More than just the sum of its parts – exploring phenotypic diversity in microbial communities
Published in Microbiology
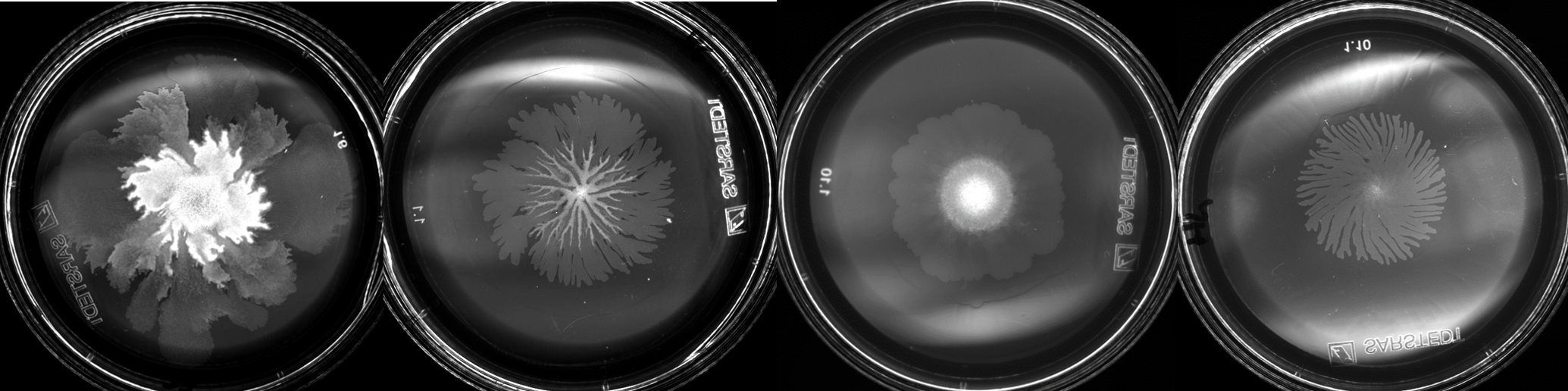

Creating something bigger than yourself, a thriving community, takes time, resources, communication and a diversity of complementary skills. We as humans tend to pride ourselves in having developed modern-day communities from small villages to big cities composed of many different people with diverse roles, providing more functionality, comfort, safety, and an overall robustness which could not have been achieved by a single person. And yet, complex, organized communities are not limited to humankind, but can be observed even for the smallest, simplest life form on earth – microbes. Long before humans, microbes already colonized every niche on this planet by creating such communities, often composed of multiple species, sometimes across different kingdoms of life. But even when originating from a single founder cell, their arrangement tends to be spatially structured, with different cell types emerging in the different areas of the community. Historically, however, experimental designs to study microbial behavior have often focused on well-mixed conditions, neglecting spatial effects. Imaging-based methods such as microscopy are an exception to this rule, but limited in the amount of information that can be captured about a cell’s internal state, characterized for example by gene expression, metabolite or protein abundance, which can be measured with so-called “-omics” methods. Experimental techniques to obtain spatially informed -omics measurements are therefore needed to advance the field, and in recent years, several approaches have been made in this direction, characterized by inventive experimental designs and a creative combination of existing methods.
