Shedding light on the elusive neurons of comb jellies
Published in Ecology & Evolution
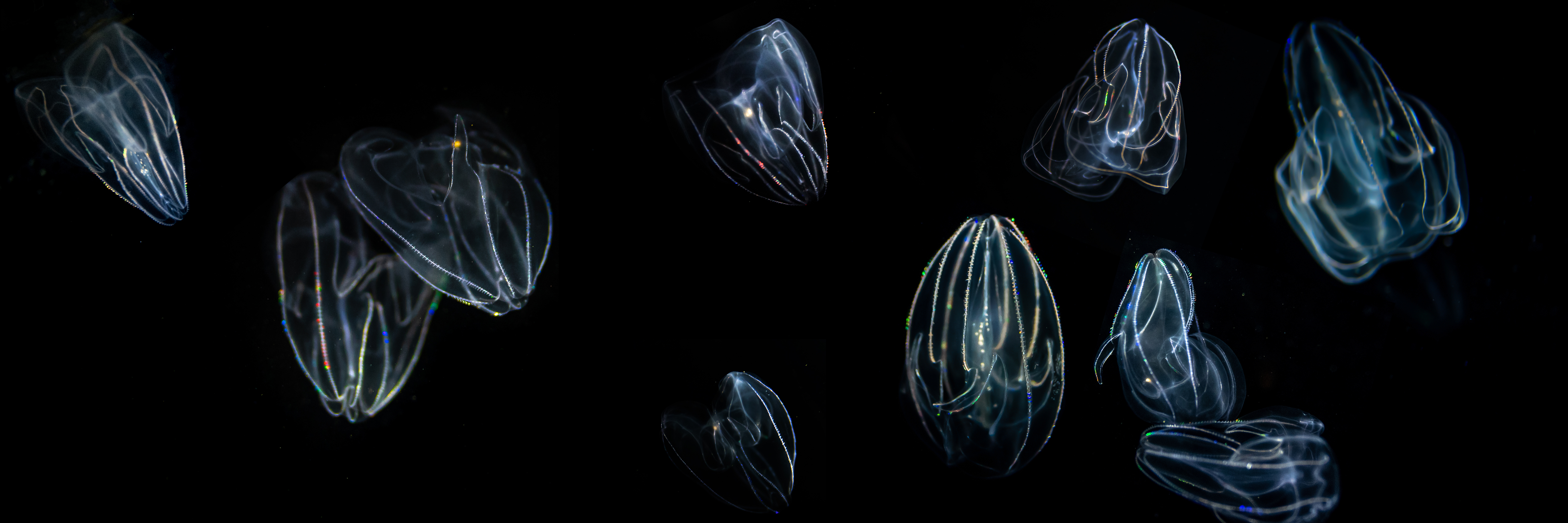

Ctenophores (pronounced “teen-o-fours”), also known as comb jellies, are semi-transparent gelatinous marine invertebrates belonging to the phylum Ctenophora. The phylum derives its name from one of the distinctive features of its members - a body with eight rows of comb plates made up of fused cilia (Gr. ctene, comb, + phora, bearer). These cilia move rhythmically to propel the animal through the water. Ctenophores are distributed in all oceans and at all depths, though the best described species are found in shallow waters near shores. Other common names of ctenophores are sea walnuts, sea gooseberries, and Venus' girdles.
Ctenophores are fascinating creatures yet there’s a lot we don’t know about them. They are the oldest animals with a nervous system, making them even more interesting from an evolutionary point of view. For a few years now, we have been studying the intriguing nervous system of ctenophores in our laboratory, the Evolutionary Neurobiology Unit in the Okinawa Institute of Science and Technology (OIST), Japan. In collaboration with Professor Kazuo Inaba of Shimoda Marine Research Center (University of Tsukuba), we have successfully established lab cultures of two ctenophore species: the predominant ctenophore in Japanese coastal waters, Bolinopsis mikado, and the comb-less benthic ctenophore, Vallicula multiformis.
