Targeting purine synthesis to curtail Staphylococcus aureus virulence
Published in Microbiology
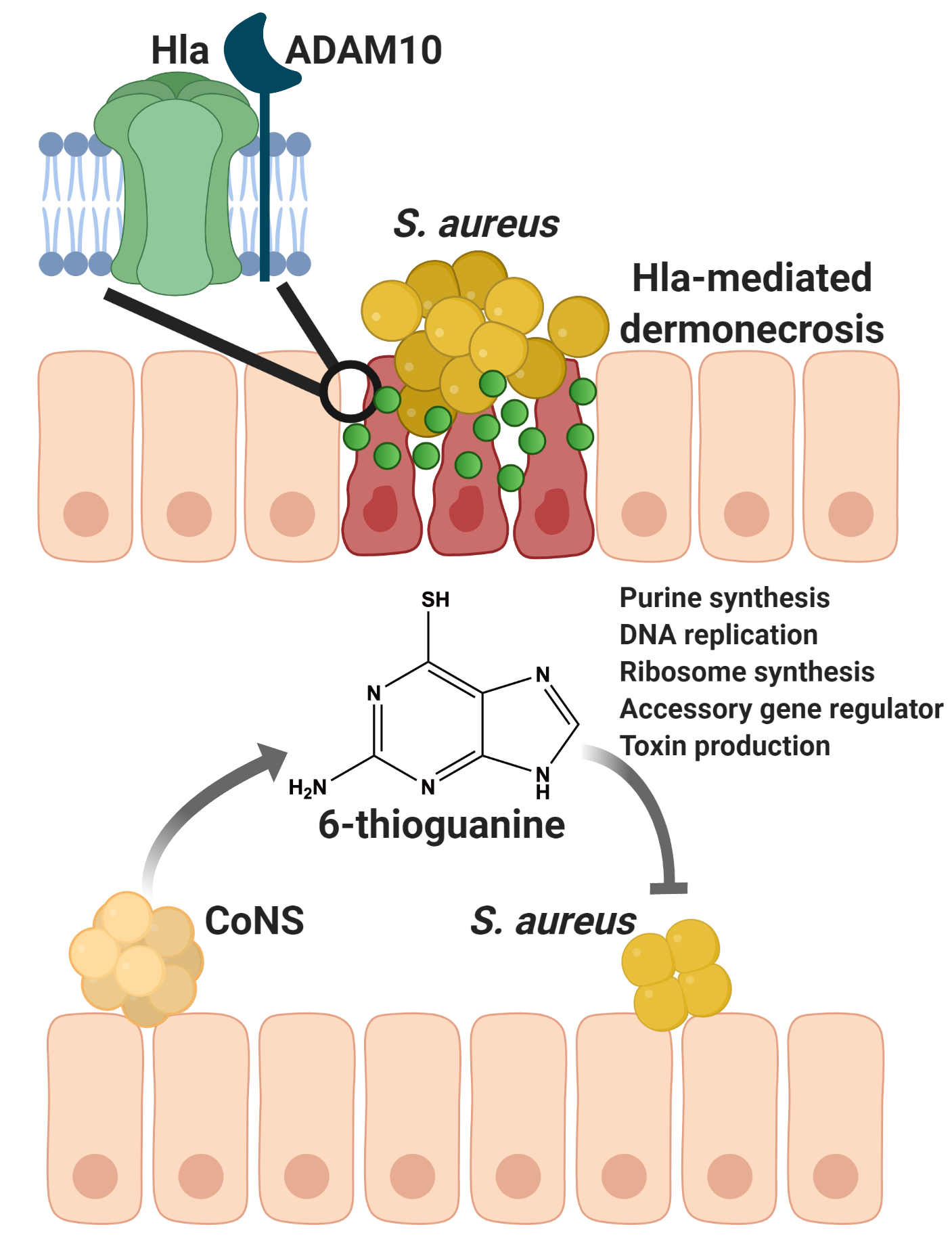
There is an increasing appreciation for the biological complexity that occurs within microbial communities. In these communities, microbes are constantly in competition to meet nutritional requirements in every environment, including the human host. Staphylococcus aureus is a bacterium that colonizes the skin of approximately 30% of the human population and is a major cause of skin and soft tissue infections. Left untreated, these infections can become invasive, allowing S. aureus to enter the bloodstream and cause serious disease. S. aureus proliferation is controlled by the body’s innate immune system and by the microflora, including other staphylococci that reside on the skin. Indeed, to gain a competitive advantage several species with the coagulase-negative staphylococci (CoNS) secrete different effector molecules that inhibit S. aureus growth and virulence. For instance, auto inducing peptides secreted by CoNS block S. aureus quorum sensing and reduces its virulence, allowing these CoNS to better colonize the host. Other effector molecules secreted by CoNS directly kill S. aureus, including small antimicrobial peptides termed bacteriocins. Importantly, these CoNS have recently been used as bacteriotherapy in a phase I clinical trial to treat atopic dermatitis caused by S. aureus. Finally, a novel cyclic peptide lugdunin was discovered from Staphylococcus lugdunensis and has been shown to kill S. aureus and modulate the host immune response.
We hypothesized that other as yet undetermined anti-bacterial mechanisms exist within the CoNS and so we screened a panel of staphylococci we had in our laboratory for inhibitory activity by co-culturing them with S. aureus. In this screen, we came across a strain of Staphylococcus chromogenes that prevented S. aureus growth. S. chromogenes is a cause of subclinical mastitis in cows, in other words, causing infection without loss of milk