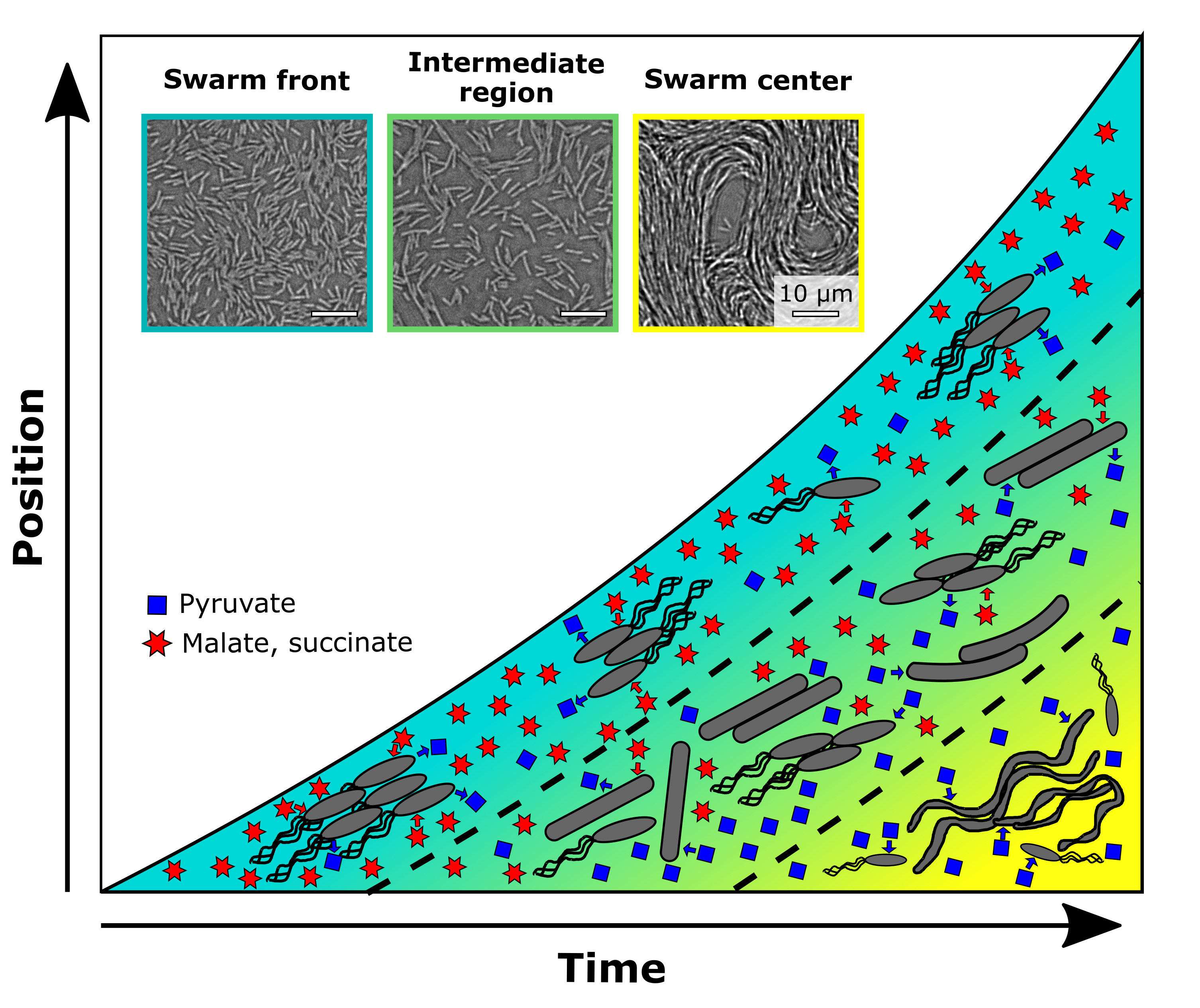
Inspired by these endeavors, we set out on our own adventure to achieve spatial –omics measurements for a spatially structured microbial community, to understand the cellular interactions and development processes within the community. We decided to focus on a Bacillus subtilits swarm, a two-dimensional community reaching macroscopic size, developing a “megacity” composed of millions to billions of cells with functionally distinct sub-populations in space and time. We developed a method to capture both microscopy and transcriptome data of swarming cells in a spatiotemporal manner and revealed the emergence of three subpopulations during swarming colony development, each characterized by distinct cell morphology, motility, metabolism, and gene regulation (see Fig. 1, Fig. 2). The leading subpopulation, located at the outer edge of the circular swarming colony, actively swarms and explores uncharted areas of the medium by consuming rich nutrients and secreting waste products. Following closely behind, the second intermediate subpopulation plays a crucial role in recycling the waste metabolites left behind by the first group. Additionally, it assists the front-runner cells by supplying surfactin, which facilitates colony expansion. Finally, the third subpopulation, situated at the center of the swarming colony, initiates biofilm formation and sporulation, likely in response to nutrient limitations. These cells might prepare for sudden changes in their surrounding environment, ensuring the adaptability of the entire microbial community. Our study successfully unraveled the emergence of these subpopulations and their distinct roles within a spatially structured microbial community and produced comprehensive microscopy and transcriptome datasets that we hope will contribute to the progress of this field. In this blog post, we will share our endeavors and struggles "behind the paper" and the lessons we learned from this project, which are not often highlighted in a published scientific paper.
