Skin basal cell carcinomas assemble a pro-tumorigenic spatially organized and self-propagating Trem2+ myeloid niche
Published in Cancer
Background:
Basal cell carcinomas (BCCs) are hedgehog-signaling-driven cancers, with targeted hedgehog inhibitors (HHi) being primarily used to treat patients with advanced non-surgically resectable cases1. In a large fraction of patients, resistance can develop, which has led to the approved utilization of immune checkpoint blockade (ICB) strategies to treat these HHi resistance cases2. Although ICB has shown promise, a large fraction of patients do not respond, reinforcing the need to better understand both tumor intrinsic and extrinsic mechanisms of growth and resistance2.
Our previous work has identified that resistant tumor epithelial populations that do not respond to targeted HHi are preexisting within the naive non-drug treated tumor and occupy distinct spatial localization within the tumor3. This population, which is marked by the surface marker LY6D, represents a plastic cellular state reminiscent of squamous cell carcinoma (SCC), with divergent molecular pathways where growth is not dependent on canonical HH signaling3.
While key BCC epithelial states are defined, it is not clear what elements of the tumor microenvironment (TME) drive these distinct states. As such, we set out to define the spatial organization of the TME in relation to these epithelial states, identifying TREM2+ macrophages as a key spatially organized population that supports the growth of a distinct BCC-associated (non-SCC) epithelial state.
Key Results:
BCCs are spatially organized into neighborhoods with distinct cell types including TREM2+ macrophages occupying spatially distinct regions.

Using a multiplexed imaging approach, we spatially profiled human BCC tumors and identified that there are distinct neighborhood regions which we called, the immune swarm (IS), upper tumor (UT), and lower tumor (LT). A key feature we noted was that there was enhanced proliferation within the LT region leading to the hypothesis that there might be a distinct stromal cell composition within the LT