Of cats and men
Published in Ecology & Evolution

Cats are the stars of the internet world. Today cats are everywhere, but how did they get there? Little is known about their conquest of the world and of their domestication - and I think most cat owners agree that it isn’t even clear whether they are really domesticated!
Most of what we knew comes from ancient Egypt, in particular from the iconography of Pharaonic Egypt where the cat had become sacred, an embodiment of the goddesses Bastet and, later, Isis. This embodiment was not unusual, however, since most animals in ancient Egypt had a divine side to them.

Cats in archaeological contexts are rare and the find of a cat in a child burial in Neolithic Cyprus, where no cat remains had been found prior to the arrival of the first farmers, suggested that they had transported this cat to the island, meaning that it probably had been tamed. This would hint to a long, at least 10,000-year-old, relationship between cats and humans, but this is still enigmatic. We knew only that cats were an important asset of early societies in controlling rodent pests and venomous snakes and scorpions. We also knew that all modern domestic cats share the mitochondrial DNA lineage with the wildcats from Northern Africa and the Levant.
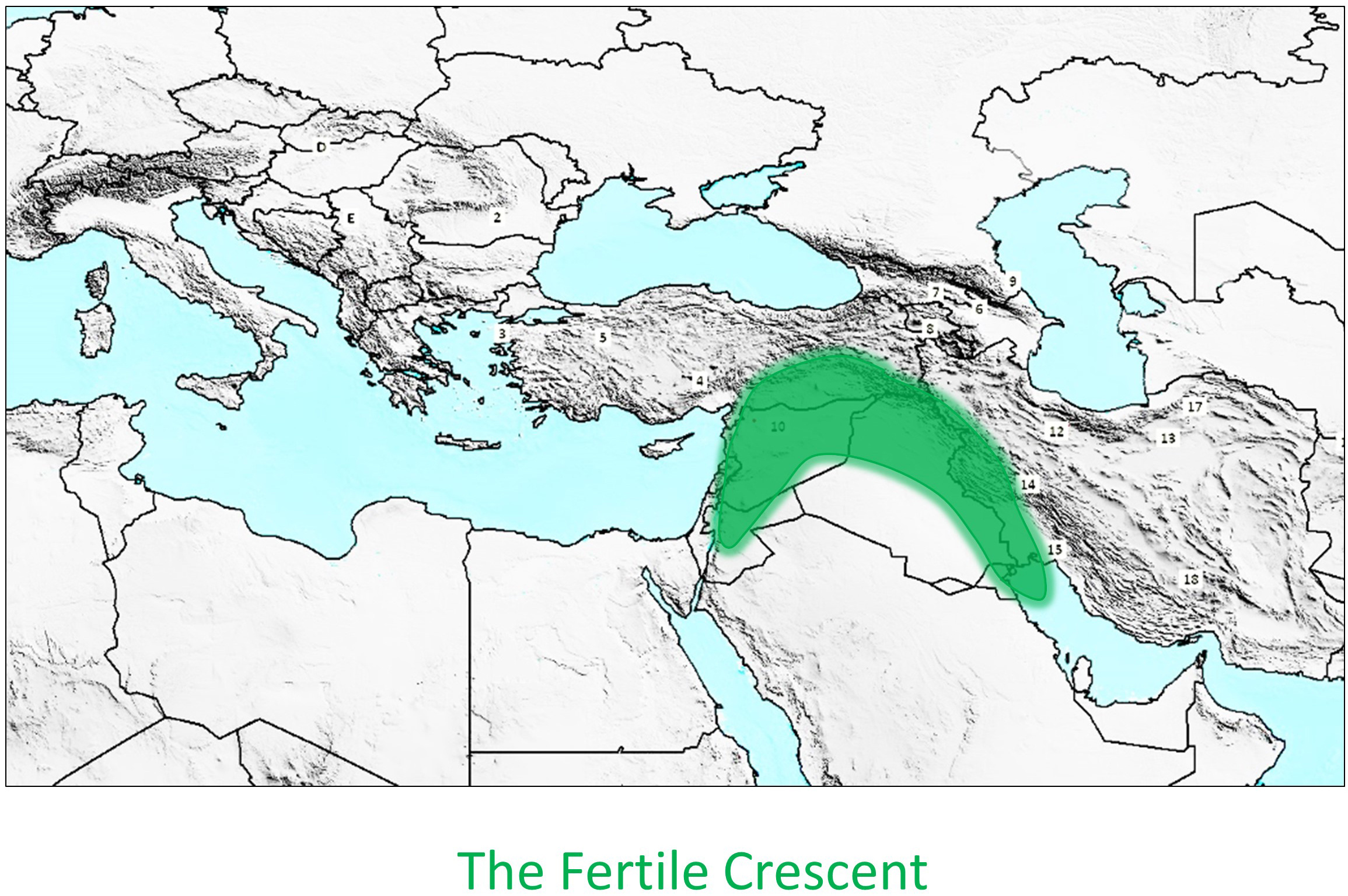
Needless to say that as a palaeogeneticist interested in animal domestication, I found this situation both intriguing and challenging. As such, I was enthusiastic when, years ago, my colleague Wim van Neer, an zooarchaeologist from the Royal Belgian Institute of Natural Sciences who had excavated cat skeletons from an elite cemetery in predynastic Egypt, proposed at a meeting in the United Emirates that we team up and tackle the poorly known cat domestication process using a paleogenetic approach. The start of the project, however, was delayed due to lack of funding, a perennial problem in archaeological research. Happily, a few years later Wim and his colleague Ronny Decorte from Leuven University obtained funds to hire a young colleague, Claudio Ottoni, for our project. The project was finally able to get going. Wim, Bea de Cupere and I first gathered cat samples by activating our networks of zooarchaeologists. Claudio came to the lab that I lead with Thierry Grange and together, the three of us worked out the most efficient strategy to maximize results from as many samples as possible. Since our group had been working for many years on retrieving DNA data from archaeological samples from the heart of the animal domestication zone, the Fertile Crescent in the Near and Middle East (see map), we knew this was going to be a challenging task. Not only the cat remains were old, but they were not cold! Indeed, DNA preservation correlates in part to temperature and is less well preserved in hotter climates, where obtaining results is more difficult. Furthermore, cat bones are small and fragile and contain only few small compact regions, if any, where DNA preserves. To make things even more challenging, there was only one large-scale study to date, performed by Driscoll and colleagues in 2007 who analyzed present-day domestic and wild cats throughout the world. We decided to follow their path and collect data from at least a hundred ancient samples to get a global view of the cat domestication process.
Since we had limited funding we needed to make it cost-efficient. We decided to use multiplex PCR to meet the challenge, but we had to do it in a way that it would work with poorly preserved DNA, which means targeting very small DNA fragments and ensuring that each PCR amplifies DNA even when only a few molecules are preserved. We targeted all the informative SNPs that were responsible for the topology of Driscoll et al.’s phylogenetic tree based on several kilobases of the mitochondrial genome. We systematically designed primer pairs and used robotic quantitative PCR to select those that performed well. After a second round of bioinformatic analysis and quantitative PCR, primers could be combined in multiplex PCRs without losing too much of their sensitivity. To sequence these short PCR products, we used at the time pyrosequencing, an approach we had successfully implemented in the course of our cattle domestication project that was carried out in the lab by Silvia Guimaraes. She initiated Claudio into the intricacies of the experimental procedure and we collected the first results using this approach. And it worked! We obtained data from about half of the ancient samples we had analyzed and were able to put them within the framework of the large-scale modern study.

