During virus infection a drastic remodeling of the host cell takes place. Existing cellular machinery is repurposed to design an optimal viral factory, akin to reshuffling a set of Lego blocks. This reorganization has been studied at different levels: from large scale structural rearrangements such as replication organelle formation, to changes in protein expression caused by alterations in transcription or cap-dependent translation. However, it is often overlooked that in a cell the most complex behavioral changes start with the subtlest of changes: post-translational modifications. And if a system exists, you can bet viruses learned to make use of it.
To study the full extent to which viruses make use of post-translational regulatory networks, we performed a system-wide, unbiased proteomics and phospho-proteomics screen on cells infected with the picornavirus CVB3. CVB3 belongs to a group of enteroviruses that include many human pathogens as well as newly emerging global health threats such as EV-A71 and EV-D68. At multiple time points after CVB3 infection, we traced the up- or downregulation of protein levels as well as changes in their phosphorylation status by mass spectrometry. An impressive change in protein levels was observed, affecting over 25% of the detected proteins. Yet this change was exceeded by an even more astounding impact of the infection on the phosphoproteome, with over 85% of identified phosphosites significantly altered.


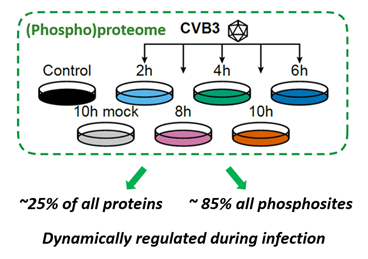
Figure 1. Experimental set up to map the dynamic regulation of the proteome and phosphoproteome during infection.
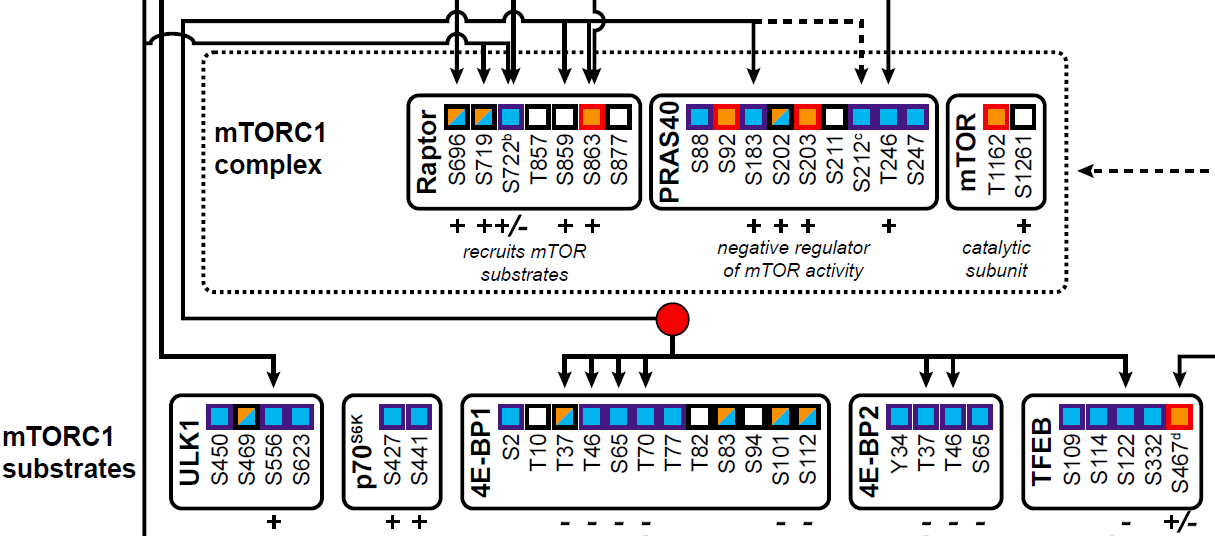
This extensive alteration of phosphosites suggests a functional role in infection. Among the detected sites, we saw an enrichment for sites regulating signaling cascades relating to apoptosis/cell death, transcription/translation, and the PI3K/AKT/mTOR and MAPK signaling pathway. In this paper, we zoomed in on the regulation of the mTORC1 network to demonstrate how these data enable us to better understand the dynamics of the regulatory rewiring that occurs in a cell upon infection. We were able to create a detailed and comprehensive oversight of a complex network leading to a decrease in mTORC1 activity and activation of downstream targets. Among these, our analysis revealed the activation of the downstream mTORC1 target TFEB, a regulator of lysosome and autophagy homeostasis. Through dedicated experiments, we discovered that TFEB enhances the release of virus particles within extracellular vesicles. This finding seems to stem from the ability of TFEB to enhance the induction of secretory autophagy and brings us one step closer to understanding the regulation of non-lytic virus release.
Interestingly, many of the identified phosphorylation sites altered by CVB3 infection do not yet have annotated functions. This knowledge gap complicates data interpretation. However, at the same time it emphasizes the wealth of knowledge that can still be gleamed with these system-wide, unbiased screening approaches. We therefore encourage data mining of our findings and look forward to future efforts that will further elucidate the functional role of the detected phosphosites during infection as complement to the regulatory pathways we have highlighted here.
Piero Giansanti
Jeroen Strating
Kyra Defourny
Frank van Kuppeveld
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026







Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in