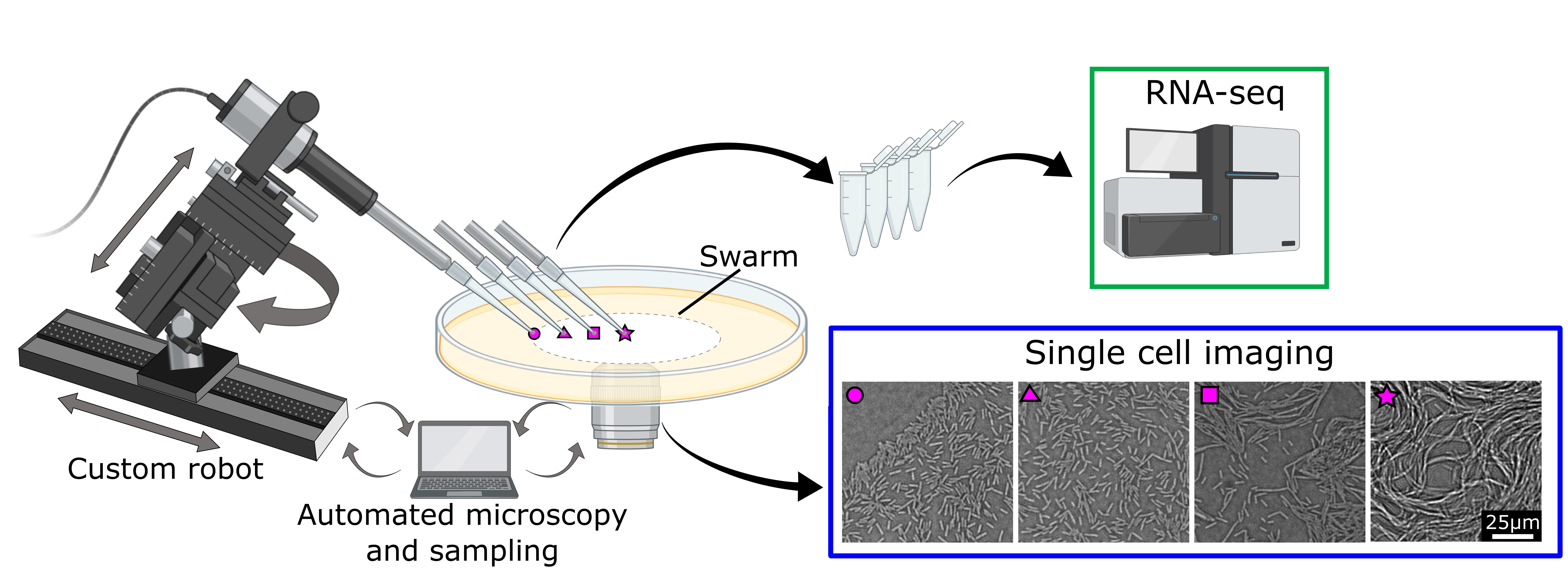
The story begins in spring 2018 with PhD student Hannah Jeckel, who had already established an adaptive light microscope for a previous project. To be able to acquire transcriptome measurements in addition to imaging, Hannah wanted to integrate an automated sampling process into this adaptive microscope. She teamed up with Konstantin Neuhaus, an expert in microscopy and robotics, as well as Horst Henseling, whose expertise as the leader of the mechanical workshop was invaluable to construct the experimental apparatus. Together, they embarked on a year-long journey to design, construct, test, re-design, modify, partially replace, test again (…) a robotic arm coupled to the adaptive microscope to extract cells in synchrony with imaging, while preserving the integrity of the swarm. When they eventually succeeded, it was time to tackle the next question – how can we obtain transcriptome data from collected cells?
