Multicopy plasmids accelerate evolution in bacteria
Published in Ecology & Evolution
I´ve always been intrigued by the role small plasmids play in bacterial ecology and evolution. Plasmids can be generally divided into large plasmids (ca. > 15 kb) that have relatively few copies per bacterial cell, and small plasmids (ca. < 15 kb) that have many copies per cell. In general, it is accepted that plasmids are crucial for bacterial evolution because they are able to transfer accessory genes between bacteria by horizontal gene transfer (HGT). Large plasmids are frequently able to self-transfer by conjugation and they have been extensively studied over the last few decades because of their important role in the spread of antibiotic resistance genes among pathogenic bacteria. Small multicopy plasmids, however, are not able to self-transfer, so they require a conjugative plasmid or a phage to help them spread. Despite their limited ability of horizontal transfer small plasmids are extremely common in prokaryotes. The ubiquity and high frequency of small multicopy plasmids in bacteria has always puzzled me. How can they be that prevalent? Are they conferring any extra evolutionary advantages to bacteria apart from HGT?
One particularly exciting illustration of how small multicopy plasmids seem to play additional roles to HGT in bacteria is the case of endosymbiont bacteria living inside eukaryotic cells. Some of these endosymbiotic species such as Buchnera aphidicola completely lack HGT, however they carry multicopy small plasmids coding for essential genes for their symbiotic association. This observation made me think of the different possible advantages associated with carrying a gene on a multicopy plasmid. For example, carrying a gene on a plasmid with multiple copies produces higher levels of expression, and this may be beneficial for the host. Another possible advantage is the variation in plasmid copy number existing among single cells, which generates heterogeneity in gene expression between plasmid-bearing bacteria, leading to potentially beneficial phenotypic plasticity. Finally, the most tantalizing possibility is that multicopy plasmids could accelerate the evolution of the genes they carry by increasing their mutation rate due to the increase in copies per cell.