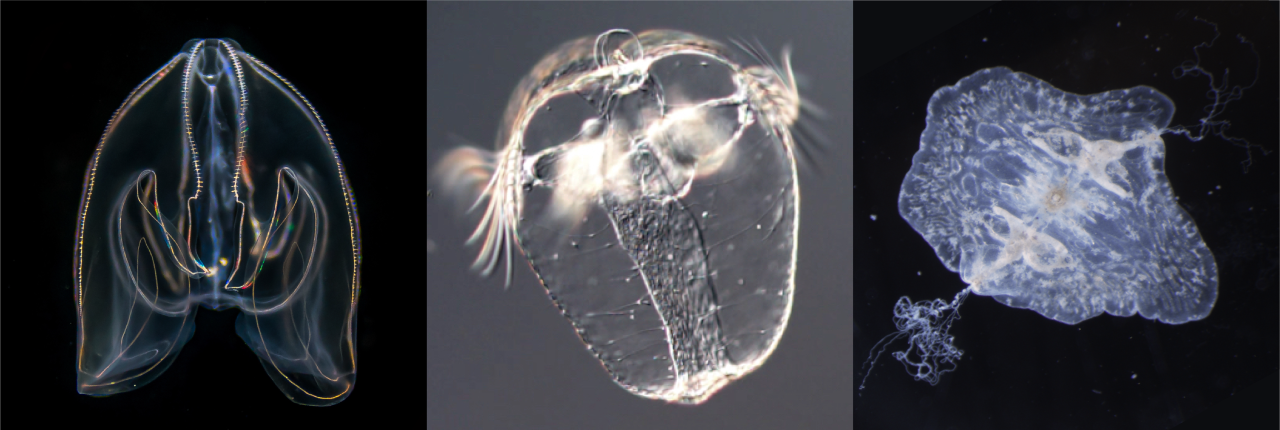
Ctenopore species housed in the Evolutionary Neurobiology Unit, OIST (Japan) — Bolinopsis mikado (adult-left, larva-middle) and Vallicula multiformis (right).
Problem: ctenophore’s nervous system is still enigmatic
It is ironic how much is known about the complex human brain, while very little is known about the simple nervous system of ctenophores. The difficulty in examining ctenophore neurons arises from several problems. First, many “neural” genes predate the appearance of the nervous system, and their expression and function in non-neural cells in early-branching animals like ctenophores complicates the molecular characterization of ancestral neurons. Second, the neurotransmitters used by ctenophores are not yet well defined, hence it is not possible to label and examine the neurons as easily as in other animals. Third, efforts to identify neuropeptides (a type of neurotransmitter) in ctenophores are largely based on our knowledge about sequence and processing features of known neuropeptide genes isolated from distantly related animal species1,2. However, such an approach predicts only the peptide precursor gene candidates but does not provide information about authentic mature peptide structures, thereby decreasing the reliability of peptide identification.
Our solutions
In contrast to previous analyses, our recent study used mass spectrometry (MS)-based method to comprehensively identify and validate ctenophore (neuro)peptides. We then demonstrated that the peptide markers can be used to visualize the ctenophore nervous system architectures. We further characterized the peptide-expressing cells by analyzing the single-cell expression data3, performing functional analysis, and predicting peptide-receptor pairs using machine learning.
Ctenophore’s nervous system shares common features with all other animals
As we expected, the genes encoding for ctenophore peptides have low sequence homology with any of the known neuropeptides. However, we found that the precursor peptides undergo proteolytic processing at acidic residues sites, similar to the neuropeptides of cnidarians (another group of evolutionarily ancient animals). We took advantage of the sequence information obtained from our MS analysis and performed a series of immunostainings. We found that the validated ctenophore (neuro)peptides may participate in volume transmission as in bilaterian animals (human, mouse, fish, fly). The cells containing the peptides also express most of the genes that are responsible for maturation, secretion, and degradation of neuropeptides in cnidarians and bilaterians. Furthermore, our functional analysis using the Bolinopsis larva has revealed that the VWYamide and NPWamide neuropeptides can trigger muscle contraction, thereby eliciting a conserved function of neuropeptides.
For these multifaceted findings that show an unexpected level of similarity between ctenophore and cnidarian/bilaterian nervous systems, we believe that there is a common evolutionary origin of the animal peptidergic nervous system.
