Mutations in the splicing factor SF3B1 are linked to frequent emergence of HLA-DRlow/neg monocytes in lower-risk myelodysplastic neoplasms
Published in Cancer and Biomedical Research
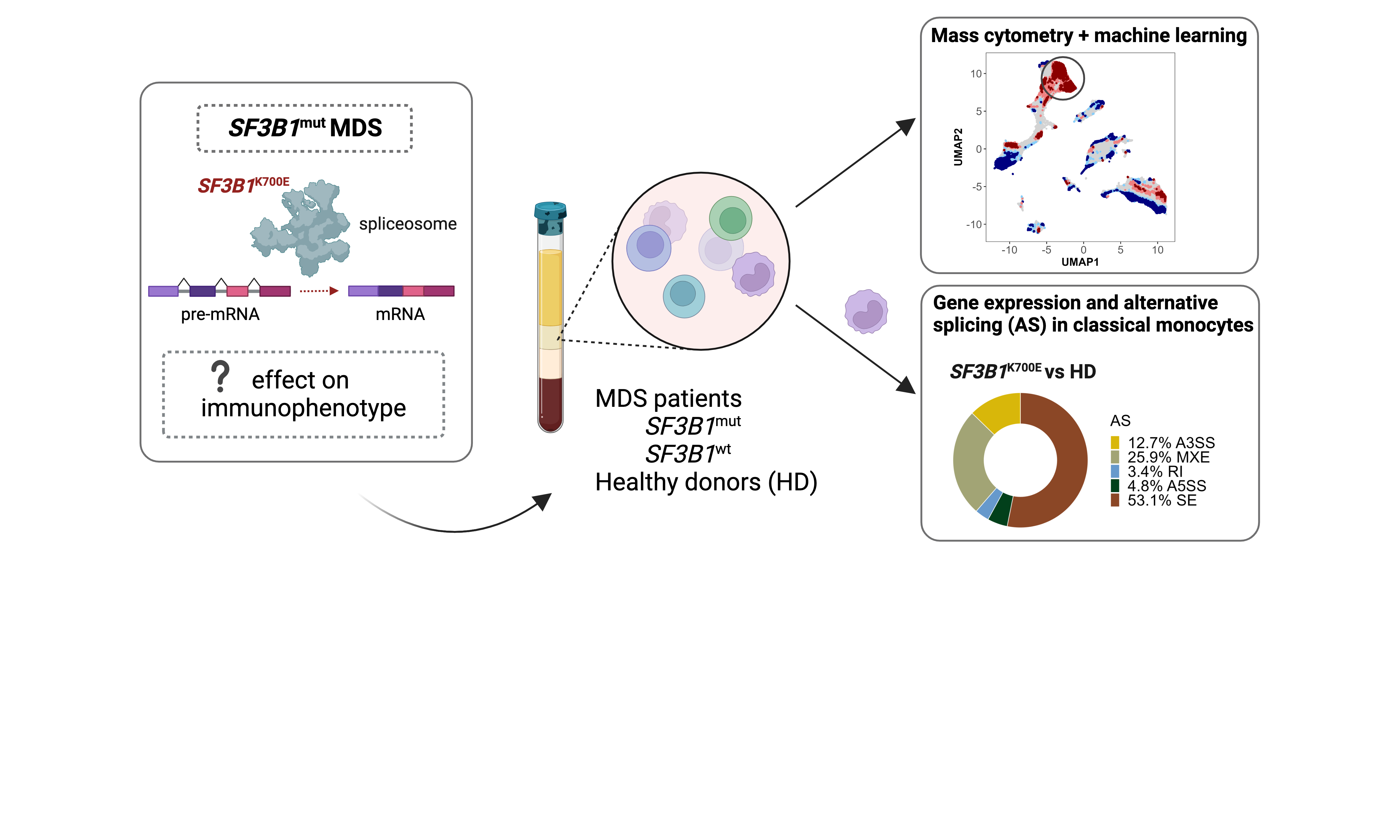
Myelodysplastic neoplasms (MDS) are a group of hematologic malignancies characterized by genetic abnormalities in hematopoietic stem and progenitor cells (HSPCs), dysfunction of blood cell production and maturation in the bone marrow, and subsequent peripheral cytopenias. Manifesting with symptoms such as fatigue, weakness, susceptibility to infections, and easy bruising or bleeding, MDS poses a significant health challenge. MDS can also progress to a more severe form of blood cancer, acute myeloid leukemia (AML). It primarily affects the elderly population, and due to aging demographics and improved diagnostic capabilities, the incidence of MDS is expected to increase significantly in the coming years. Treatment for MDS aims to improve blood cell maturation and counts, manage symptoms, and prevent progression to leukemia. The only potentially curative treatment is allogeneic hematopoietic cell transplantation (alloHCT). As research continues to advance, improving patient stratification and identifying novel therapeutic strategies remains paramount in enhancing outcomes for MDS patients.