that drives this enhanced proliferation. After extracting the stromal cells from all three regions, we were able to directly compare the cellular compositions. A key difference we noted was the enrichment of HLA-DR+ cells within the LT region. In addition to HLA-DR+ cells, we found that there were additionally large numbers of FOXP3+ T regulatory cells within the LT region.
Using a combination of multi-omic data, we were able to find that a large proportion of the HLA-DR+ cells are macrophages, and a subset of these macrophages are TREM2+, which have been recently implicated in fostering an immunosuppressive TME4-6. We find that the TREM2+ cells represent a more terminal state in the predicted lineage trajectory. Quite interestingly, we show that this lineage trajectory is spatially organized spanning from the UT to the LT region.
Trem2+Vcam1+ skin cancer-associated myeloid cells (SCAMs) are distinct from other non-cancer-associated Trem2+ myeloid cells.
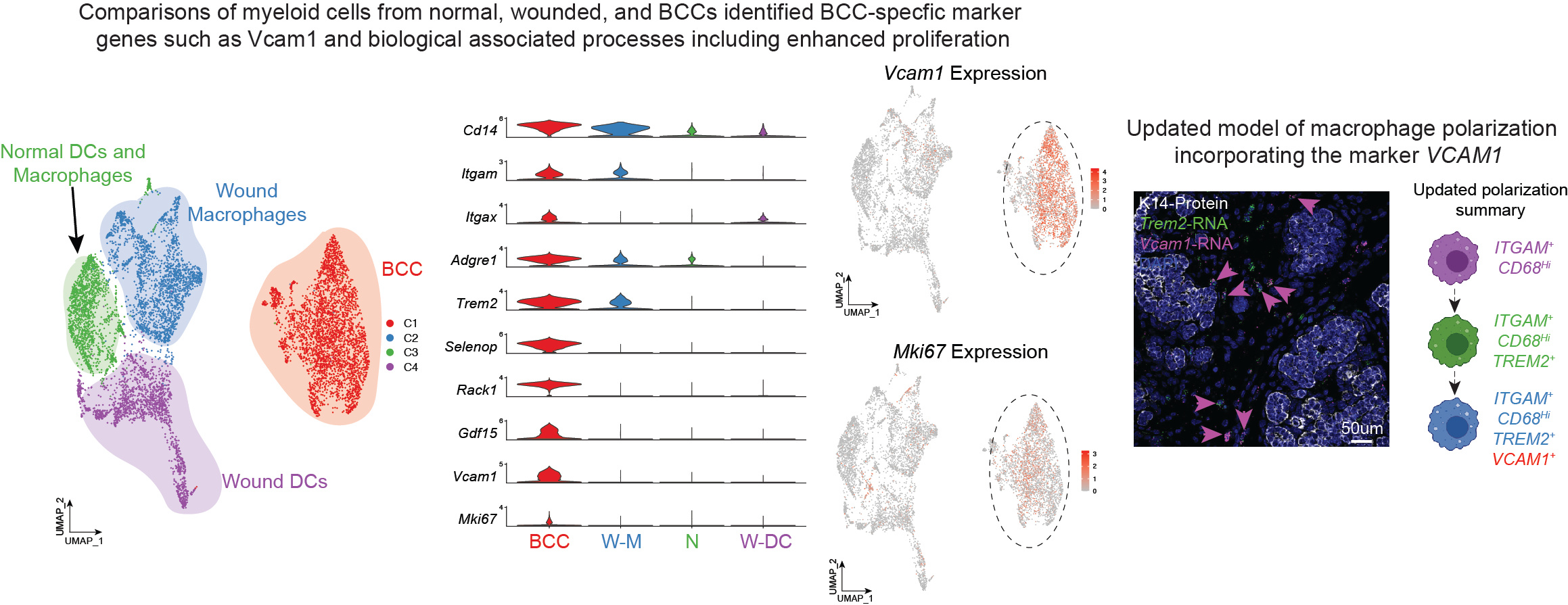 TREM2+ cells have been identified in multiple contexts within the skin during states of normal tissue regeneration, tissue repair, and now in the context of BCCs7,8. Comparative analysis between myeloid cells from these different distinct states identified key features associated with Trem2+ cells from BCCs such as the expression of the surface marker Vcam1 and the observation that these cells seem to have enhanced proliferation. Based on these features, we decided to call these BCC-specific Trem2+Vcam1+ cells skin cancer-associated myeloid cells (SCAMs).
TREM2+ cells have been identified in multiple contexts within the skin during states of normal tissue regeneration, tissue repair, and now in the context of BCCs7,8. Comparative analysis between myeloid cells from these different distinct states identified key features associated with Trem2+ cells from BCCs such as the expression of the surface marker Vcam1 and the observation that these cells seem to have enhanced proliferation. Based on these features, we decided to call these BCC-specific Trem2+Vcam1+ cells skin cancer-associated myeloid cells (SCAMs).
