Precision mouse model for patients with gallbladder cancer that enables analysis of local tumor immune responses
Published in Cancer
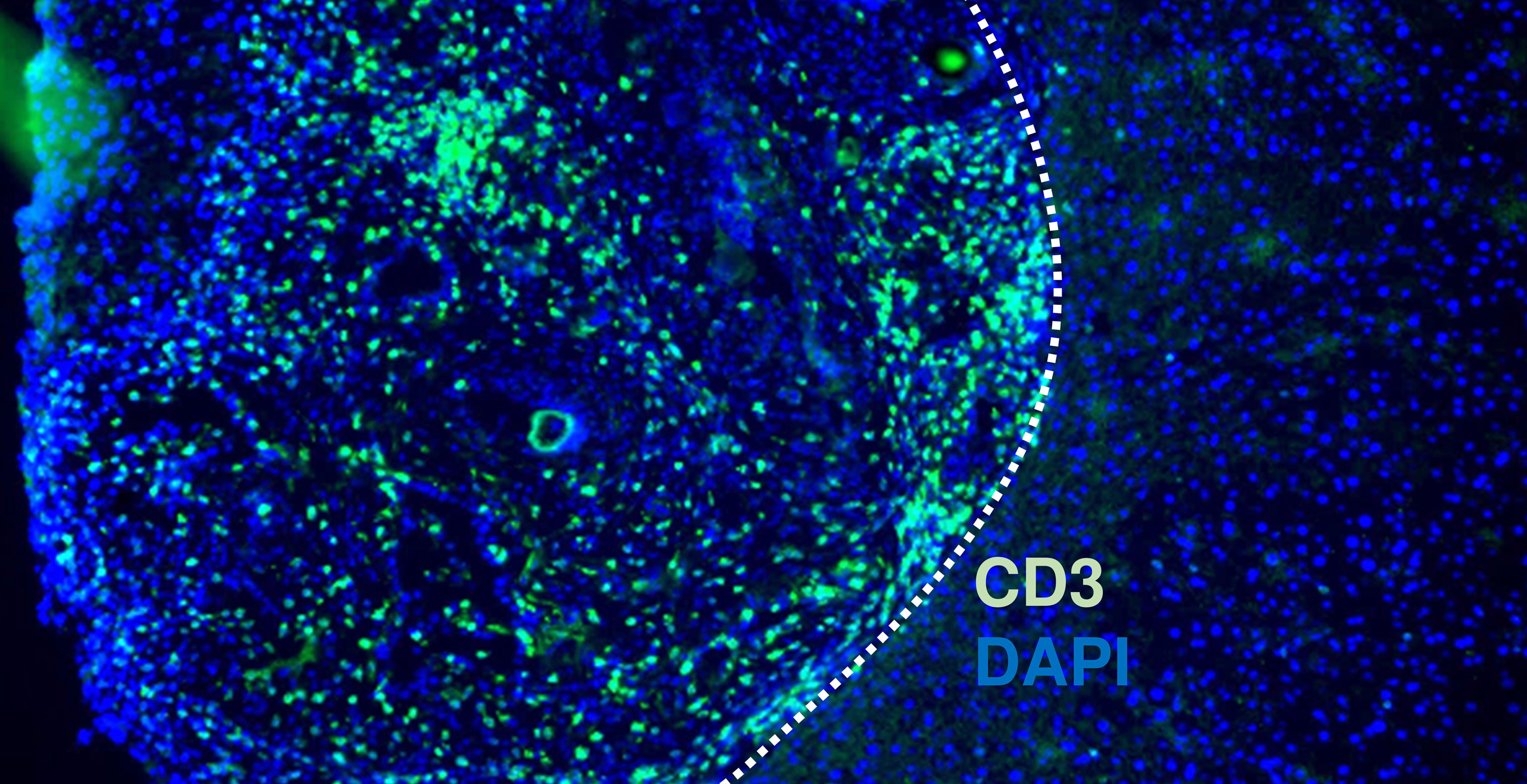
We developed a new mouse model that can mimic not only cancer development but other biological characteristics of cancer patients as well. Given the poor prognosis and availability of few models due to the lack of gall bladder (GB)-specific Cre mouse strain, we selected the GB as a target organ for the proof-of-concept study.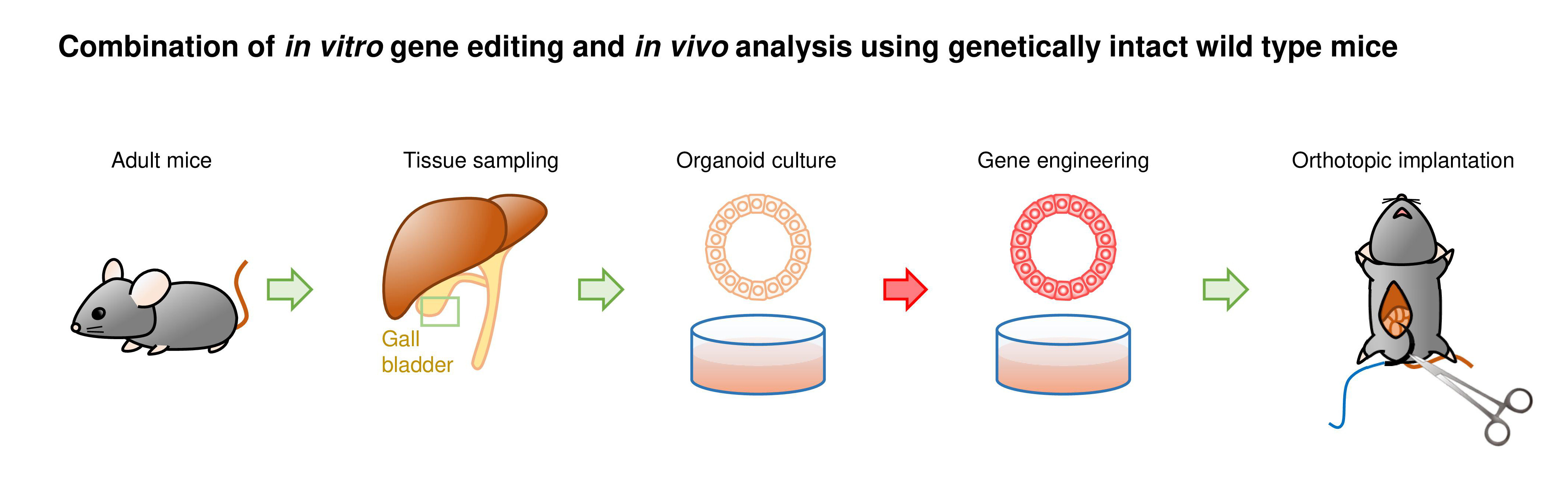
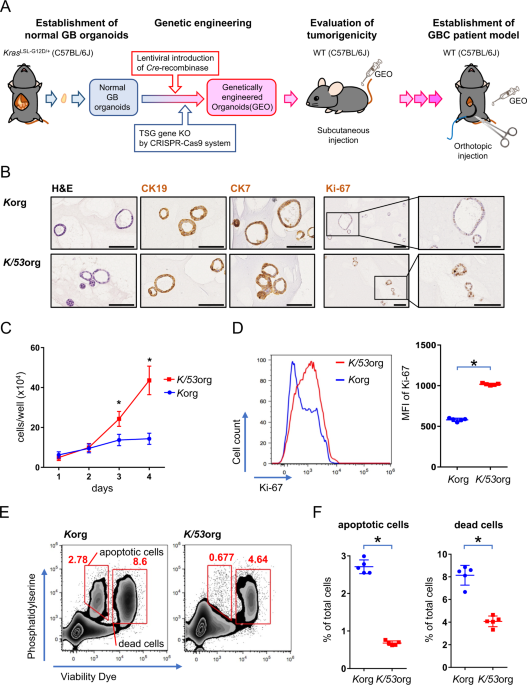
We first generated GB organoids with mutant Kras and Trp53 loss by lentiviral Cre transduction and CRISPR/Cas9 gene editing, respectively (hereafter referred to as K/53org). We aimed to develop orthotopic tumor models via intra-GB injection. However, with K/53org resuspended in Matrigel, injection into the GB lumen of syngeneic mice retained the Matrigel in a liquid state, allowing organoids to readily and freely flow out into the lower bile duct. To avoid such leakage, we ligated the GB neck with a thread prior to injection, which retained the organoids inside the GB. However, all mice eventually died due to stenosis-induced lethal cholecystitis. Subsequently, we ceded the development of an orthotopic GB cancer model using a single-step implantation approach.