Programming protein switches through DNA strand displacement
Published in Chemistry
The paper in Nature Chemistry is here.
Inspired by amazing protein switches evolved by nature, synthetic biologists have sought to engineer synthetic versions to carry out novel and useful tasks for humanity. I often explain our protein switch work to non-experts as toying with Legos, except with protein blocks. Much like Legos, proteins provide an infinite space for creativity.
However, the reality is that piecing together proteins to reprogram their function is not as easy as snapping together Legos. Evolved through time, proteins are less like Legos and more like custom handcrafted wood pieces one finds at small artisanal stands. These “blocks” are carved to play an exact role in seamless connection with its partners. Often our group has rearranged and linked proteins with suboptimal initial results. Through endless tinkering, we often arrive at our desired solution, but these answers are case-by-case and not generalizable.
The unpredictability can be extremely frustrating, and for this reason, our group began exploring the benefits of nucleic acids. The predictable nature of Watson-Crick base pairing makes DNA a much more “Lego-like” material. Early work led us to create DNA scaffolds for docking enzyme cascades and to design molecular beacons for sensing. The first breakthrough towards this current study came when Daniel Blackstock (my co-first author) developed a technique to covalently label and purify protein-DNA conjugates by utilizing the HaloTag protein. This enabled us to link any protein of interest with any DNA sequence of interest. We could circumvent our previous use of zinc finger proteins, which have lower affinities than hybridization can provide and are a hassle to engineer to different binding sequences. The biggest breakthrough for the dynamic nature of our strategy came when my advisor, Wilfred Chen, attended a toehold-mediated strand displacement presentation. He realized that DNA could not only serve as a static scaffold, but as a fluid and dynamic medium that could be wired into complex circuits.
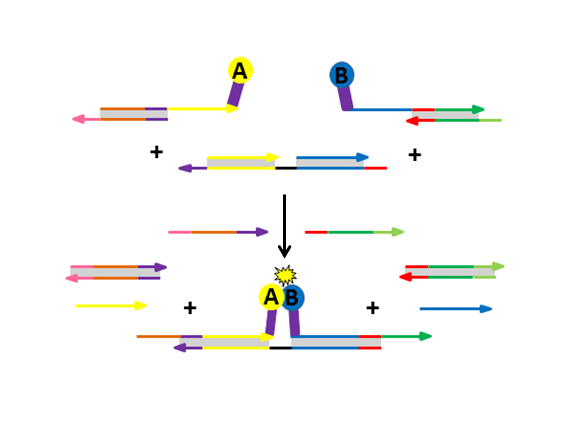
A funny anecdote comes from when Daniel first began exploring the use of protein-DNA conjugates for strand displacement. He was finding success dynamically assembling DNA-labeled CFP and YFP for FRET assembly to some extent, but he often observed a significant level of background leak. Struggling to eliminate the leak, Daniel’s fortunes changed when a back injury put him on bedrest for a week. Upon his return, he tested samples he had assembled prior to his injury and found that there was absolutely no leak! Turns out, DNA hybridization occurred slowly in the buffer he used so many complexes were inefficiently blocked. Upon changing to an optimized DNA hybridization buffer, this issue was completely resolved. From there, he was quickly able to expand his designs to incorporate more complex architectures quite smoothly. Now, when someone in lab is struggling, we joke that they should throw out their back to change their luck.
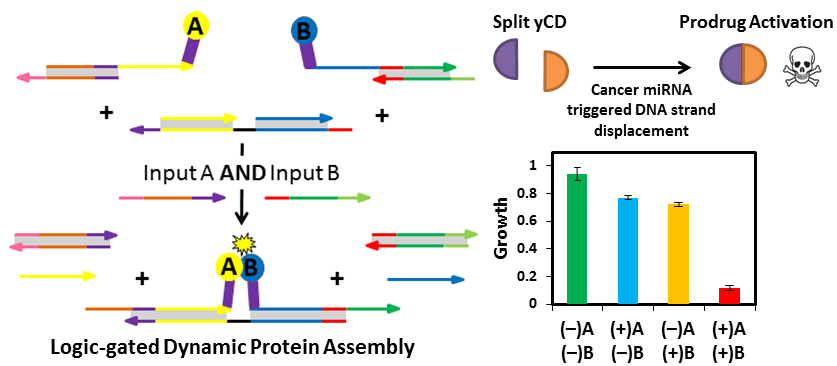
With the framework Daniel established, we explored other output proteins. Qing Sun applied the system to successfully control an enzyme cascade reaction. I joined the project while working on reconstituting the activity of split yeast cytosine deaminase (yCD), a prodrug activating enzyme, in the presence of cancer-specific RNA. Struggling with RNA binding proteins, I realized adopting a strand displacement strategy would be perfect. Given the simple design rules of our circuits, I was able to successfully reprogram the circuit to target cancer miRNA very smoothly. Upon labeling the circuit with split yCD, I found that prodrug activation directly corresponded to the circuit being turned on by the target miRNA. This exciting result proved how modular and tunable our system is towards any input and output of interest. The implication being that this technology has a powerful application in disease therapeutics and personalized medicine.
In hindsight, the idea seems simple: link proteins with DNA and take advantage of both. However, at the time, using strand displacement to assemble proteins was an out-of-the-box idea that had never been explored before. This idea has spawned a wealth of new projects within our lab, and we continue to apply this technology and, more generally, this line of thinking to other synthetic biology problems. Often times the best ideas come from simple views of problems from a creative angle.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in