Prediction of response to neoadjuvant chemotherapy in ovarian cancer using an integrated machine learning framework for multimodal data
Published in Biomedical Research
What is the background?
High grade serous ovarian carcinoma (HGSOC) is frequently diagnosed at an advanced stage complicating treatment for a large proportion of patients. Neoadjuvant chemotherapy (NACT) followed by delayed primary surgery (DPS) is a common treatment strategy for patients with advanced HGSOC. Predicting response to NACT is difficult because of high levels of heterogeneity of HGSOC at multiple scales, both within patients and between patients,. Clinical response is variable and approximately 40% of patients may not benefit from NACT. Non-responding patients could benefit by being identified early as immediate primary surgery (followed by adjuvant chemotherapy) might be preferable over NACT and DPS. Predictors of response have mainly focused on single data types but integrated approaches may have higher predictive power. Although the value of integrated approaches are well documented for many cancer types, they have not yet been fully embraced for HGSOC, particularly because of the complexity of applying these approaches to multisite heterogenous disease. In addition, an integrated approach requires rich and well annotated data sets that are rarely available.
What we did
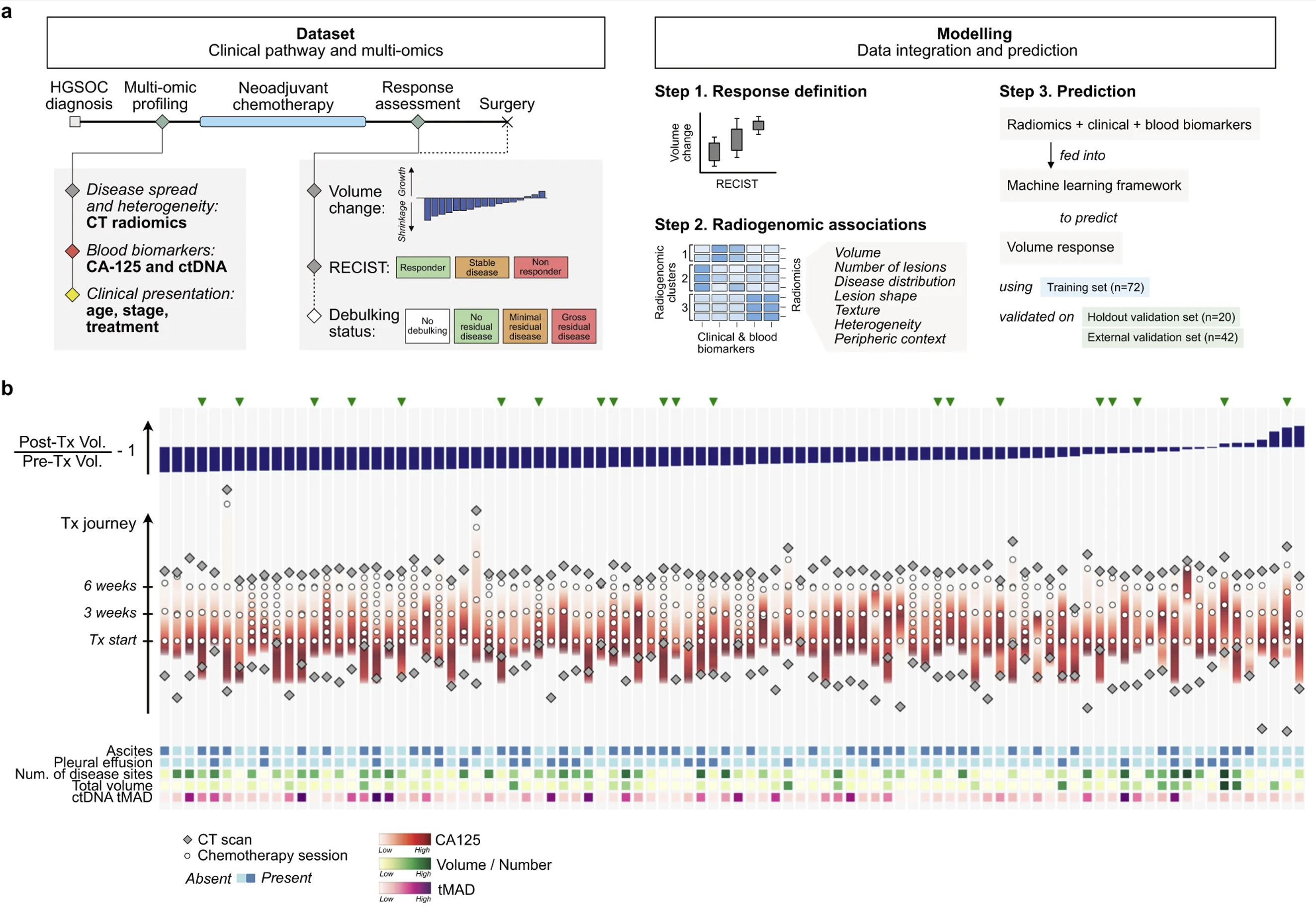
We put together two independent data sets with 134 patients in total (92 cases in the first data set, split into 72 for training and 20 as a hold out validation set; and 42 in a second independent test set). For all patients at baseline (prior to treatment), we obtained clinical data including demographics and details of their treatment and blood-based biomarkers such as CA-125 and circulating tumour DNA (ctDNA) and extracted quantitative features from computed tomography (CT) scans by manually segmenting all primary and metastatic tumour sites.
Our results
Tumour in the omentum and pelvic/ovarian locations (the most frequent locations forovarian cancer spread) accounted for the majority of the disease burden at baseline. Omental deposits showed significantly better response to NACT than pelvic disease.
Both, TP53 mutant allele fraction (TP53 MAF) assessed on ctDNA and CA-125 correlated with the overall disease burden prior to treatment but neither baseline CA-125 nor TP53 MAF correlated significantly with volumetric response. Baseline TP53 MAF correlated significantly with pelvic/ovarian disease volume measured both before and after chemotherapy but not with omental disease at either time point. This suggests that high TP53 MAF at baseline could be a specific indicator for high disease burden in the ovaries or pelvis, which tends to show poorer response.