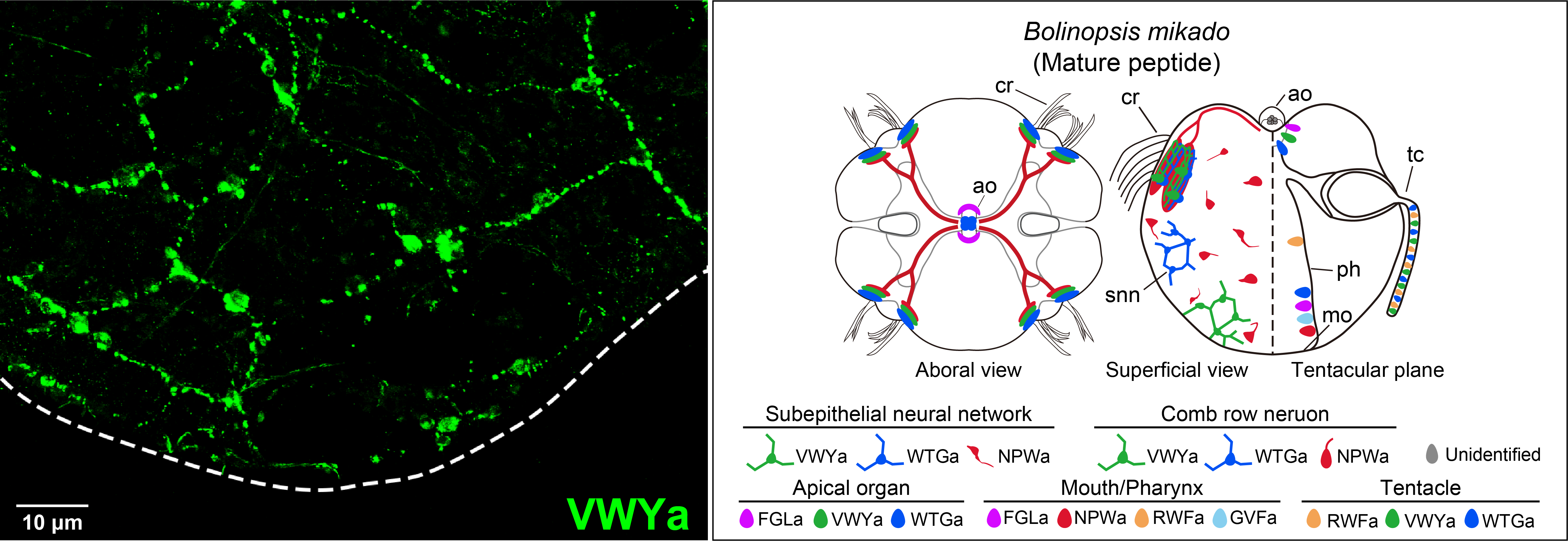
Nerve net of the ctenophore Bolinopsis mikado labelled with anti-VWYamide antibody (left). Schematic diagram of the spatial distribution of peptide-expressing neurons/cells in Bolinopsis larva (right).
With our work, we shed a light on the “hidden” neurons of ctenophores. Now, the neurons can be easily identified, enabling us to examine their physiological properties. This is a fundamental step in the study of the ancestral nervous system and how it increased in complexity throughout animal evolution. We are excited to see how the many enigmas of ctenophores will be resolved.
Images: Banner photo and adult Bolinopsis (Soumen Jana), Bolinopsis larva (Osamu Horiguchi), Vallicula (Dr. Kurato Mohri).
References:
- Sachkova, M. Y. et al. Neuropeptide repertoire and 3D anatomy of the ctenophore nervous system. Curr Biol, doi:10.1016/j.cub.2021.09.005 (2021).
- Jager, M. et al. New insights on ctenophore neural anatomy: immunofluorescence study in Pleurobrachia pileus (Muller, 1776). J Exp Zool B Mol Dev Evol 316B, 171-187, doi:10.1002/jez.b.21386 (2011).
-
Sebe-Pedros, A. et al. Early metazoan cell type diversity and the evolution of multicellular gene regulation. Nat Ecol Evol 2, 1176-1188, doi:10.1038/s41559-018-0575-6 (2018).
Follow the Topic
-
Nature Ecology & Evolution
This journal is interested in the full spectrum of ecological and evolutionary biology, encompassing approaches at the molecular, organismal, population, community and ecosystem levels, as well as relevant parts of the social sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Biodiversity and ecosystem functioning of global peatlands
Publishing Model: Hybrid
Deadline: Jul 27, 2026
Understanding species redistributions under global climate change
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in