Trem2+ macrophages can promote tumor epithelial growth via an immunosuppression-independent mechanism via the secretion of Osm.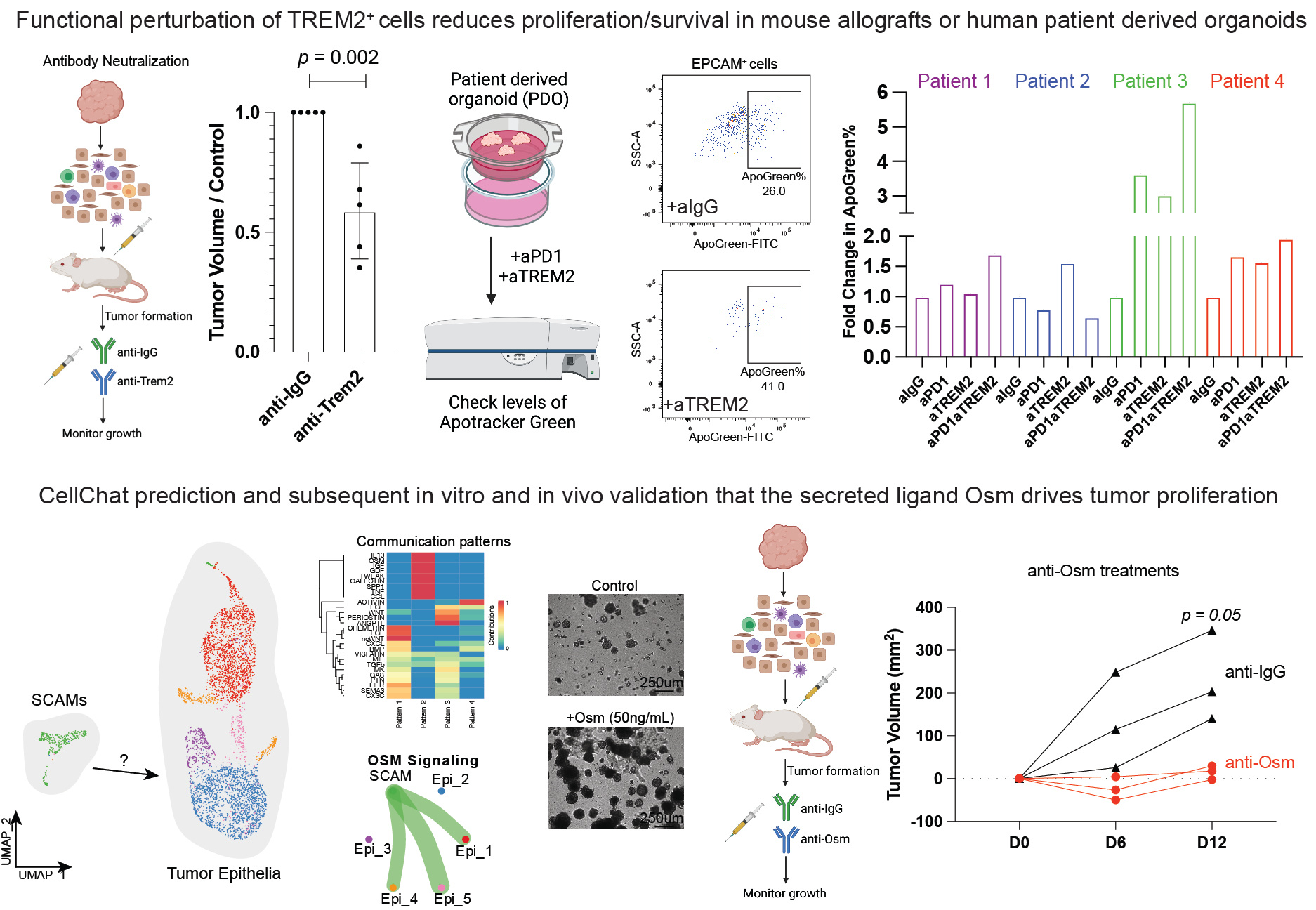
Functional experiments in our mouse allograft system showed that targeting SCAMs reduced tumor growth. To validate and confirm findings, we utilized our human patient-derived organoid system, where we could acutely treat pieces of a human tumor with aTREM2 and/or aPD1 antibodies and then assess the effect on the tumor epithelium. In some samples, aTREM2 or aPD1 treatment alone led to enhanced cell death as read out by ApoGreen. Interestingly, dual treatment of aTREM2 with aPD1 did seem to further enhance the ApoGreen levels in the tumor epithelium.
To determine how SCAMs cells might be promoting tumor growth, we utilized CellChat, a powerful receptor/ligand inference tool9. We computationally identified the Osm/Osmr signaling pathway (where Osm is secreted by SCAMs and Osmr is expressed on the tumor epithelium) and subsequently validated our findings in both in vitro organoid and in vivo mouse models.
The TME creates and drives a Trem2+ self-propagating niche.