Clustering based on the imaging features of the lesions revealed six clusters with distinct biological and clinical characteristics indicating that multi-lesion radiomics features of disease extent and heterogeneity might convey similar information as clinical and molecular tumour markers. For example, one cluster was associated with CA-125 levels and comprised mostly lesion volume features whereas clusters associated with ctDNA values were generally dominated by features quantifying lesion heterogeneity and context.
Our integrated framework is an ensemble of three machine learning pipelines, each of which includes feature selection steps together with a classifier (elastic net, support vector regression, or random forest). Models were trained using 5-fold cross-validation repeated five times with different seeds. We chose the relative change in total disease volume as the response metric and trained four models by successively adding clinical and molecular features: (i) age, FIGO stage, and treatment; (ii) CA-125; (iii) radiomics features; and (iv) ctDNA. After training was completed, the models were then tested on the hold-out and independent test sets. Only those models including radiomics features were able to produce response scores that were significantly correlated with the observed volume response. The addition of ctDNA values did not significantly improve performance levels. When analysing the relative contribution of different features, we found that CA-125 was frequently dropped from models containing radiomics features and lesion context. The most important lesion context represented features at the periphery of a lesion and was consistently selected. These findings highlight the importance of the including context features in models and the relative redundancy of CA-125 when these features are used.
In conclusion…
…We show that radiomics features are critical for predicting response to NACT and demonstrate site-specific patterns of differential response and correlation of radiomics features with ctDNA detection in ovarian cancer. We applied radiomics in the context of a robust ensemble-machine learning framework to heterogeneous real-world CT data and were able to integrate multimodal data to improve prediction performance. From a clinical perspective, the proposed framework addresses the unmet need for early identification of patients unlikely to respond to NACT who could be offered immediate surgery and could be applied to stratify patient risk in the context of clinical trials.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Healthy Aging
Publishing Model: Open Access
Deadline: Dec 31, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in