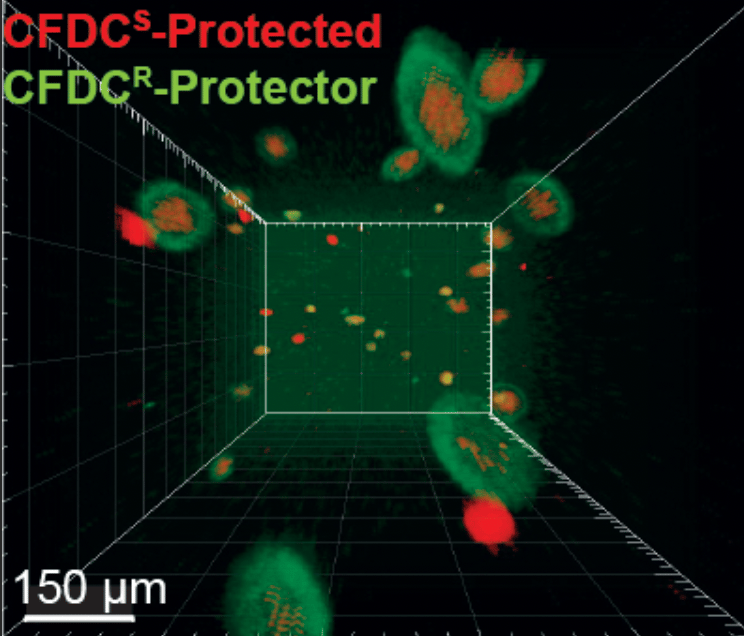

Antibiotic cross-protection arises when the concentration of antibiotics in the environment decreases locally due to the presence of antibiotic-resistant individuals within the population. Cross-protective cooperative behavior has been linked to the secretion of β-lactamases produced by resistant cells that degrade β-lactam antibiotics, protecting their susceptible siblings. Therefore, we expected that cross-protective populations would overexpress β-lactamase biosynthesis genes. Surprisingly, the expression of β-lactamase genes was unchanged. Instead, cross-protective populations upregulated CPX two-component system genes, as well as genes under CPX regulation, such as the muxABC-opmB efflux pump. Hence, we discovered a novel mechanism of cross-protection independent of enzymatic antibiotic degradation.
Although cefiderocol is not a substrate for MuxABC-OpmB, this efflux pump plays a role in resistance to novobiocin, aztreonam, macrolides, and tetracycline, and its overexpression in related bacteria results in increased secretion of a Pseudomonas siderophore, called pyoverdine. Therefore, we reasoned that pyoverdine, secreted by resistant “protector” cells, could act as a public good protecting the cefiderocol susceptible cells. Indeed, we observed that pyoverdine itself can not only prevent cefiderocol killing in P. aeruginosa, but also in other bacterial species including E. coli, Klebsiella pneumoniae, Burkholderia cenocepacia, and B. multivorans. The pyoverdine protective effect led us to ask whether their production in co-culture could drive interspecies cross-protection. A P. aeruginosa clinical isolate that produced massive amounts of pyoverdine cross-protected both K. pneumoniae and E. coli, while a pyoverdine non-producer did not. Together these findings support the hypothesis that pyoverdine drives cefiderocol cross-protection. But the question remained: how does this siderophore mediate this cooperative phenotype?
As previously mentioned in this blog post, pyoverdine is one of P. aeruginosa’s secret weapons in the tug-of-war for iron, and can displace iron from host innate immune proteins, such as lactoferrin. Likewise, we showed that pyoverdine can directly steal iron from cefiderocol. Since cefiderocol has to be bound to iron to hijack bacterial iron transporters, pyoverdine iron displacement decreases the bacterial antibiotic uptake, and, therefore, susceptible P. aeruginosa cells and members of polymicrobial communities are cross-protected by the diffusion of pyoverdine benevolently produced by resistant “protectors” (Fig 2).
