Recipe for success - interaction proteomics becomes a household item
Published in Protocols & Methods
The affinity purification coupled mass spectrometry (AP-MS) and proximity-labeling methods such as BioID have been wildly applied to detect protein interaction networks. Both methods are complementary and provide a comprehensive view of a protein's interactome. We are developing an integrated approach named MAC (Multiple Approaches Combined) –tag system, which utilizes both AP-MS and BioID in a single construct to parallel explore the protein interactome. Moreover, application of MAC-tag system on subcellular localization markers provided a proximity cellular protein landscape (Figure 1), which serves as a reference grid to probe the molecular level distribution of the protein of interest. Therefore, the original publication (https://www.nature.com/articles/s41467-018-03523-2) that released in March 2018 has drawn massive attention in the community. Especially the feature of assigning the protein localization based on interactome data has proven the MAC-tag system a really powerful and sensitive tool to investigate molecular mechanisms of disease-causing mutation proteins.
As a continuation, in current work, we gathered all the feedback and suggestions from our collaborators and users, to update online platform of MAC-tag system. The updated version of MAC-tag system (www.proteomics.fi) consists of four main modules: data input, SAINT analysis, data filtering, and MS-microscopy visualization, greatly facilitating the data processing. The spatial information of reference database covers most of the cellular organelles including 19 cellular compartments, and 3 sub-organelle regions of mitochondria.
The development and application of MAC-tag system has never been stopped. Recently, we have adapted the MAC-tag technology to expand the knowledge and understanding of COVID-19 and the collection of related MAC-tag plasmids available via Addgene.org (link)
We welcome any comments or suggestions for our MAC-tag platform, in order to help us to improve our services to the community. Meanwhile, we will continue our efforts with MAC-tag system development to explore protein-protein interaction networks and their implications for cell biology.
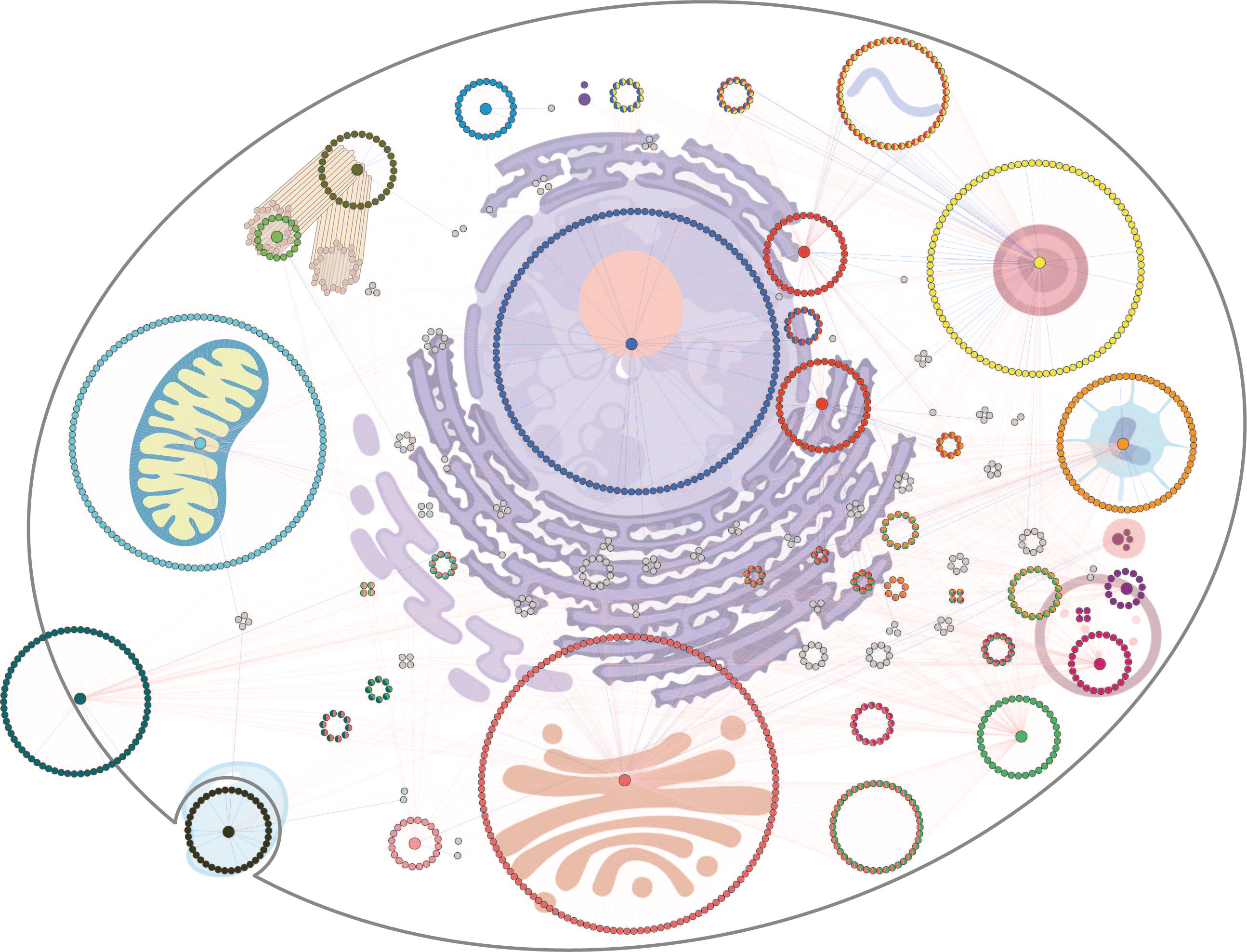
Figure 1. Subcellular localization markers provided a proximity cellular protein landscape.
The study was financially supported by the Finnish Academy, University of Helsinki, The Sigrid Juselius Foundation, Instrumentarium Science Foundation, Biocentrum Helsinki and HiLIFE.
Reference:
Xiaonan Liu, Kari Salokas , Rigbe Weldatsadik , Lisa Gawriyski and Markku Varjosalo
Nature Protocols
Follow the Topic
-
Nature Protocols
This journal publishes secondary research articles and covers new techniques and technologies, as well as established methods, used in all fields of the biological, chemical and clinical sciences.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in