Phase Separation of Scaffold Protein Regulates Microbial Asymmetric Cell Division
Published in Microbiology
By polarizing different cell fate determinants at opposite cell poles, asymmetric cell division that produces distinct daughter cells is an evolutionarily conserved mechanism to generate cellular diversity in both eukaryotes and prokaryotes.
As a well-established model to study bacterial asymmetric cell division (ACD), Caulobacter crescentus produces a motile swarmer cell and a sessile stalked cell during each cell cycle. In the pre-division cell stage, the polar localization of two distinct membraneless signaling complexes, particularly the phosphatase PleC and the kinase DivJ, coordinates to modulate the phosphorylation levels of a set of downstream signaling proteins (including the master regulator CtrA) and determinate the cell fate of C. crescentus. However, the biomolecular basis and regulatory mechanism of the polar signaling complexes remain largely unclear.
Scaffold proteins are known to physically tether client proteins to specific cellular areas, functioning in spatial regulation of biological processes including signaling transduction, cytokinesis, morphogenesis, and ACD. In C. crescentus, the kinase DivJ is recruited to the stalked cell pole (old cell pole) through a PopZ-SpmX-DivJ sandwich, while the phosphatase PleC is localized to the swarmer cell pole (new cell pole) by the PodJ scaffold.
Recently, a research team led by Wei ZHAO from the Shenzhen Institutes of Advanced Technology (SIAT) of the Chinese Academy of Sciences proposed that the PodJ scaffold in the new cell pole forms biomolecular condensates with physiological functions via phase separation, which helps to establish and regulate the asymmetry of bacterial cells.
The study was published online in Nature Communications on November 23, 2022. (https://www.nature.com/articles/s41467-022-35000-2)
The researchers found that phase separation plays an essential role in the C. crescentus PodJ-signaling hub assembly. Both the coiled-coil 4-6 region and the intrinsically disordered region (IDR) in PodJ functionally contributed to generating biomolecular condensates and forming the PodJ-client complexes. Moreover, a negative regulation of PodJ phase separation by the old-cell-pole scaffold protein SpmX was observed. SpmX inhibited PodJ condensate formation and impeded its cell-pole accumulation and client recruitment (Figure 1).
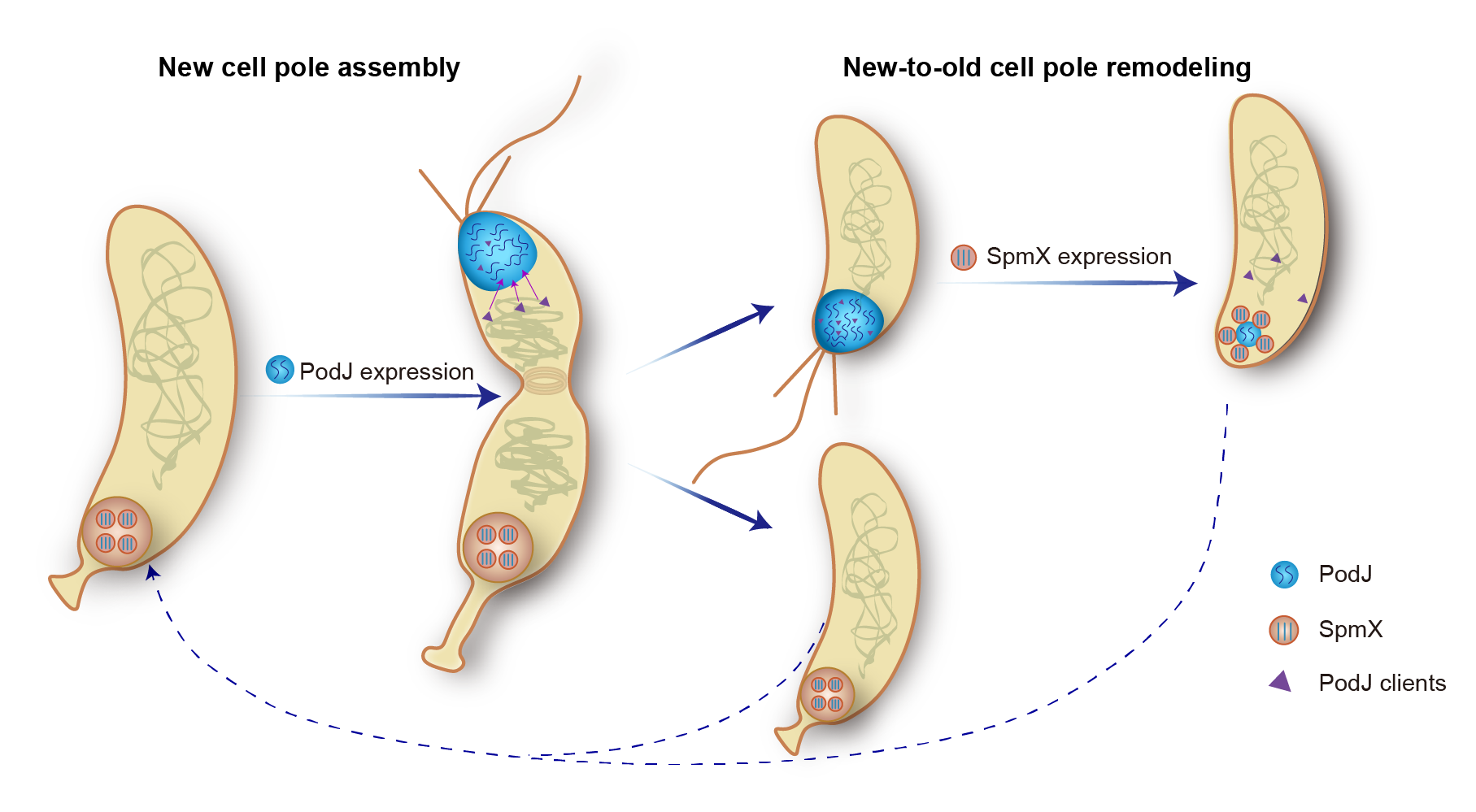
Figure1. Schematic of PodJ phase separation and the proposed model for regulating bacterial asymmetric cell division
The results revealed that phase separation modulates the assembly and dynamics of scaffold-signaling hubs in C. crescentus. Phase separation may serve as a general biophysical approach for assembling scaffold-signaling complexes and regulating ACD. Similar methods could be employed for the rational engineering of artificial organelles and other membraneless biocatalytic compartments.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Healthy Aging
Publishing Model: Open Access
Deadline: Dec 31, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in