Of course, some samples were more difficult than others, e.g., Claudio had to crush a lot of bones from Egyptian cat mummies before he could get some to work (6 out of 74!). The strategy was good enough, however, to make the project a success. Due to funding requirements and difficulties, Claudio had to go back to Leuven and work on other projects as well. Meanwhile in Paris, we also worked on other projects and introduced next-generation sequencing and bioinformatics in the lab. In particular, Thierry Grange along with Julien Daligault and Mélanie Pruvost developed a high-throughput strategy to quickly and economically collect multiplex PCR data from ancient and degraded samples. We had the workflow working when Claudio came back with extracts from another ca. 100 bones that Wim and Bea had collected in the meantime. In the lab, Claudio and Julien worked frantically to gather data from all these samples. This was a highly productive harvest of data that amazingly took place in less than 2 months. And the isolated jigsaw pieces suddenly yielded a picture! We could see these ancient cats moving all over the planet! We could see the ancient Egyptian cats becoming very popular in the Classical Era as they spread all over the ancient world at the time of the “Pax Romana” and later even jumped on Viking drakkars (longships). We could monitor in “real-time” the arrival of Indian cats in a Roman harbor at the Red Sea in Egypt. We saw that trade and raiding routes, documented in the historical records, turned into dispersal motorways for cats. The cats that now conquer the world using the information superhighways of the internet were simply expanding into a new ecological niche: in the ancient world they were already using the fastest person-to-person connections in existence. It couldn’t be clearer that domestication is a path for evolutionary success, and cats were clever enough to achieve this while sacrificing a limited amount of freedom and self-determination, in contrast to most other domesticates - except maybe mice and rats!
Our paper in Nature Ecology & Evolution can be found here.

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in