There are a variety of standardized protocols and kits to preserve and extract mRNA from biological samples. Unfortunately, none of them really works when you have only a couple of thousand cells collected by our sampling robot. This could have been the end of the project, if we did not have an ace up our sleeves: Newly-arrived postdoc Kazuki Nosho, whose knowledge and creativity surrounding RNA-work was exactly what was needed to bring the project to the next stage. And the hard work was far from over, as new challenges arose to integrate the requirements of low-biomass RNA-seq into our setup. How can we ensure that collected cells are dropped into an ice-cold buffer when the experiment is taking place in a 30 °C environment? How do we increase the collected biomass and, equally important, accurately estimate the number of collected cells to enable necessary corrections during post-processing? We optimized our sample-collection tip design, introduced pre- and post-imaging of the sampled area, tested and re-tested the sampling process. It turned out that one person monitoring the experiment and replenishing materials such as collection tips and sample tubes was not sufficient, so Hannah and Kazuki squeezed into the cramped microscope room, carefully optimizing each step and coordinating respective workflows. Weeks were spent plating the collected cells and comparing results with the imaging-based estimate. When things stopped working, we suspected the weather and cursed the inhomogeneous airflow of the air conditioning. Eventually, having spent another year on optimization, we felt ready to start our final sampling in the early spring of 2020. Which coincided perfectly with the Covid-19 pandemic and lockdowns.
Planning and performing experiments during a pandemic came with new and unanticipated challenges. Interruption of in-person work was followed by a series of shortages of different materials, particularly for RNA work, putting new pressure on making each experiment count. Working in close proximity was no longer an option, so we re-calibrated our workflow, passing samples and materials from a safe distance. We moved our schedule to work in a shift-like manner, one of us arriving well before sunrise to inoculate plates, while the other one stayed into the late evening to prepare all components for the next day’s experiment. Our goal to collect samples from six independent replicates seemed easy enough to achieve at first, but unexpected complications – a software refusing to start (or worse, starting and then stopping to work mid-experiment), contamination in our overnight culture, a set of tips failing to collect the sufficient number of cells or a tired PhD student dropping carefully prepared components to the floor – meant that the final sampling phase spread over several weeks. Eventually, we made it work, and we can only imagine each other’s triumphant smiles behind the masks on the day of our final experiment. Of course, this was before we knew the struggle of analyzing the data and making sense of the results to enable the formulation – and testing! – of hypotheses.
Looking back today, another 3 years later, we can recognize how time, resources, communication and combining the diverse skill sets of people involved were key to success. And we believe that the phenotype of cross-generational sharing of metabolites in space and time that we ended up discovering in this project teaches us yet another important lesson about successful communities: That sharing not only occurs between the people we interact with right now, but also between generations. In the swarms we were studying, a first generation of surface explorers consumed the preferred carbon sources malate and succinate, but they left another resource, pyruvate, behind, which served as a nutrient for subsequent cells. In the same manner, we benefit from the knowledge past generations of researchers have accumulated and taught us, but we ourselves also create new resources to share. These can be datasets, computational tools, shared protocols as well as simply ideas and inspiration. It is important to recognize that, much like microbes, we as researchers rely on our communities to be successful, and that the best strategy to collectively thrive is often to combine knowledge and expertise across disciplines, with each one of us playing to our strengths.
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
Very nice work showing the generation level selection for carbon source . Congrats whole team involved.