Recombinant DNA: how simple can we make it?
Published in Bioengineering & Biotechnology

Science can be complicated, which is perhaps why scientists have always been fascinated with simplicity. Famous examples include: “Simplicity is the ultimate sophistication” (Leonardo de Vinci), “Truth is ever to be found in the simplicity, and not in the multiplicity and confusion of things” (Isaac Newton) and “Everything should be made as simple as possible, but not simpler” (Albert Einstein).
Everyone who has tried working in a laboratory knows the pain of a failed or irreproducible experiment. Some even say that a field like synthetic biology is facing a reproducibility crisis. One thing is for sure: a major bottleneck in biotechnology is the complexity of both biology and molecular biology protocols.
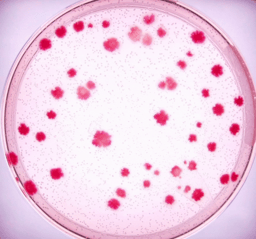
Two complementary ways to address these challenges is to introduce standards in molecular biology and to simplify the methods. With this mind, we created SEGA: the standardized genome architecture. With SEGA, recombining DNA doesn’t get any simpler: We just add DNA to cells! The DNA automatically integrates on the genome and successful recombinant clones are identified based on the color of the bacterial colonies.
Sounds cool, right? But surely this is not compatible with making advanced genetic constructs directly on the genome of E. coli? Wrong! This is where standards (and years of hard work in our lab) come into play. We generated more than hundred different “landing pads” on the genome hosting a variety of well-characterized genetic modules controlling transcription, translation, and down-streaming processing of your gene- or protein-of-interest. Everything is open source. The strains are available at the Belgian Co-ordinated Collections of Microorganisms (https://bccm.belspo.be), all sequences can be downloaded in Genbank format and SBOL format at https://synbiohub.org, and detailed protocols are available at protocols.io64.
Watch Carolyn explain more about SEGA here : https://sega-genomes.com & https://www.youtube.com/watch?v=ZnsXpZflOF4
As a scientific principle, Ockham’s words are perhaps better known with slight modifications as Occam’s razor: “If two explanations are plausible, the simplest explanation, usually the one with the fewest assumptions, is preferred”. I like the other version better – it is simpler! And I think that the only necessary things for making recombinant DNA should be: DNA and cells.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in