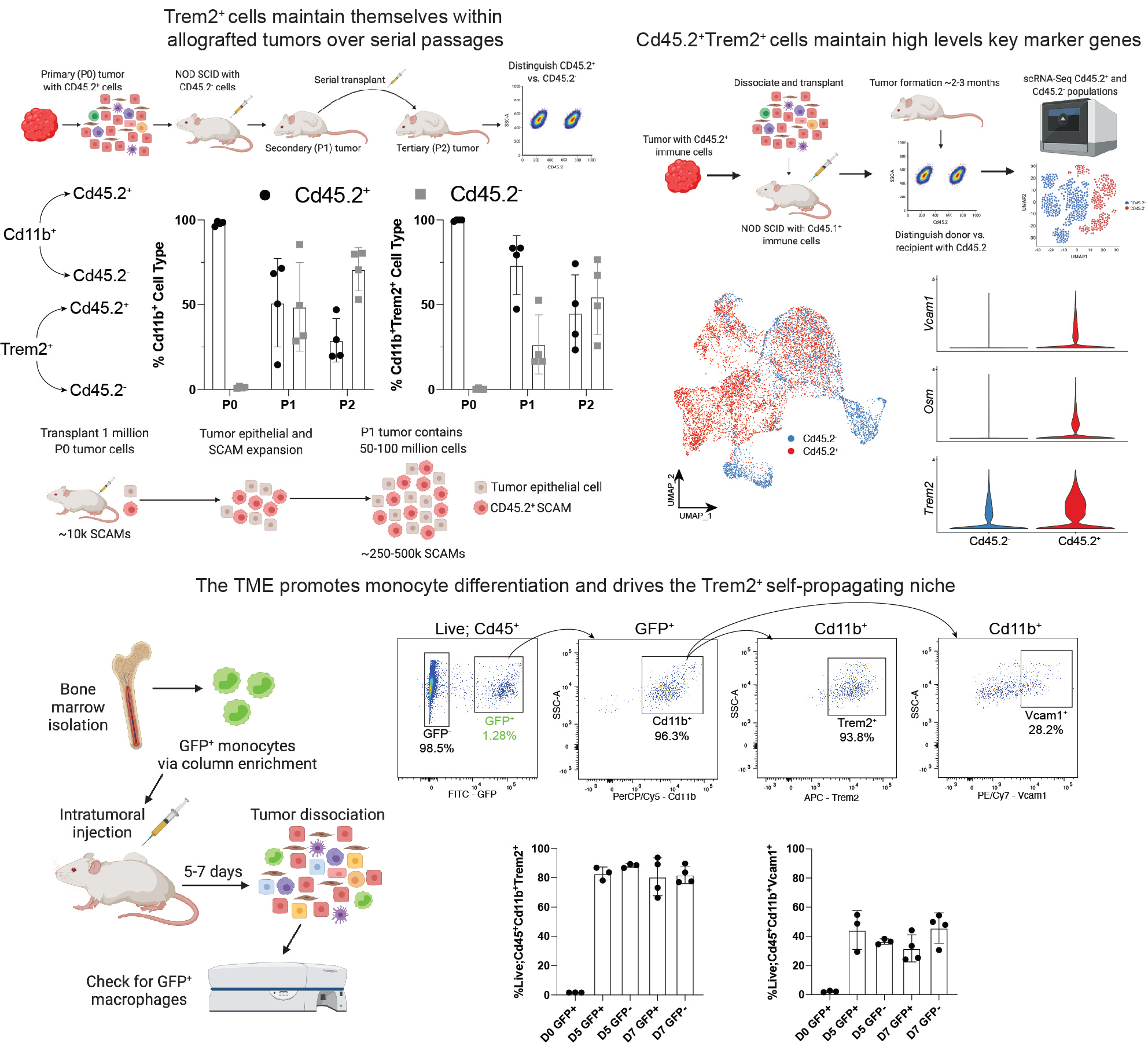 A key observation from the comparative analysis between SCAMs and other myeloid cells found in the skin during normal tissue regeneration or during repair was the noticeably enriched proliferation. We found that SCAMs actively proliferate within the allograft tumors and through tracing via the Cd45.1/Cd45.2 system (where SCAMs are Cd45.2+ and recruited immune cells from the allograft hose would be Cd45.1+), we found that SCAMs from the initial primary tumor maintain themselves long-term over serial passages. Isolating SCAMs from the allograft tumors and subsequent analysis found that these cells maintain key features of SCAMs such as high expression of Trem2, Osm, and the key surface marker Vcam1.