production. Interestingly, studies have found an inverse correlation with the presence of S. chromogenes and S. aureus, suggesting S. chromogenes may hinder infection by S. aureus. It took approximately a year of experimentation but we eventually were able to determine that the inhibitory compound produced by S. chromogenes is the purine analog 6-thioguanine (6-TG), known to inhibit purine biosynthesis and a molecule used in humans to treat leukemias and inflammatory bowel disease.
Surprisingly, the genetic basis for 6-TG biosynthesis had been elucidated in the plant pathogen Erwinia amylovora although those studies did not characterize its possible antibacterial role. Nonetheless, we used this information to identify the genes required for its synthesis in S. chromogenes. Teaming up with Shayna Deecker, a doctoral student in Dr. Alex Ensminger’s laboratory at the University of Toronto, we identified the 6-TG biosynthetic operon in available genome sequences among four other coagulase-negative staphylococcal species. This suggested to us that 6-TG synthesis may be an important anti-bacterial strategy. In examining this directly in the laboratory, we found that 6-TG producing strains of staphylococci antagonized lesion formation by S. aureus in a murine co-infection model of subcutaneous infection.
With these results in hand, we were curious as to whether we could use 6-TG as an antimicrobial, or anti-virulence compound, against S. aureus skin infection. Strikingly, physiologically relevant amounts of the pure compound significantly reduced skin damage caused by S. aureus. To elucidate the mechanism of action, we used RNA-seq to determine the changes that 6-TG was exerting on S. aureus and found that it exerts many effects on the pathogen. First, it blocks de novo purine biosynthesis, thereby preventing DNA replication and bacterial growth. Moreover, it reduces transcription from genes encoding for the global virulence regulator agr and for ribosome synthesis, resulting in a significant reduction in toxin production.
