Small deer with giant fusion chromosomes
Published in Ecology & Evolution

Muntjacs aren’t the familiar deer species that occasionally hang out in one’s backyard, so I was quite surprised when seeing it appeared on a Berkeley local news one time when I was still a postdoc in UC Berkeley. The Chinese muntjacs (Muntiacus reevesi) are called ‘the little muntjac’ in Chinese (Figure 1A), while in English, they are also called ‘barking deer’ as they bark like a dog. The Chinese muntjacs become vulnerable in their native country, they are flourishing in the English countryside. These two characters (and they are quite tamed) make them attractive as an exotic pet for some people. However, some muntjac species, e.g., black muntjac (Figure 1B) and giant muntjac in fact are endangered species due to disruption of their habitat. The muntjacs have inspired much interest from evolutionary biologists ever since the discovery of their extreme karyotype diversity in the 1970s. Back to the era of cytogenetics, one important way of defining a species is relying on their observed differences in chromosome numbers and morphology or to a finer scale, banding patterns (so that homologous chromosomes and chromosomal rearrangements can be inferred) between species. It was already known that karyotypes of gibbons range from 2n=38 to 2n=52 within their about 6 million years of divergence[1]. But muntjacs push the extent of chromosome number variation, and how fast such variations evolved to another level: within the estimated 3 million years’ divergence time, the chromosome number ranges from only 2n=6 (female) or 7 (male) chromosomes of Indian muntjac[2], 2n=8 (female) or 9 (male) of black muntjac [3] to 2n=46 of Chinese muntjac[4] (Figure 1B), making the Indian and black muntjacs with the lowest chromosome number known so far in mammals.

Figure 1A Chinese muntjac (Muntiacus reevesi) and 1B Black muntjac (Muntiacus crinifirons) Photo credit: Baowei Zhang
Why can some mammals like muntjacs live with so few chromosomes? What caused such rapid and dramatic changes and what are the consequences for the genome when chromosome numbers are greatly reduced? What roles do such chromosome number changes play during the rapid speciation of muntjacs? The answers to these questions obviously are relevant to a broader spectrum of biological topics that are beyond the less-known deer. But in the late 1970s, when the Sanger sequencing had just been developed, there weren’t many means for determining species’ genetic content or comparing it between different species. The earliest clue about how these different muntjacs’ chromosomes are related to each other, came from a pioneer study on a F1 hybrid between Indian and Chinese muntjacs (the existence of such a living hybrid between two species that differ so much in their chromosome numbers is already surprising enough) found in a zoo in Shanghai. Liming Shi, who also established the first cell bank in China for wildlife that is still actively running today in Chinese Academy of Science in Kunming, made the discovery using cells of the hybrid that repeated tandem fusions (centromere end of one chromosome to be fused with telomere end of another) probably accounted for the drastic reduction of chromosome numbers from 2n=46 to 2n=6/7 [5], by aligning the chromosome banding pattern of one species to another. This finding was later described by Barabra McClintock in her Nobel lecture in 1983, as an example of ‘genome shock’ that initiated later ‘stunning’ changes of chromosome organization[5,6]. And the tandem fusion mechanism that shaped the muntjac chromosome evolution was later confirmed by genome-wide chromosome painting and higher resolution of BAC mapping[7], and findings of both telomeric and centromeric repeats at the chromosome fusion sites[8].
This early work initiated a tradition of studying muntjac deer in Shi’s institute in Yunnan province, where over half of all the plant and animal species across China can be found, including four muntjac species. Another important discovery made later by Shi and his students is the unusual way by which the black muntjac chromosomes pair with each other during male meiosis. Shi showed that males of Indian or black muntjacs have one extra chromosome than their females, because of a translocation between the X chromosome and an autosome, leaving the Y chromosome as the ‘extra’ chromosome in males [3]. And five chromosomes of male black muntjac pair with one another through parts of homologous regions (Figure 2). It was shown later by chromosome painting and BAC clone mapping in black muntjacs[7], that X chromosome was translocated to an autosome chr4 (X+4), and that of the other chr4 was translocated to the short arm of chr1(1p+4) followed by an inversion, creating a pair of chromosomes that do not have homologous recombination across two thirds of its length specifically in males. Such complicated rearrangements only occurred in black muntjac after its divergence from its relative species Gongshan muntjac in less than 1.5 million years. This made this pair of chromosome evolve exactly like a pair of canonical XY chromosome, specifically lacking recombination in males, but much younger (hence they are called ‘neo-sex chromosomes’).