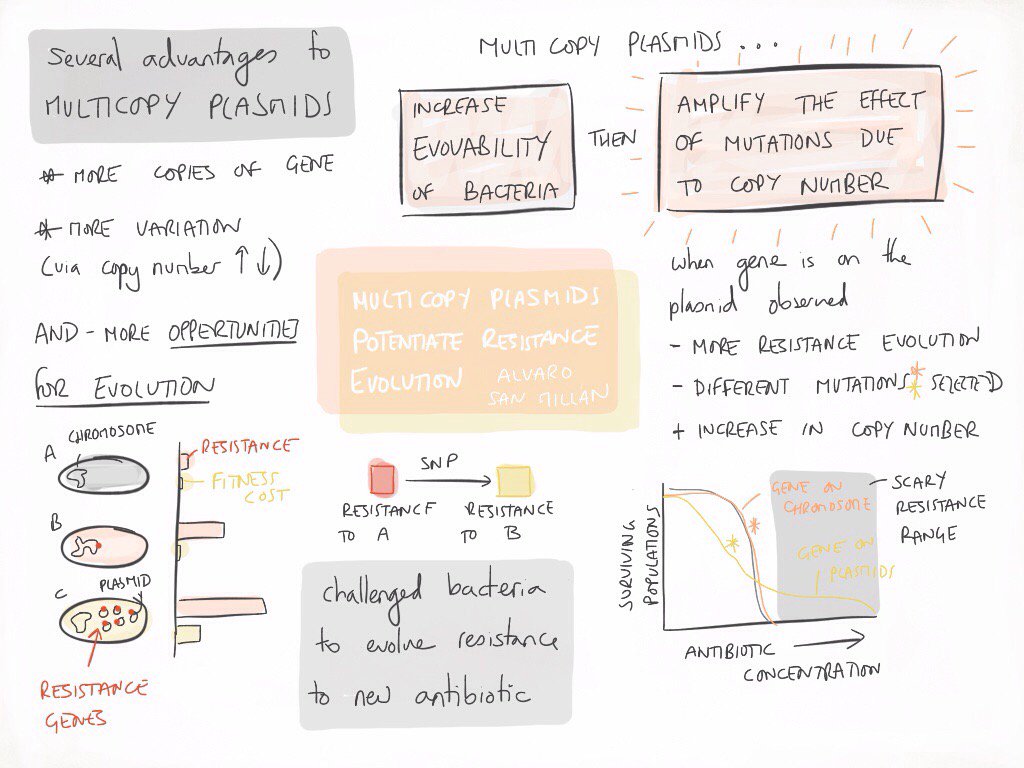
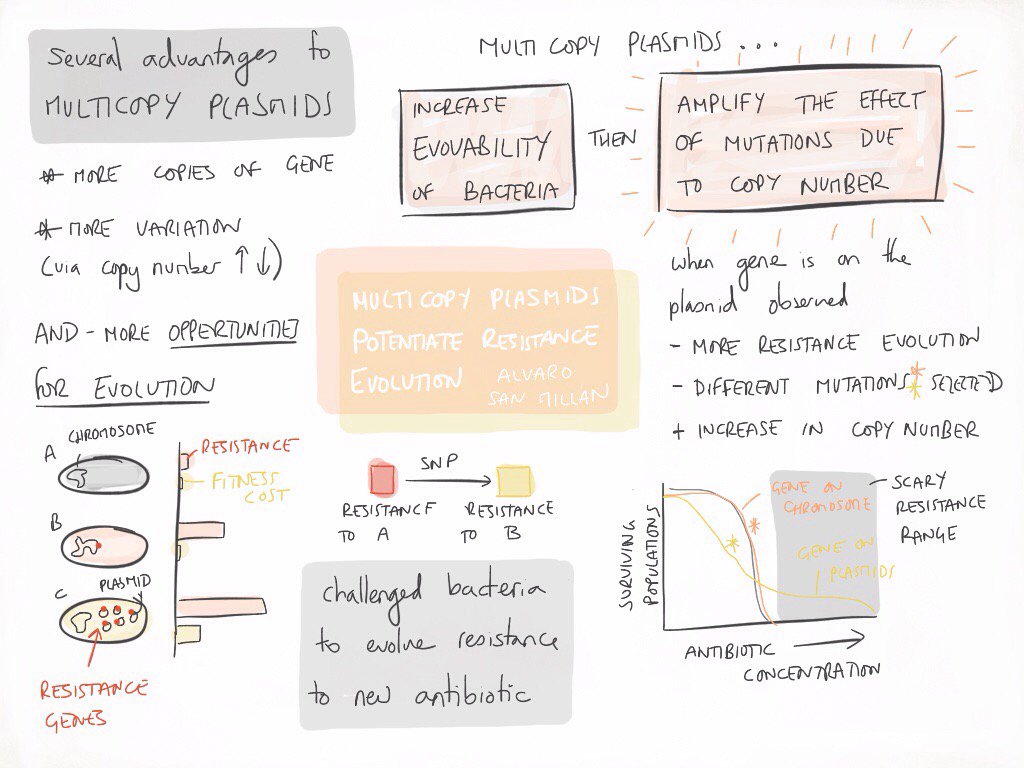
We decide to experimentally test what are the advantages for the host bacteria when a gene is located on a multicopy plasmid instead of on the chromosome. To answer this question we constructed a model system using Escherichia coli carrying the β-lactamase gene blaTEM-1 either on the chromosome or on a multicopy plasmid with approximately 20 copies per cell (pBGT). We analysed the advantages and disadvantages associated to each gene location in terms of antibiotic resistance and fitness levels, and we use experimental evolution and whole genome re-sequencing to investigate the evolvability of ceftazidime resistance in the different populations carrying blaTEM-1 (see attached image, courtesy of Ellie Harrison).
Our results demonstrate that multicopy plasmids provide an evolutionary advantage above and beyond mediating HGT. Over the short term, carrying blaTEM-1 on a multicopy plasmid is associated with an increase in fitness under conditions of strong selection for beta-lactam resistance. Over the long term, plasmid pBGT acts as evolutionary catalyst that facilitates the evolution of novel variants of blaTEM-1 and allows bacterial populations to evolve clinically relevant levels of ceftazidime resistance. The plasmid accelerated resistance evolution by a dual effect: (i) increasing the rate of appearance of novel TEM-1 mutations conferring resistance to ceftazidime first, and then (ii) amplifying the effect of TEM-1 mutations due to increased gene dosage. Our study challenges the established idea of plasmids being simple vehicles of HGT by demonstrating the role of multicopy plasmids as hotbeds gene evolution.
The article in Nature Ecology & Evolution can be found here: http://go.nature.com/2fxSwk6




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in