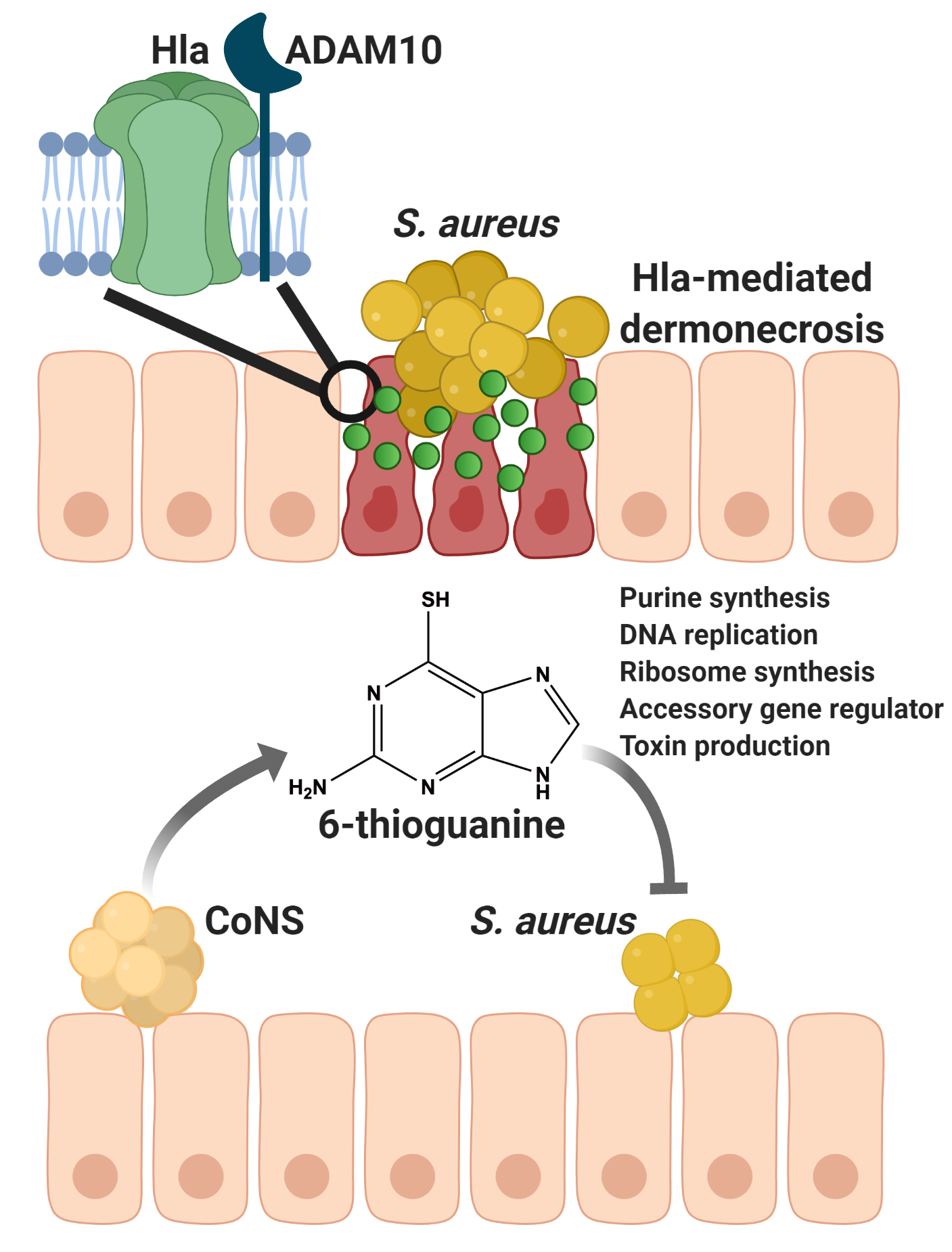
Treatment of S. aureus infections is complicated by the emergence and spread of multi-drug resistant S. aureus strains. As we strive to stay ahead of resistance, one route to the discovery of new antistaphylococcals is to continue to investigate genomes and metabolomes of the microflora, including other staphylococcal species, that are in constant competition with S. aureus in their natural environment. This has the potential to uncover not only new antibiotics, like the recently discovered lugdunin, but also anti-virulence compounds such as 6-TG.
The article can be accessed at the following link:
https://www.nature.com/articles/s41467-021-22175-3
Cover image created with https://biorender.com
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Healthy Aging
Publishing Model: Open Access
Deadline: Dec 31, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in