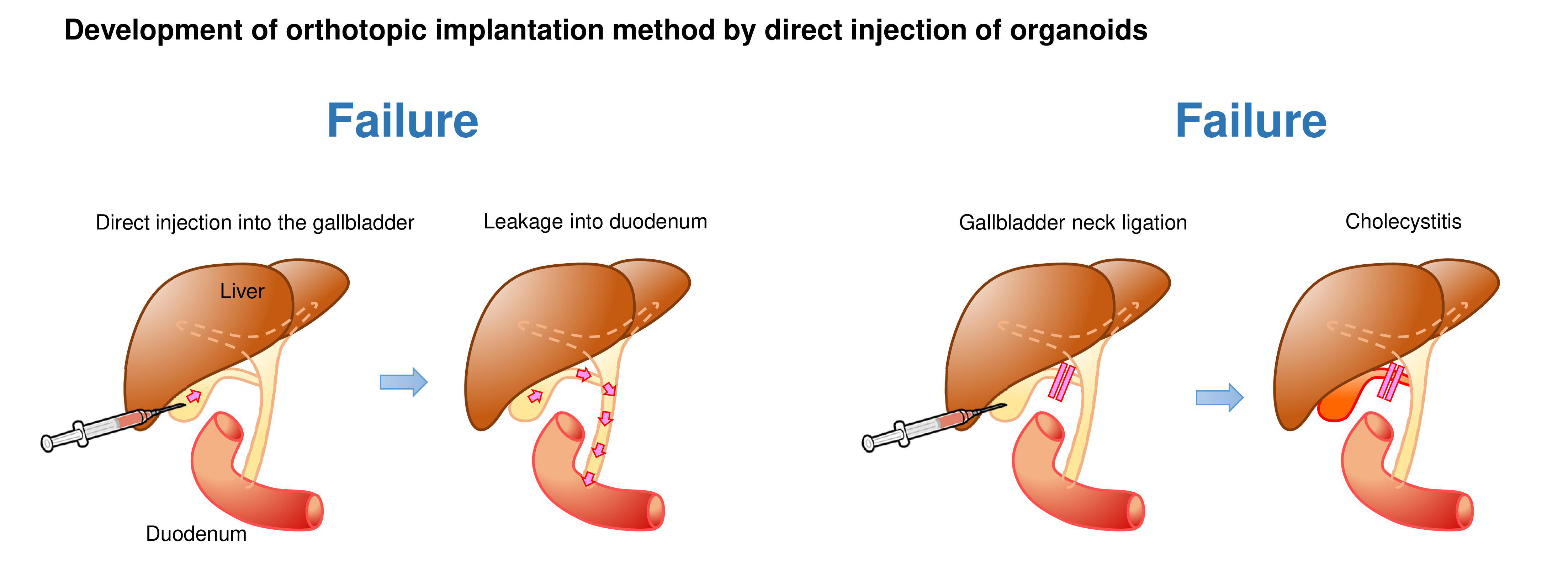
We then tested a two-step implantation approach in which K/53org-derived subcutaneous tumors were orthotopically implanted in syngeneic mice. Tumors were typically minced into 2-mm fragments, which we named “tumor buds (TB)”. Although a TB was initially inserted into the GB with a cut and closed tip, the TB remaining inside the GB did not develop tumors, presumably due to digestion by bile. To avoid direct exposure to bile, we fixed the TB outside the GB by suturing. This method, which we designated as the “implantation of organoid-derived tumor bud (IoTB),” was effective, allowing the transplanted TBs to grow into solid tumors that resembled human GB cancer.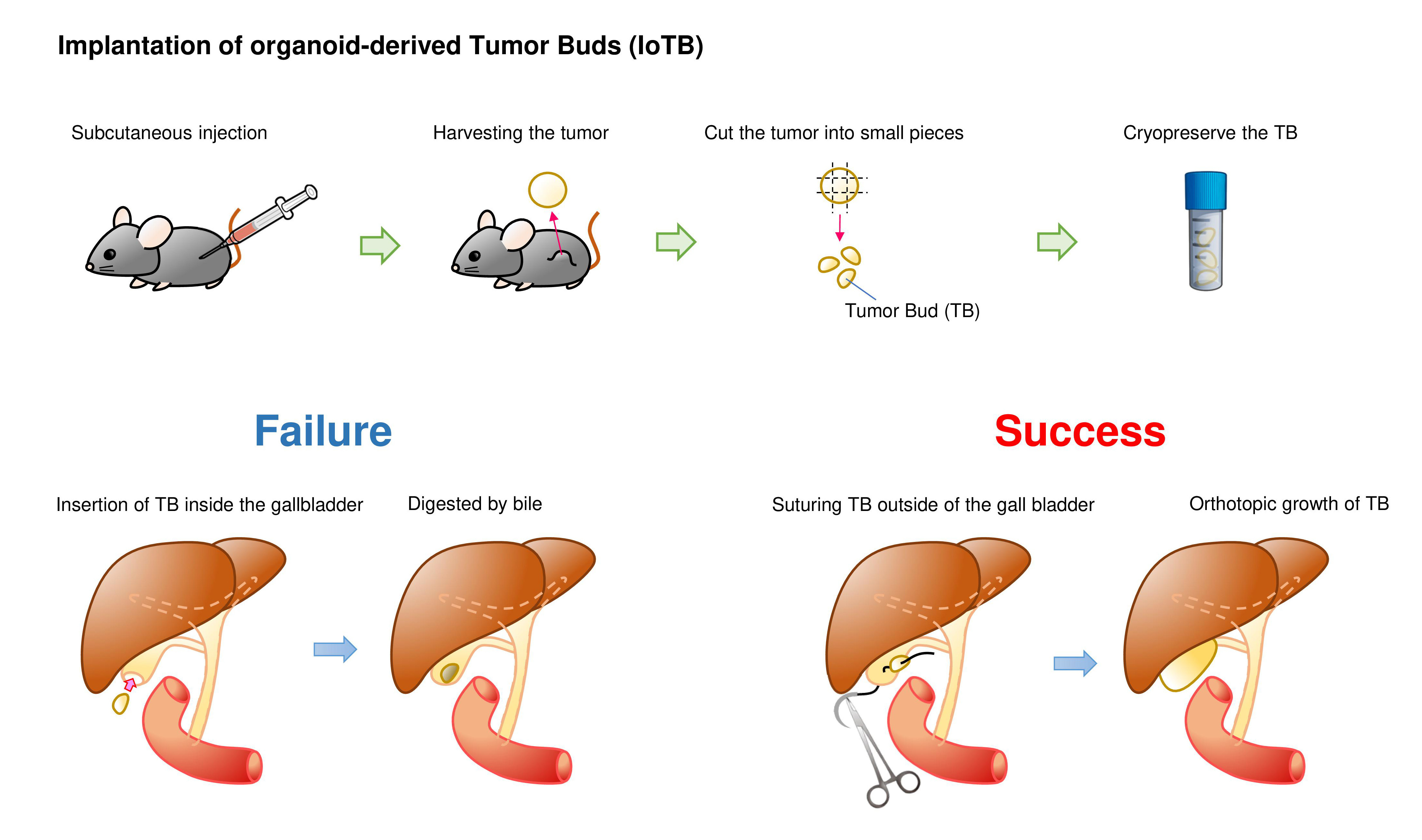
This IoTB model revealed the infiltration of similar subsets of immune cells in both subcutaneous and orthotopic tumors, confirming the physiological immune environment during carcinogenesis. In addition, the in vivo efficacy of gemcitabine, a common therapeutic agent