Certain concepts in biology – evolution by natural selection, for instance – have taken decades to become entrenched, only to then seem so intuitive that it’s hard to understand why people needed much convincing at all about their veracity. This may be a personal shortcoming, but I recall having a difficult time wrapping my head around the fact that there was debate in the early 20th century between mutationists and selectionists. “But selection acts on genetic variation,” I thought in utter disbelief at the obtuse pedantry of my scientific predecessors. Oh, the luxury of hindsight!
In a few years, or perhaps already, a new generation of biologists may similarly look back in surprise at a time when the phrase “junk DNA” was thrown around with casual certainty when speaking of transposons and other seemingly non-functional parts of the genome. I may be among the last of the old-timers, who first saw a study demonstrating host recruitment of transposons in 2016 and registered awe.
It turns out it isn’t always simple to identify or classify “junk” in genomes.
Transposons are mobile genetic elements that are often described as selfish because of their tendency to “jump” and propagate themselves throughout the host genome, risking disruption by insertion into functional regions like genes. Host genomes typically adapt over time to restrict the dispersal of transposons through various means, but past transposon expansions have left their imprint on the genomes of almost all known organisms.
In humans, transposons comprise about 45% of the entire genome, and as it currently stands in the arms race between these jumping elements and host genome integrity, the vast majority of them have been rendered inactive. But beyond simply neutralizing their enemies, the triumph of our evolutionary ancestors extended to also recycling some of their genomic remains to evolve novel functions and traits, some of which set us on the path toward becoming human.
A parasite of parasites
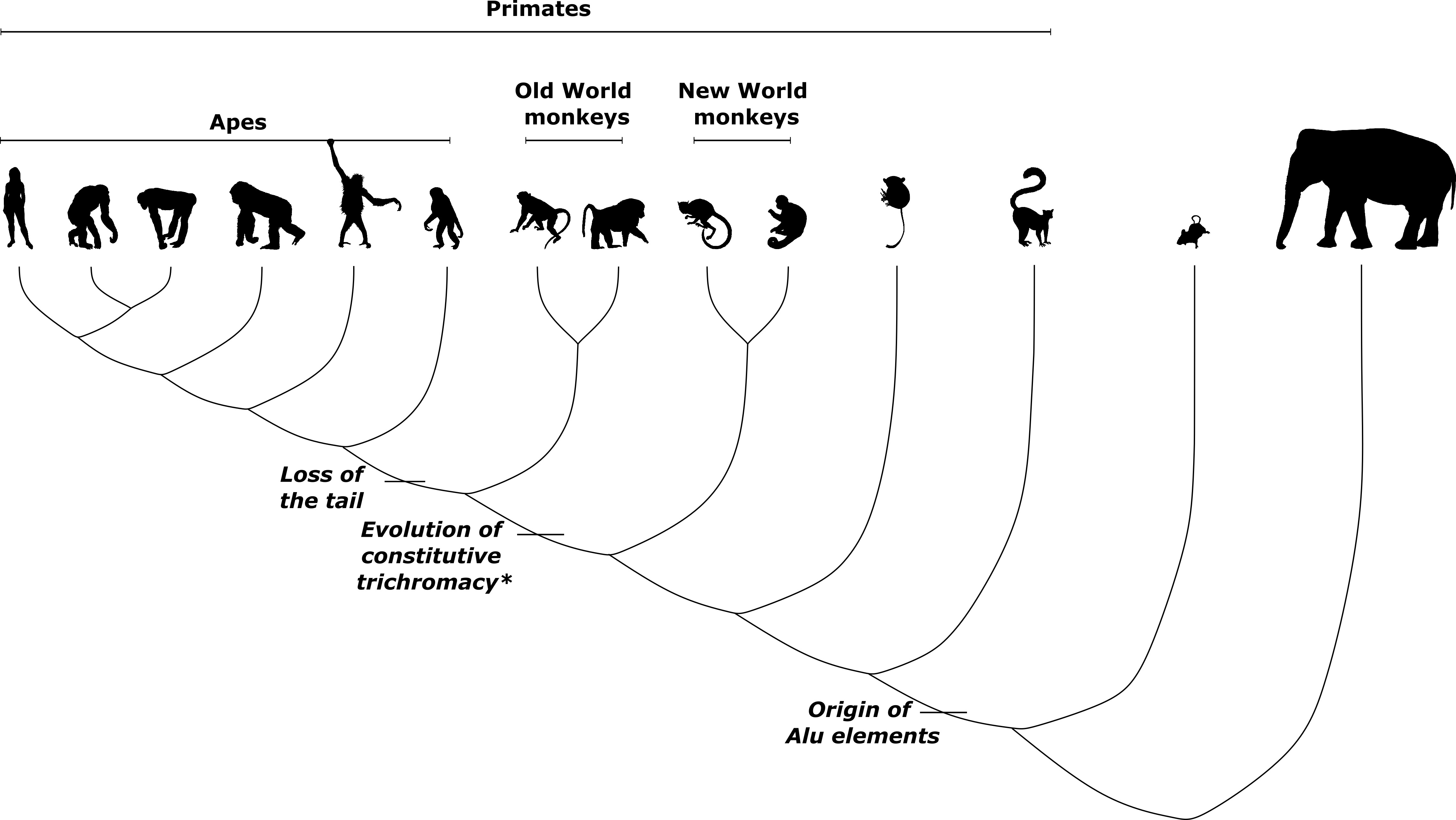
Transposons come in a variety of flavors, classified according to distinct modes of genome-wide dispersal. In recent years, we have learnt that transposons belonging to essentially every class have been coopted in animals and plants to perform roles in gene regulation, immunity, and development. One group of transposons, Alu elements, recently made the news by featuring centrally in a new preprint on the loss of the tail in apes – which includes us.
Alu belongs to a class of transposons known as short interspersed nuclear elements or SINEs. Transposons rely on enzymes – usually ones they encode themselves – to cut or copy themselves and mediate their insertion into new parts of the genome. SINEs are defined by the absence of their own enzymes, relying instead on those produced by other transposons for their dispersal – making them true parasites of parasites.
Alu elements are thought to have emerged in an ancestral primate about 65 million years ago, subsequently undergoing multiple expansions. The human genome contains more than 1 million copies of Alu, but only about 5,000 are unique to humans, suggesting that much of the action took place before the origin of hominins.
With an average length of about 300 base pairs – the human genome is about 3 billion base pairs long – Alu elements make up 10% of the genome, contributing to disease and genomic disruption, but also helping to bring about some dramatic transitions in our evolutionary past.