A key observation from the comparative analysis between SCAMs and other myeloid cells found in the skin during normal tissue regeneration or during repair was the noticeably enriched proliferation. We found that SCAMs actively proliferate within the allograft tumors and through tracing via the Cd45.1/Cd45.2 system (where SCAMs are Cd45.2+ and recruited immune cells from the allograft hose would be Cd45.1+), we found that SCAMs from the initial primary tumor maintain themselves long-term over serial passages. Isolating SCAMs from the allograft tumors and subsequent analysis found that these cells maintain key features of SCAMs such as high expression of Trem2, Osm, and the key surface marker Vcam1.
Finally, to determine if it is indeed the TME that drives cells to a SCAM fate, we took freshly isolated bone marrow-derived monocytes and injected them intratumorally. We found that within 5 days after injection, these cells expressed both Trem2 and Vcam1 at high levels suggesting that they were able to take on a SCAM-like phenotype.
Take home message:
Overall, this work identifies the role of TREM2+ cells in BCCs, highlighting their development and spatial location within the tumor. Importantly, this work identifies an immunosuppression-independent role of TREM2+ cells within the TME.
References:
1 Atwood, S. X. et al. Smoothened variants explain the majority of drug resistance in basal cell carcinoma. Cancer Cell 27, 342-353, doi:10.1016/j.ccell.2015.02.002 (2015).
2 Stratigos, A. J. et al. Cemiplimab in locally advanced basal cell carcinoma after hedgehog inhibitor therapy: an open-label, multi-centre, single-arm, phase 2 trial. Lancet Oncol 22, 848-857, doi:10.1016/S1470-2045(21)00126-1 (2021).
3 Haensel, D. et al. LY6D marks pre-existing resistant basosquamous tumor subpopulations. Nature Communications 13, 7520, doi:10.1038/s41467-022-35020-y (2022).
4 Katzenelenbogen, Y. et al. Coupled scRNA-Seq and Intracellular Protein Activity Reveal an Immunosuppressive Role of TREM2 in Cancer. Cell 182, 872-885 e819, doi:10.1016/j.cell.2020.06.032 (2020).
5 Molgora, M. et al. TREM2 Modulation Remodels the Tumor Myeloid Landscape Enhancing Anti-PD-1 Immunotherapy. Cell 182, 886-900 e817, doi:10.1016/j.cell.2020.07.013 (2020).
6 Binnewies, M. et al. Targeting TREM2 on tumor-associated macrophages enhances immunotherapy. Cell Rep 37, 109844, doi:10.1016/j.celrep.2021.109844 (2021).
7 Wang, E. C. E., Dai, Z., Ferrante, A. W., Drake, C. G. & Christiano, A. M. A Subset of TREM2(+) Dermal Macrophages Secretes Oncostatin M to Maintain Hair Follicle Stem Cell Quiescence and Inhibit Hair Growth. Cell Stem Cell 24, 654-669 e656, doi:10.1016/j.stem.2019.01.011 (2019).
8 Haensel, D. et al. Defining Epidermal Basal Cell States during Skin Homeostasis and Wound Healing Using Single-Cell Transcriptomics. Cell Rep 30, 3932-3947 e3936, doi:10.1016/j.celrep.2020.02.091 (2020).
9 Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat Commun 12, 1088, doi:10.1038/s41467-021-21246-9 (2021).
Follow the Topic
-
Nature Communications

An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Mar 24, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in