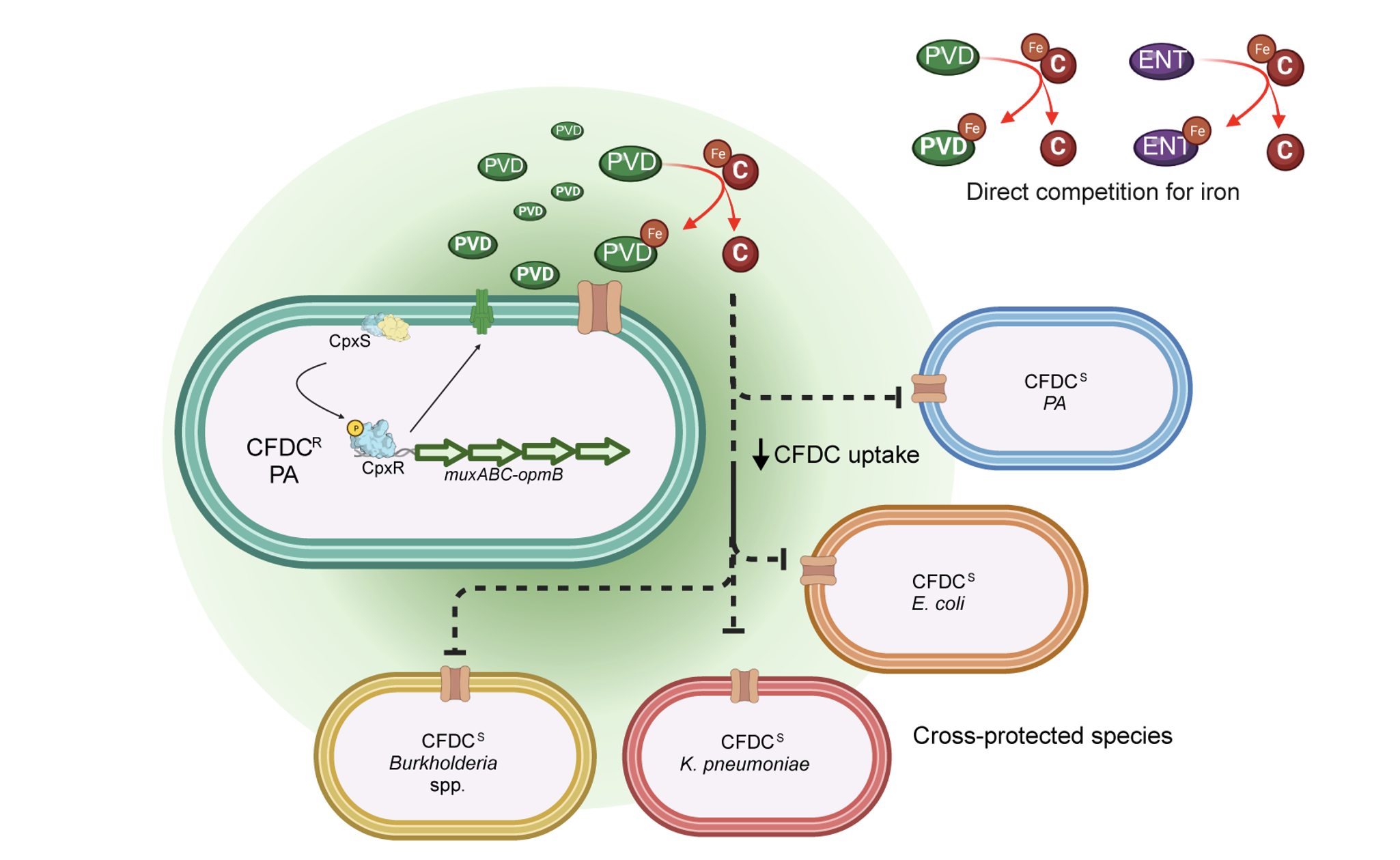
Figure 2- Pyoverdine drives cefiderocol cross-protection. Cefiderocol must be in its ferric form to be transported inside bacterial cells. Pyoverdine siderophore secreted by P. aeruginosa with high affinity for iron. We showed that cefiderocol selects for variants with increased pyoverdine secretion. The secretion of pyoverdine can directly steal iron from cefiderocol, which limits its uptake by bacterial cells, allowing susceptible neighbors to grow in a antibiotic concentration that would be otherwise lethal.
Although the cross-protective dynamics may pose challenges for cefiderocol treatment, we demonstrated that cefiderocol resistance impose a growth disadvantage in drug-free environments and allows susceptible cells to outcompete the resistant cells. Thus, we anticipate that adopting an intermittent therapeutic strategy could be beneficial in suppressing the prevalence of cefiderocol resistant populations.
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in