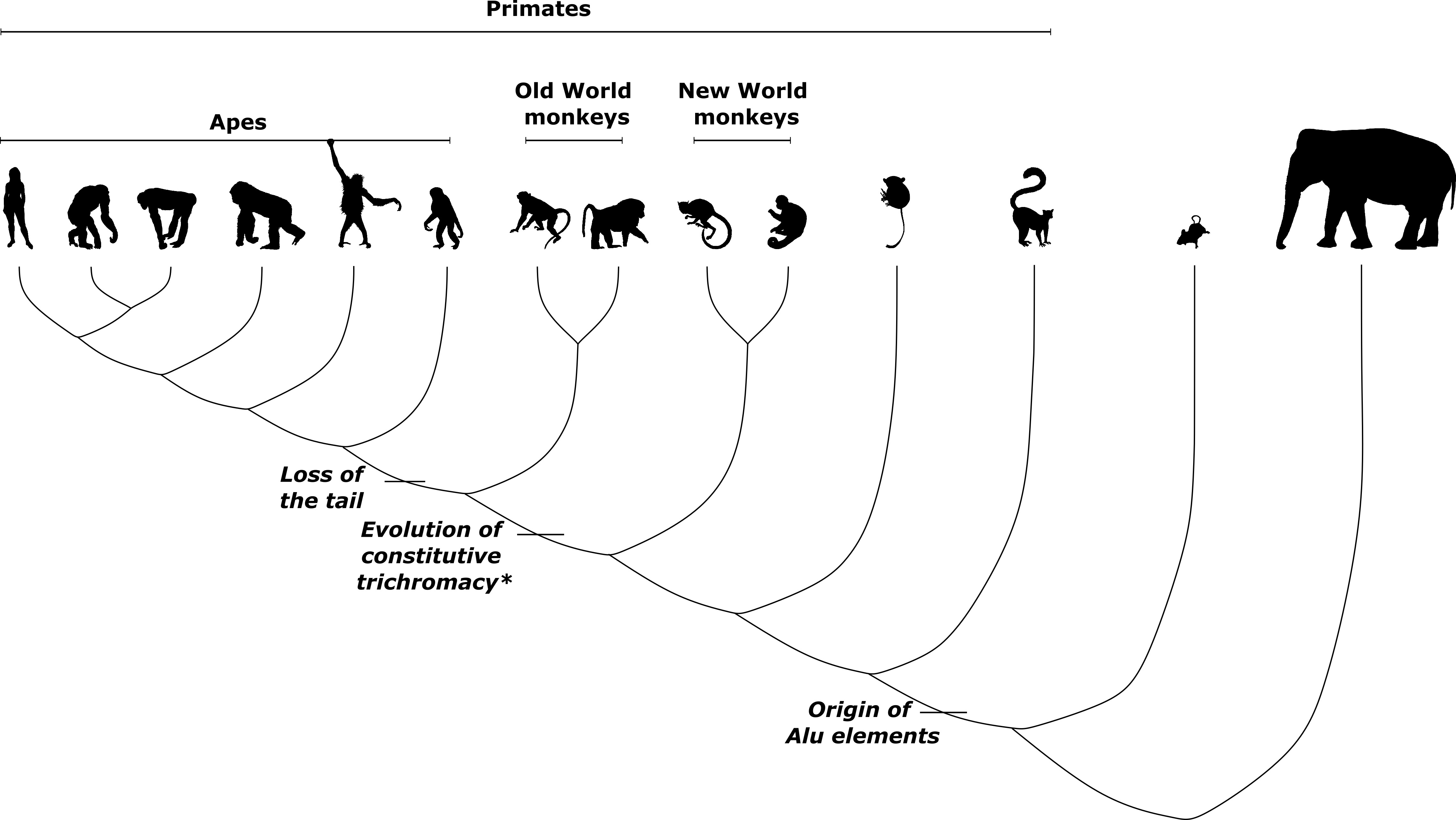
* Polymorphic and female-specific trichromacy exists in New World monkeys and prosimians, while constitutive trichromacy independently evolved in the ancestor of howler monkeys, a New World taxon. Animal silhouettes from PhyloPic
Seeing red again
Humans, along with some other primates, are unique among mammals in their ability to perceive color in three dimensions. The evolution of trichromatic color vision in primates is something of a reacquisition of an ancient trait: the common ancestor of tetrapods – which also includes birds, amphibians, and reptiles – is thought to have had tetrachromatic vision, which is still retained in all tetrapods except mammals. The first mammals evolved in the age of dinosaurs almost 200 million years ago. Keeping largely to the shadows, these nocturnal shrew-like creatures came to lose much of their color vision. To this day, non-primate mammals only have dichromatic vision, leaving them red-green colorblind.
In sexually reproducing organisms, recombination between paternal and maternal chromosomes allows a synthesis of parental genetic content to be passed on in the gametes. Such recombination relies on regions of high similarity between paternal and maternal homologous chromosomes, so that equivalent regions can come together and mediate exchange. Alu elements represent similar sequences scattered throughout chromosomes and increase the risk of asymmetrical recombination. Occasionally, this can lead to duplication of genomic regions on one chromosome, coupled with a deletion on its recombining partner.