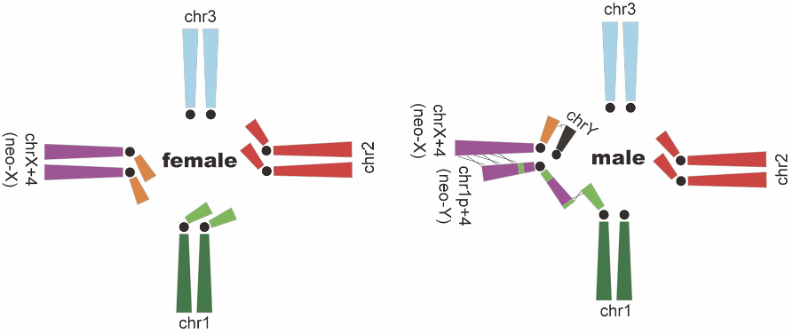
Figure 2 Karyotype of male and female black muntjacs There was a translocation between the eutherian X chromosome (orange) and the autosome 4 (purple), and another translocation between the short arm of autosome 1 (light green) and autosome 4, followed by an inversion. Therefore, seven chromosomes pair with one another through parts of homologous sequences.
Muntjacs thus provide a great paradigm to study the mechanisms of mammalian chromosome rearrangements, evolution of young sex chromosomes, and functional consequences of the former two on the entire genome. When I was a Ph.D. student, I had the chance to lead a project studying muntjac’s sex chromosomes, based on partial sequences of a couple of BAC clones in early 2000s. Now over 40 years later, the former students and students’ students of Professor Liming Shi worked together and finished the chromosome sequences of several muntjac species, and dissected the sequence composition at the chromosome fusion sites, and provided unequivocal evidence for the idea that tandem fusions have led to the chromosome numbers to change from 2n=46 to 2n=6. And they also found the three-dimensional genome architecture did not dramatically change after the fusion, this may explain why viable hybrid between Indian and Chinese muntjac species can still exist, after such a rapid speciation event.
Read the new paper of muntjac genomes here: https://www.nature.com/articles/s41467-021-27091-0
References
- Roberto R, Capozzi O, Wilson RK, Mardis ER, Lomiento M, Tuzun E, et al. Molecular refinement of gibbon genome rearrangements. Genome Res. 2007;17: 249–257.
- Wurster DH, Benirschke K. Indian muntjac, Muntiacus muntjak: a deer with a low diploid chromosome number. Science. 1970;168: 1364–1366.
- Ma K, Shi LM. [Comparative studies on synaptonemal complexes in spermatocytes of Chinese muntjac Muntiacus reevesi, black muntjac M. crinifrons and Indian muntjac M. muntjak]. Yi Chuan Xue Bao. 1988;15: 282–289.
- Wurster DH, Benirschke K. Chromosome studies in some deer, the springbok, and the pronghorn, with notes on placentation in deer. Cytologia . 1967;32: 273–285.
- Liming S, Yingying Y, Xingsheng D. Comparative cytogenetic studies on the red muntjac, Chinese muntjac, and their F1 hybrids. Cytogenet Cell Genet. 1980;26: 22–27.
- McClintock B. The Significance of Responses of the Genome to Challenge. 1984.
- Huang L, Chi J, Wang J, Nie W, Su W, Yang F. High-density comparative BAC mapping in the black muntjac (Muntiacus crinifrons): molecular cytogenetic dissection of the origin of MCR 1p+4 in the X1X2Y1Y2Y3 sex chromosome system. Genomics. 2006;87: 608–615.
- Tsipouri V, Schueler MG, Hu S, NISC Comparative Sequencing Program, Dutra A, Pak E, et al. Comparative sequence analyses reveal sites of ancestral chromosomal fusions in the Indian muntjac genome. Genome Biol. 2008;9: R155.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in