for GB cancer, was accurately validated in large cohorts. Taken together, this model may serve as an avatar for patients with GB cancer in drug discovery and precision medicine. In addition, the application of this approach is warranted for other cancer subtypes.
Follow the Topic
-
Oncogenesis
A peer-reviewed open access online journal that publishes articles exploring mechanistic insight and molecular basis of cancer and related phenomena. It seeks to promote diverse and integrated areas of molecular biology, cell biology, oncology, and genetics.
Related Collections
With Collections, you can get published faster and increase your visibility.
Metabolic Reprogramming in Cancer
The field of cancer metabolism has expanded rapidly, revealing how metabolic reprogramming drives tumour progression, therapy resistance, and cellular adaptation. Beyond local interactions, accumulating evidence shows that circulating immune cells and blood‑borne factors—including metabolites, cytokines, and extracellular vesicles—actively influence tumour metabolic states. These systemic signals interact with tumour‑resident cells to shape metabolic plasticity, modulate survival pathways, and affect treatment responses.
Within the tumour microenvironment itself, metabolic interplay between cancer cells, immune cells, and stromal components remains a central determinant of tumour behaviour. Altered glycolytic flux, lactate‑driven microenvironmental changes, and cancer stem cell–associated metabolic adaptations contribute to tumour aggressiveness, immune evasion, and resistance mechanisms. Together, these local and systemic dimensions provide an integrated view of how metabolism underpins cancer progression.
This collection supports United Nations SDG 3: Good Health & Well-Being.
Topics of interest include:
- metabolic interactions within the tumour microenvironment
- cancer stem cell metabolism
- glycolytic pathways in tumour promotion
- tumour–immune cell metabolic interplay (local and systemic)
- interplay between circulating blood cells and tumour metabolism
- drug resistance and metabolic shifts
- protein expression in metabolic reprogramming
- role of transcription factors in metabolism
- signal transduction pathways affecting cancer metabolism
Publishing Model: Open Access
Deadline: Dec 31, 2026

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in