Approximately half of MDS patients carry somatic mutations in splicing factor genes, with SF3B1 (splicing factor 3b subunit 1) being the most commonly mutated gene. SF3B1 mutations are associated with specific characteristics such as ring sideroblasts (RS), higher white blood cell counts and lower bone marrow blasts. Recent updates in MDS classification recognize cases with mutated SF3B1 as a separate entity with better clinical outcomes1,2. These mutations disrupt splicing of key genes throughout erythroid differentiation. In terms of treatment, patients with SF3B1 mutations may respond well to luspatercept, a medication that helps red blood cells mature. However, they may not respond as well to immune-suppressive treatments3,4. Understanding the immune system changes triggered by SF3B1 mutations remains an area of ongoing research.
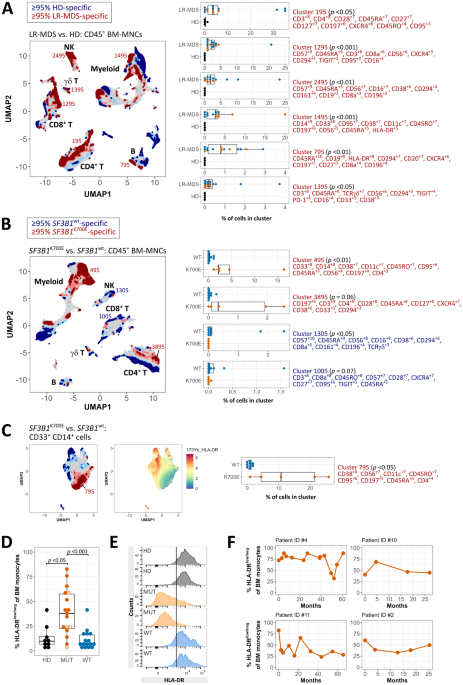
Our study was motivated by the need to bridge the gap in understanding how SF3B1 mutations affect the immune system in MDS patients. The primary goal was to conduct a comprehensive investigation of immune cell phenotypes in the bone marrow. Utilizing a machine learning workflow for the analysis of high-dimensional single-cell mass cytometry (CyTOF) data, we identified a prominent cluster of HLA-DRlow/neg classical monocytes in SF3B1-mutant lower-risk MDS. This discovery was validated through retrospective analysis of diagnostic flow cytometry data and confirmed in independent patient cohorts, showing an increased frequency of HLA-DRlow/neg monocytes in SF3B1-mutant compared to other MDS patients and healthy controls. Notably, we found a strong correlation between the frequencies of HLA-DRlow/neg monocytes in the bone marrow and peripheral blood of patients. Furthermore, our results showed that patients with a higher prevalence of HLA-DRlow/neg monocytes in the bone marrow maintained consistently elevated levels over time. The significance of our findings lies in the potential immunoregulatory properties associated with HLA-DRlow/neg monocytes, including the inhibition of effector T-cells, decreased antigen presentation, and defective dendritic cell maturation.
Moreover, to gain further insight into the transcriptional changes linked to SF3B1 mutations, we performed RNA sequencing on classical monocytes isolated from patients with an isolated SF3B1K700E hotspot mutation (data are publicly available via GEO accession number GSE236535). Our analysis revealed downregulated expression of genes involved in inflammatory cytokine signaling, as well as differential mRNA splicing of various genes involved in the regulation of the defense response and cytokine signaling (e.g., MAP3K7), mRNA metabolism, apoptotic signaling, and mitotic cell cycle.
Despite these alterations, classical monocytes with a heterozygous mutation in SF3B1 demonstrated adequate responses to in vitro lipopolysaccharide (LPS) stimulation, indicating their capability to secrete pro- and anti-inflammatory cytokines essential for initiating the host defense. However, further research with larger sample sizes is imperative to elucidate the functional and stimulus-dependent deficits resulting from dysregulated expression and/or mis-splicing of the identified genes.
Importantly, future studies comparing HLA-DRlow/neg and HLA-DRhigh classical monocytes from patients with an isolated SF3B1K700E mutation will help to clarify their respective roles in the inflammatory process. Our ongoing commitment to deciphering the immune landscape of MDS holds promise for advancing personalized therapeutic approaches and improving patient outcomes in this challenging disease paradigm.
The header image was created in BioRender.com.
1 Khoury JD, Solary E, Abla O, Akkari Y, Alaggio R, Apperley JF et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/Dendritic Neoplasms. Leukemia 2022; 36: 1703–1719.
2 Arber DA, Orazi A, Hasserjian RP, Borowitz MJ, Calvo KR, Kvasnicka HM et al. International Consensus Classification of Myeloid Neoplasms and Acute Leukemias: integrating morphologic, clinical, and genomic data. Blood 2022; 140: 1200–1228.
3 Stahl M, DeVeaux M, De Witte T, Neukirchen J, Sekeres MA, Brunner AM et al. The use of immunosuppressive therapy in MDS: Clinical outcomes and their predictors in a large international patient cohort. Blood Adv 2018; 2: 1765–1772.
4 Zhang Q, Haider M, Al Ali NH, Lancet JE, Epling-Burnette PK, List AF et al. SF3B1 Mutations Negatively Predict for Response to Immunosuppressive Therapy in Myelodysplastic Syndromes. Clin Lymphoma Myeloma Leuk 2020; 20: 400-406.e2.
Follow the Topic
-
Leukemia
This journal publishes high quality, peer reviewed research that covers all aspects of the research and treatment of leukemia and allied diseases. Topics of interest include oncogenes, growth factors, stem cells, leukemia genomics, cell cycle, signal transduction and molecular targets for therapy.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in