Single cell transcriptome analysis of eye in multi-species.
Published in Healthcare & Nursing

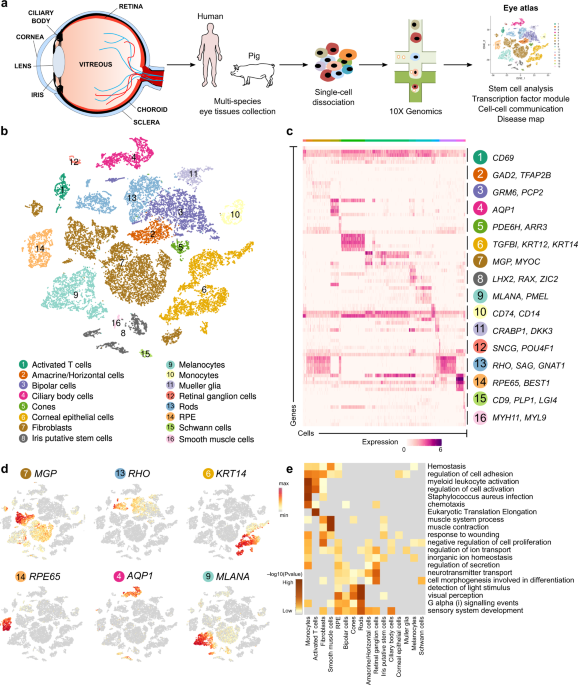
The vertebrate eye has evolved by undergoing significant changes. However, much of the functioning of the eye and its compartments remain conserved across species. There have not been many cross-species studies of the eye. With the advances in single-cell RNA sequencing, retinal single-cell atlases have been generated in mice1, zebrafish2, non-human primates3, chicks4, and humans5,6. With such datasets, it provides an opportunity to understand cross-species differences across species using single-cell RNAseq technologies.
The human eye is a complex organ comprising of cell types from different developmental origins. In our current work, we tried to understand the diverse cell types of the human eye using single-cell RNA sequencing. In the process, we identify and catalogue more than fifty thousand cells across tissues like the retina, iris, cornea, choroid, sclera, and retinal pigmented epithelium. We performed a comparison of our retina single-cell RNAseq datasets with earlier published datasets5,6 and observed high transcriptomic similarity across cell types. In human eye, discrete transcriptional factor modules, which are specific to cell type, were found. Subsequently, we checked the conservation of such modules across species. Published retinal single-cell RNAseq is available in mice, zebrafish, non-human primates, and pigs. Therefore, we explored an opportunity to compare retina across species. In this way, we focussed on the core transcription factors (TFs) that identify a cell type across species in the retina.
In the non-retinal cell populations, we identified putative stem cells present in the iris tissue of both pigs and humans. Even though such cells express markers of retinal progenitor cells or retinal stem cells, these cells have to be verified for their capability in generating retinal neurons in vivo before revisiting the annotation as retinal stem cells. Therefore, we have annotated them as putative stem cells. Such cells express a promising panel of markers (WNT2B, MDK, FGFR1), similar to retinal progenitor cells. We also observed TFs that populate such cell types (MSX1, MSX2) and the active signalling pathways (FGF signalling, MDK signalling). Even though the work lack functional in vivo studies, this is the first time such cells with unique transcriptional factor makeup, cell surface receptors, and gene markers are being reported in the adult human eye. Such cell types could be of immense interest to the scientific community.
Finally, we wanted to check the perturbation effects of one core TF KLF7, conserved across species in retinal ganglion cell cells. After perturbing KLF7 gene expression during retinal ganglion cells (RGC) differentiation from human embryonic stem cells, current studies concluded that KLF7 plays a significant role in the maturation of RGC cells during differentiation.
In summary, our work catalogues the different cell types in the anterior and posterior compartments of the eye. In addition, we found rare cell populations in iris tissue and labelled them as putative stem cells because of their unique markers and TF makeup. We hope our work will provide resources to scientists working with retinal biology, eye disorders, single-cell genomics, and adult stem cells.
1 Macosko, E. Z. et al. Highly Parallel Genome-wide Expression Profiling of Individual Cells Using Nanoliter Droplets. Cell 161, 1202-1214, doi:10.1016/j.cell.2015.05.002 (2015).
2 Hoang, T. et al. Gene regulatory networks controlling vertebrate retinal regeneration. Science 370, doi:10.1126/science.abb8598 (2020).
3 Peng, Y. R. et al. Molecular Classification and Comparative Taxonomics of Foveal and Peripheral Cells in Primate Retina. Cell 176, 1222-1237.e1222, doi:10.1016/j.cell.2019.01.004 (2019).
4 Yamagata, M., Yan, W. & Sanes, J. R. A cell atlas of the chick retina based on single-cell transcriptomics. Elife 10, e63907 (2021).
5 Lukowski, S. W. et al. A single-cell transcriptome atlas of the adult human retina. Embo j 38, e100811, doi:10.15252/embj.2018100811 (2019).
6 Menon, M. et al. Single-cell transcriptomic atlas of the human retina identifies cell types associated with age-related macular degeneration. Nat Commun 10, 4902, doi:10.1038/s41467-019-12780-8 (2019).
Follow the Topic
-
Nature Communications

An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in