In a 1999 study, Dulai et al. proposed that the common ancestor of apes and Old World monkeys likely underwent such an Alu-mediated duplication about 40 million years ago, based on the presence of ancient Alu elements near the duplication in humans and monkeys. This yielded a new photopigment-producing opsin gene which evolved to produce, alongside two ancestral opsins, the ability to see red and green that is shared today among Old World primates.
Transposons as sources of genetic variation
The trichromatic vision story demonstrates one of several ways in which transposons may spur functional innovation in the host. In this instance, the role of the Alu elements was simply in mediating a historical accident – the duplication – by virtue of its tendency to proliferate throughout the genome. Alu elements have also been found to be recruited in gene regulatory functions.
One of the ways in which hosts control transposon activity is by adding a methyl group to transposon sequences. Methylation is used across the genome to suppress gene expression – maybe this too represents a cooption, from a strategy that originally evolved to control transposons. In any case, if a host gene located near a transposon becomes suppressed as a byproduct of transposon methylation, and if the resulting change in gene expression is adaptive, that represents recruitment of the transposon as a host regulatory sequence – as might have taken place with some Alu elements. Sequences found within Alu elements have also been adopted as binding sites for transcription factors in promoter regions, with Alu transposition possibly facilitating rapid genome-wide spread of these binding sites.
Alu elements, and transposons in general, appear to have frequently acted as sources of genetic variation that were exploited by host genomes in the evolution of functional novelty.
The lost tail of two Alu elements
Genes are a mosaic of protein-coding sequences called exons and non-coding regions called introns. Introns are spliced out during translation of mRNA into protein, but exons too can be similarly removed. Alternative splicing of mRNA enables the use of different combinations of exons from the same gene to express several different versions of a protein. Transposons occurring within introns have been shown to alter alternative splicing, often by disrupting splice sites – which define the boundaries between retained and discarded regions.
A recent preprint by Xia et al. found alternative splicing mediated by Alu elements – which has been referred to as Aluternative splicing by scientists who presumably couldn’t resist – to be behind a long-standing mystery in primate evolution: the loss of the tail in the ancestor of modern apes about 25 million years ago.
Comparing between apes, which include human, chimpanzee, bonobo, gorilla, orangutan, and gibbon, and Old World monkeys, the authors identified an ape-specific Alu element in an intron in the TBXT gene, which has been linked to tail development in cats, dogs, and mice. The authors provide evidence that this Alu element can form a loop with another neighboring Alu element – this one is older and not specific to apes – upon transcription, thereby trapping an exon and causing it to be spliced out. The ability to produce different splice variants of TBXT, and the truncated form in particular, may be driving the tail-loss phenotype.
The authors sought to validate this in a mouse model by simultaneously expressing truncated – with the equivalent exon removed to mimic the alternatively spliced primate TBXT – and full-length forms of the mouse TBXT gene. They found that the truncated form was sufficient in the presence of the full-length form to produce an absent or shortened tail in the mice.
The ape-specific Alu insertion may thus have markedly reduced tail length or occasionally removed the tail in the ape ancestor, but other genetic changes were likely necessary to stabilize the complete loss of the tail. Even so, such a large-effect phenotypic change resulting from the insertion suggests that tail loss was rapid rather than gradual, possibly helping the ape ancestor adapt to new habitat challenges.
Of course, all of the ways in which Alu elements have led to novel functions also enable the disruption of existing functions, occasionally resulting in disease. For example, Alu-mediated genomic rearrangements have been observed to cause diseases like hemophilia A and Alström syndrome, and Alu-mediated alternative splicing in the CD58 gene may increase the risk of multiple sclerosis.
The transposons that do, however, get coopted into actively functional roles are less likely to be lost from the host genome, representing evolutionary success for these selfish genetic elements. You could do a lot worse than being present in more than 7 billion individuals spanning the entire planet.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in